Abstract
The extreme rarity of asexual vertebrates in nature is generally explained by genomic decay due to absence of meiotic recombination, thus leading to extinction of such lineages. We explore features of a vertebrate asexual genome, the Amazon molly, Poecilia formosa, and find few signs of genetic degeneration but unique genetic variability and ongoing evolution. We uncovered a substantial clonal polymorphism and, as a conserved feature from its interspecific hybrid origin, a 10-fold higher heterozygosity than in the sexual parental species. These characteristics seem to be a principal reason for the unpredicted fitness of this asexual vertebrate. Our data suggest that asexual vertebrate lineages are scarce not because they are at a disadvantage, but because the genomic combinations required to bypass meiosis and to make up a functioning hybrid genome are rarely met in nature.
Similar content being viewed by others
Main
Asexual lineages present a paradox to biology. Overwhelmingly, theory predicts that asexual reproduction would have several main disadvantages. First, the classical model of demise, Muller’s ratchet1,2, states that deleterious mutations cannot be purged without meiosis, and their accumulation will lead to genomic decay and eventually extinction3,4. Thus, obligate asexuals are predicted to be evolutionarily short-lived: a strictly clonal vertebrate population is unlikely to survive more than 104–105 generations in the face of incessant mutational pressure4. Second, the Red Queen hypothesis5,6 predicts that an absence of meiosis and formation of new genotypes in the zygote hinders the creation of genetic diversity, which is a precondition for adaptation to changes in the physical and biological environment. Third, recombination can uncouple beneficial and deleterious mutations, allowing selection to proceed more effectively with sex than without7. On the other hand, eukaryotic parthenogenetic lineages are all-female. Because they do not have to produce males, 100% of their offspring contribute to population growth, giving them a two-fold reproductive rate compared with sexual propagation8. However, all theories agree that the disadvantages of asexual propagation quickly outweigh this advantage, and that clonality should eventually lead to extinction3,9. Mixed support for this prediction exists: although some asexual species, for example, the obligate asexual waterflea, show deleterious mutation accumulation and are evolutionary extremely short-lived10, other asexuals are older than predicted and successful colonizers in their natural habitats11,12,13,14.
Clonal lineages are numerous amongst unicellular eukaryotes15, plants16 and invertebrates17,18, but vertebrates were long thought to be unable to exist as asexuals. However, in 1932, the Amazon molly, Poecilia formosa, was the first unisexual vertebrate to be described19, followed by the discovery of more than 50 naturally occurring fish, amphibian and reptile species, and more than 50 others that at least occasionally reproduce clonally20. P. formosa became one of the paradigmatic cases that appear to violate the age predictions of Muller’s ratchet and the dynamics of the Red Queen hypothesis. It is a highly successful colonizer of diverse habitats over a wide geographical range and mitochondrial DNA-based estimates postulated a much longer existence, exceeding the theoretical extinction time10,12,14.
As with most vertebrate asexuals, P. formosa shows features of an interspecific hybrid from distantly related sexual species, predicted to be an Atlantic molly (P. mexicana) mating with a Sailfin molly (P. latipinna)14,21. Its mode of reproduction is gynogenesis22, an elaborate form of parthenogenesis where sperm from males of sympatric sexual species is ‘stolen’ (kleptospermy) to trigger embryonic development from unreduced diploid eggs (Fig. 1a). The sperm DNA is usually excluded from the developing egg; thus offspring are true clones of their mothers11. In this study we investigate the ancestral history of the P. formosa genome and reveal its novel features.
a, Through hybridization of a female P. mexicana (genome composition m/m) and a male P. latipinna (l/l), a hybrid (m/l) was produced that continued to reproduce through gynogenesis. Sperm from males of either one of the parental species (P. mexicana, haploid m genome, or P. latipinna, haploid l genome) or another sympatric species, P. latipunctata22, is used to trigger parthenogenetic development of the diploid oocyte (m/l), but the genetic content of the sperm is excluded (red cross) from the oocyte. In the laboratory, other Poecilia host males, for instance the ornamental black molly (b/b), are used. b, Map of the sampling sites and origins of stocks of P. formosa, P. mexicana and P. latipinna specimens used in this study. Multicoloured circles indicate sympatry of two (location 10, 16) or all three (location 9) species. 1, Pme_Cav; 2, Pme_Azu; 3, Pme_Ver; 4, Pfo_Ta-1, Pfo_Ta-2, Pfo_Ta-3, Pfo_Ta-4; 5, Pfo_Ma; 6, Pfo_Vic-1, Pfo_Vic-2; 7, Pfo_Lim; 8, Pfo_Cha, Pla_Cha, Pme_Cha; 9, Pla_Tam; 10, Pfo_Gua, Pme_Gua; 11, Pfo_Pad; 12, Pfo_Bar-1, Pfo_Bar-2, Pfo_Bar-3; 13, Pfo_Br, Pfo_Br-1, Pfo_Br-2; 14, Pfo_D1; 15, Pfo_Olm, Pla_Olm; 16, Pfo_SM, Pla_SM; 17, Pla_Flo; 18, Pla_Fld. For location details see Supplementary Table 2. The natural range of the three species is indicated by the coloured lines. At location 16, P. formosa has been introduced by human activities. Map dimensions: ca. 102–78° W longitude; ca. 17–32° N latitude.
Results
Genome assembly and gene annotation
To understand how a vertebrate genome evolves when the maternal genome is simply copied from generation to generation we sequenced a single P. formosa female. Total sequence coverage of 95-fold produced an assembly (Poecilia_formosa-5.1.2) with N50 contig and scaffold lengths of 57 kb and 1.57 Mb, respectively (Supplementary Tables 1 and 2). Two independent sets of protein coding genes (Ensembl: 23,615; NCBI: 25,474) were produced from the assembly. Both sets of genes were used throughout these analyses.
To measure the parental genome contributions to P. formosa we assembled P. latipinna and P. mexicana genomes using mostly assisted alignment23 to the Poecilia_formosa-5.1.2 reference (P_latipinna-1.0 and P_mexicana-1.0) to total sizes of 815 and 803 Mb, respectively (Supplementary Tables 1 and 2). The number of protein-coding genes for each was similar to P. formosa: 25,220 for P. latipinna and 25,341 for P. mexicana. For population-level analyses we sequenced the genomes of an additional 19 P. formosa, 5 P. latipinna, and 4 P. mexicana collected from various locations over each species’ range (Fig. 1b, Supplementary Table 3).
Transposable element history and activity
The absence of meiosis has been hypothesized to impact transposable element (TE) colonization of the genome. Theories predict that TEs should be either very few in number or absent from the genomes of asexuals because new integrations after zygote formation cannot occur and existing ones decay24; or that TEs accumulate at unrestrained rates after the emergence of asexuality, because as the genome is subject to Muller’s ratchet they cannot be eliminated through recombination25. This could eventually lead to extinction of asexual lineages. We detected no relevant difference in TE composition (Supplementary Note 1, Supplementary Tables 4 and 5) and transposition history between the genomes of P. formosa and the parentals (Fig. 2a). Most superfamilies are rather ancient and expanded long before the origin of P. formosa. The presence of TE sequences in the transcriptome indicated that some TEs are still active (Supplementary Note 1, Supplementary Table 6), in particular Gypsy elements (Fig. 2b), supported by genomic evidence for post-hybridization transposition events in P. formosa (Supplementary Note 1).
a, Copy-divergence analysis of TE classes in five poeciliid genomes, based on Kimura 2 parameter distances. The percentages of TEs in genomes (y axis) are clustered based on their Kimura values (x axis; K values from 0 to 50; arbitrary values). Older copies are located on the right side of the graphs while recent copies are located on the left side. DNA transposons (TcMariner, hAT and all other DNA superfamilies) are shown in red tones. LTR retrotransposons (Gypsy and all others) are in green tones. LINE and SINE retrotransposons are in blue and purple tones, respectively. b, The proportion of TE superfamily representation in the genome and the transcriptome of P. formosa. The proportion of each TE superfamily was initially calculated as (% of TE superfamily × 100) / total % of TEs in the genome or transcriptome, and then for the spider graph transformed to log10 values. The expression of Gypsy elements might be the result of their activity rather than of general background expression because their relative fraction is notably higher in the transcriptome than in the genome.
In summary, TEs show none of the predicted consequences of their host genome reproducing in the absence of recombination, but surprisingly look much like the original parental genomes and more broadly like other related teleosts. No increase in the TE load was detected in asexual arthropod lineages, including the water flea26. In the genome of the asexual rotifer Adineta vaga a rather low percentage of TEs was found17, which seems to be mainly shaped by an ongoing process of TE acquisition by horizontal gene transfer and the predicted loss of ancestral elements. In P. formosa we detected no TE sequences, whose absence from the parental genomes would indicate horizontal gene transfer.
Gene evolution
The standard genome-wide analyses on gene evolution (positive selection, gene duplication and gene family expansion/contraction; Supplementary Note 2, Supplementary Fig. 1, Supplementary Tables 7–9) did not reveal any unusual features of the P. formosa genome in comparison with other teleosts that reproduce sexually. Genes associated with functions that are assumed to be non-essential and thus dispensable for asexual female reproduction, including genes pertinent to spermatogenesis and meiosis, showed no damaging variants (such as frameshifts, premature stops and high sequence divergence; Supplementary Table 10–12). Similarly, we find the average number of loss of function variants (LoFs) in P. formosa, compared to the parental species (Supplementary Note 2, Supplementary Table 13), to be slightly less. Moreover, LoF counts are all in the range of what has been reported for the genomes of related sexual Poeciliid species23. Genome-wide analyses to detect genes showing signs of relaxation from purifying selection revealed seven such genes in P. formosa, but also seven in P. mexicana and 11 in P. latipinna (Supplementary Table 14). For the opposite phenomenon, positive selection, we also observe similar counts: P. formosa (24), P. mexicana (22) and P. latipinna (27) (Supplementary Table 15). In the genome of P. formosa a total of 211 non-processed or duplicated pseudogenes (not overlapping with LoFs) were recorded, similar to its sexual relatives (P. mexicana 268, P. latipinna 266, P. reticulata 278). In summary, asexual genic evolution of the Amazon molly is not obviously different from the sexual parental species.
Gene copy number variation (CNV) impacts genome evolution and adaptation27. The extent of CNVs is not significantly different between the Amazon molly and its parental species (Supplementary Note 2) and gene ontology (GO) term analysis (Supplementary Tables 16–21) revealed that several of these variants are most probably expansions of certain TEs.
Segmental duplications of genes linked in a common process can give hints on specific features of P. formosa. The asexual reproduction mode, apomixis, requires that diploid oocytes are formed without meiosis. Disturbance of meiosis leads to mis-segregation of chromosomes and can even turn meiosis I into a normal mitosis28. Several genes involved in the meiosis-specific separation of homologous chromosomes show CNV in P. formosa compared with the parentals and other sexual species (Supplementary Note 3, Supplementary Table 22). In particular, genes for cyclin-dependent kinase 1, an essential regular of meiotic kinetochore–microtubuli capture, and its oocyte-specific activator, Ringo/speedy, are present in multiple copies. We hypothesize that the expression imbalance brought about by such CNVs disturbs the proper establishment of kinetochore unipolarity in meiosis I and induces a mitotic division, thereby generating diploid, ameiotic oocytes.
Ancestry and evolutionary age of the Amazon molly
A low level of genomic decay in the asexual species could be expected if new lineages are repeatedly produced from hybridization of the parental species. Previous studies based on small fractions of mitochondrial or nuclear genomes showed conflicting origins as to a single14 or multiple F1 hybrids or backcrosses14,29.
We performed ancestry estimates for P. formosa using our high-quality single-nucleotide polymorphism (SNP) data set (292,324 sites). In total 53,175 sites display fixed nucleotide differences between the parentals, of which 47,359 were inferred to be heterozygous in P. formosa for the two parental alleles confirming an F1 hybrid origin. All P. formosa isolates consistently display a heterozygosity (H) index of 0.5 (Supplementary Table 23).
We then reconstructed the complete mitochondrial genomes of all samples. Haplotypes of P. formosa were more closely related to P. mexicana than to P latipinna (Supplementary Note 4, Supplementary Fig. 2) confirming P. mexicana as the maternal ancestor. Phylogenetic trees (Fig. 3) revealed that all P. formosa are united in a highly supported single clade that is consistent with a single maternal (mitochondrial) origin. Using a time-calibrated Bayesian phylogenetic tree of mitogenomes (Supplementary Note 4, Supplementary Fig. 3) we estimate a minimum age for P. formosa of at least 100,000 years, exceeding by far the predicted age for extinction from previous calculations based on the threat from Muller’s ratchet12.
Maximum likelihood phylogenetic tree based on 35 complete mitochondrial genomes (16,587 bp), obtained with the program PhyML. Samples labeled ‘_S’ represent reference mitochondrial genomes. All other mitochondrial genomes were assembled from whole genome sequencing reads. The numbers above the branches represent bootstrap values based on 100 resampled data sets. The branching point for P. formosa from P. mexicana was estimated at about 100,000 years ago; for details see Supplementary Note 4 and Supplementary Fig. 3.
Gene conversion
Given a considerable age and single origin we looked for possible mechanisms that could affect the extent of the predicted negative consequences of asexual reproduction. One explanation could be that enhanced gene conversion reduces genetic decay. Homogenization of local sequence tracks through gene conversion will limit the spread of LoFs by exposing mutations to selection or overwriting them entirely. We find a rate of 3 × 10−8 conversion events per generation per site in P. formosa (Supplementary Note 5), which is in the range of other asexually reproducing species30 but still two orders of magnitude lower than for sexual species31,32.
Paternal introgression
Sustained genomic diversity and refreshing alleles damaged by the process of Muller’s ratchet could come from exceptional uptake of paternal sperm DNA. Microchromosomes, which occur in some P. formosa individuals and are inherited like B-chromosomes33,34, have been explained to be derived from incomplete removal of paternal DNA after insemination of P. formosa oocytes. This hypothesis critically depends on the assumption that microchromosomes are indeed of paternal origin and that they contain coding DNA. Hence, we measured non-meiotic introgression from host males, hypothesizing that within the P. formosa genome allele imbalance will exist in regions that experienced introgression due to additional copies present on the microchromosomes (Supplementary Note 6). We detected 19 putatively introgressed scaffolds across 5 P. formosa samples (Fig. 4, Supplementary Table 24). The total introgressed scaffold size ranged from 0.33 to 8.1 Mb, approximately 1% of the genome, adding up to hundreds of protein coding genes (with no obvious enrichment of GO terms; Supplementary Note 6, Supplementary Tables 25 and 26), microRNAs and other functional units from a recombining parental genome.
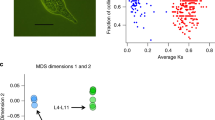
Evidence supporting paternal introgression of scaffold KI519701.1 into sample Pfo-Tan-2. a, Reference allele frequency distribution at called heterozygous sites of scaffold KI519701.1. In the absence of introgression, the reference allele frequency at heterozygous sites (density) follows a normal distribution centred at 0.5. The distribution of sample Pfo-Tan-2 deviates from the theoretical normal distribution centre, and instead presents two peaks at frequencies of ~0.33 and ~0.66, which is consistent with the presence of 2 copies of scaffold KI519701.1 that originate from one parental species and 1 copy of scaffold KI519701.1 from the other parental species. b, Copy number distributions across 1 kb repeat-free windows in scaffold KI519701.1. All samples except Pfo-Tan-2 show a copy number distribution that is centred at 2, suggesting that they are diploid. The distribution of Pfo-Tan-2 is centred at a copy number of 3, indicating it is triploid at this scaffold. c, Median frequency of P. mexicana alleles at heterozygous sites. For diploid organisms, the frequency of any allele at heterozygous sites should be centred at 0.5. However, for sample Pfo-Tan-2, the frequency of the P. mexicana-derived allele at heterozygous sites is 0.66, indicating that the introgressed scaffold KI519701.1 in Pfo-Tan-2 was derived from P. mexicana. The legend applies to all panels.
Hybrid genome constitution
One reason for the fitness and ecological success of P. formosa could be that heterozygosity of the interspecies hybrid is maintained in the asexual lineage, described in the ‘frozen hybrid genome’ hypothesis.35. Indeed, heterozygosity in the parental species was on average approximately tenfold lower than in P. formosa (Supplementary Table 27). Next we reconstructed haplotypes for all isolates to examine phylogenetic relationships (Supplementary Note 7). We observe two strongly supported clades, each consisting of the parental haplotypes and those P. formosa haplotypes derived from this parent (Supplementary Fig. 4), again supporting the single origin hypothesis.
Allele-specific gene expression analysis from three different organs of P. formosa revealed that only 1.2 to 4.1% of genes had an expression bias towards one of the parental alleles, whereas for the overwhelming majority of genes there is about equal contribution from both parental alleles (Supplementary Fig. 5). This demonstrates that P. formosa is not only a genomic but also a ‘functional’ hybrid.
To test if the increased level of heterozygosity in P. formosa might counter the predicted detriment under the Red Queen hypothesis, we looked at a system that in clonal organisms should be at major disadvantage, namely immune genotypic variability. The cell-mediated adaptive immune response is regulated by the major histocompatibility complex (MHC)36. Variability in these multicopy genes is positively correlated with immune competence37. Remarkably, we discovered high diversity in MHC class I genes (80 different alleles in 20 individuals) (Fig. 5a). MHC class II genes were less variable, but still 36 different alleles were found (Fig. 5b). MHC copy number in P. formosa is generally in the same range as in its sexual ancestors and in some clones even exceeded average levels (up to 13) with higher variation than expected from previous studies on a limited data set38,39. However, one clone (F in Fig. 5a) had only two MHC class I genes, suggesting that some functional gene copies might have been lost in this lineage. We also examined 15 critical members of the innate immune system for variability. In the sexual parental genomes within-individual interallelic diversity was low, and for the majority only a single allele was retrieved. However, P. formosa always comprised at each gene locus two considerably different alleles (Supplementary Table 28, see also Fig. 6 for neighbour-joining amino acid trees of toll-like receptors). Alleles derived from one parent were similar among the P. formosa individuals and clearly of monophyletic origin. The genetic distance of immune gene sequences within individuals was significantly higher for P. formosa (Supplementary Fig. 6a). Overall, the immune system genes of P. formosa present an unexpected high level of genetic variability.
a,b, For MHC class I (a) and MHC class II (b) gene alleles we plotted individuals that are sorted geographically from north to south on the x axis. Unique alleles are shown in grey, the coloured bars depict alleles that were found in more than one individual. Coloured letters indicate the mitochondrial haplotypes that were found in more than one individual (see Supplementary Fig. 2). Bars indicate species: blue, P. latipinna; red, P. formosa; orange, P. mexicana.
Neighbour-joining trees (unrooted) are shown based on the amino acid sequences of the largest open reading frame of innate immune genes TLR 1 (802aa) and TLR 5 (759aa). Only bootstrap support values >50% are shown (1,000 randomizations). Species are marked by coloured squares after the sample names. The colour used for the names corresponds to the mt haplotype (see Supplementary Fig. 2; black, individual haplotype). For both TLRs only very few individuals from each of the parental species showed two different alleles, whereas each P. formosa individual always has two clearly divergent alleles (L indicates P. latipinna origin, M indicates P. mexicana origin), reflecting the hybrid origin of the species.
The immune genes also demonstrate that genetic variation within P. formosa is not restricted to intra-individual heterozygosity but that one and the same parental gene copy exists in many alleles. The phylogenetic trees obtained for each of the immune genes indicate a single origin for each group of the parental alleles. Consequently, such differences between alleles must be due to mutations. Nucleotide sequence differences in the open reading frame were generally much lower than in the noncoding regions (Supplementary Fig. 6b). With the exception of CD59, which is under positive selection in all three species, most immune genes are under strong purifying selection (Supplementary Table 29).
Discussion
Analysis of the Amazon molly genome and comparison to its sexual parental species uncovered unanticipated features that may change our view on asexual organisms that practice gynogenesis. Unexpectedly, we found no widespread signs of genomic decay. This is not explained by a recent origin because our age calculations of about 100,000 years from whole mitochondrial and nuclear genome data substantiate earlier estimates of 120,000–280,000 years11. Thus, given a generation time of 3–4 months, P. formosa has existed for approximately 500,000 generations, and has survived several-fold beyond its Muller’s ratchet-based predicted extinction12.
Despite such an ancient origin we find genes that serve organs or processes that are no longer in use in the all-female fish, such as spermatogenesis, male development and meiosis genes, are not corrupted. The Mexican tetra, which inhabits a similar natural range to P. formosa, has evolved cave populations from surface fish, some not older than 30,000 years40, yet all show total loss of organs no longer in use, such as eyes. A similar trait loss situation could have been expected in the Amazon molly, but was not observed. Another simple explanation for the much lower level of genomic decay than predicted from mathematical models may be that not enough time has elapsed. In this case the Amazon molly genome sets a new time point for what is ‘not old enough’ for a vertebrate genome to undergo genetic degeneration. This result also has implications for ‘regressive’ vertebrate systems like the cavefish as it provides a baseline for how many generations have passed without any signs of neutral morphological and genetic degeneration or regression appearing. It could make an argument that comparatively fast trait loss in the cave environment is based on selection and standing variation in cavefish.
Another explanation is introgression of DNA into an asexual lineage that represents a unidirectional flow of genetic material that can compensate for harmful alleles accumulated in the genome. In the ancient asexual bdelloid rotifers occasional parasexual transfer of genetic material between individuals, known as horizontal gene transfer in bacteria, generates divergence with up to 10% of genes of putative non-metazoan origin41. Unisexual salamanders of the Ambystoma jeffersonianum-laterale complex occasionally interrupt clonality by ‘stealing’ paternal DNA from sympatric sexual species (kleptogenesis) and loss of part of the clonal genome20,42. In P. formosa our sequence-level analysis detected ‘genetic addition’ as the mode for introgression of subgenomic amounts of DNA in about 25% of the samples at a much higher resolution than is possible with cytogenetic methods, suggesting paternal introgression occurs more frequent than initially thought. These introgressed sequences are derived from genic regions of the paternal genome.
Most evaluations of Muller’s ratchet use estimates of important parameters, such as mutation rate and population size, which are unknown for P. formosa. Lower actual mutation rates and higher numbers of individuals might contribute to the discrepancy between the expected and observed rates of genomic decay. Although such differences from assumed values are difficult to evaluate, our genomic data now provide indications about processes that can affect Muller’s ratchet. Introgression has not been considered in calculations of the expected time to extinction (Tex) for asexual species, including P. formosa. We recalculated predicted extinction times for the Amazon molly according to a published method12, but considered relief from genomic decay when different amounts of genetic material are introgressed from sexual parental species at the frequency observed in our data. This resulted in a slight increase in Tex, (Supplementary Fig. 7). However, paternal introgression alone does not provide sufficient explanation for how long P. formosa has outlived its predicted extinction time. Other genomic features of P. formosa that are more difficult to quantify are traces of past events of loss of heterozygosity. It has been concluded that loss of heterozygosity can have a similar beneficial effect to segregation during meiosis43, and consequently could also counteract the ratchet.
The Red Queen hypothesis is often touted as an explanation for why sexual reproduction persists, as it posits that recombination allows species to maintain genome diversity against ecological stressors, such as pathogens. Without recombination, how do rare asexual vertebrate species remain extant? We propose that the available standing genetic variation is a sufficient starting point. The evolutionary advantage of such hybrid vigour has been shown in hybridogenetic frogs44 and it is noteworthy that all asexual vertebrates are of hybrid origin20. Our results provide support for earlier predictions that being a ‘frozen hybrid’45 with elevated heterozygosity provides a fitness advantage (assuming polymorphisms improve fitness) and a possible resource for responses to natural selection. Moreover through de novo mutations new clones can arise from this original hybrid genome.
Interacting effects of parasites and the accumulation of mutations were shown by computer simulation to enhance Muller’s ratchet in a population that experiences Red Queen dynamics46. Our genome analyses uncovered—in contrast to predictions of the Red Queen model for asexuals—a high genomic diversity. Field studies revealed that P. formosa does not have a higher parasite load than sympatric sexual species47. We therefore assume that the ratchet for Amazon mollies clicks at a lower speed than projected under the severe conditions of the hypothesis.
We propose that genetic diversity between clones offers at minimum a short-term benefit to the asexual species in coping with environmental challenges. Those clones that acquired new adaptive mutations will thrive, while others that are less fit, like the one with only two MHC class I genes, will disappear. On this basis, we posit that in the absence of recombination P. formosa can evolve by clonal selection of naturally occurring mutations and competition between clones. Because Amazon mollies do not have to pay the ‘two-fold costs of sex’8, they have an increased population growth rate and can more quickly reach large population sizes12,14. Sperm-dependent parthenogens hinge on their ecologically similar sexual host but exhibit some niche separation, as they otherwise are expected to out-compete them and ensure their own demise48. In large populations of P. formosa multiple clones can be maintained, favouring selection of advantageous clones and clearing of less fit ones.
All known obligate asexual vertebrates are hybrids and we provide evidence that a hybrid genome might be one driver of fitness of asexual lineages. There is increasing evidence that interspecific hybridization is much more frequent than previously thought; it is estimated that approximately 10% of animal species hybridize regularly with at least one other species49. Given that even after 500,000 generations in the absence of recombination the Amazon molly does quite well, one might ask why clonal vertebrates are so rare, and in particular why P. formosa arose only once despite the continuing co-occurrence of the parental species? So far, all attempts at a laboratory synthesis of P. formosa by crossing the parental species have failed33,50. Combining just the right genotypes may have been key to allow asexual reproduction to occur. In interspecific hybrids, certain combinations of parental genes can lead to Dobzhanski–Muller incompatibilities, thus many hybrid genotypes will not be favourable for generating new species51,52. In addition, finding the right combination of immune genes might be a problem. While immune gene diversity is important, too much diversity can be disadvantageous by either hindering effective pathogen detection53,54 or triggering autoimmune diseases55. In P. formosa, however, no evidence of reduced pathogen resistance or autoimmune disease is found, indicating that their highly diverse immune genes are compatible47,56.
Taken together, we favour a ‘rare formation hypothesis’ specifying that clonal species might not be rare because of their inferiority to sexual species, but because the genomic combinations that allow successful survival and reproduction are very specific14.
Methods
Biological material
The fish used in this study from aquaria housed stocks were kept and sampled in accordance with the applicable European Union and national German legislation governing animal experimentation. We hold an authorization (568/300-1870/13) from the Veterinary Office of the District Government of Lower Franconia, Germany, in accordance with the German Animal Protection Law (TierSchG).
Sequencing and assembly
The P. formosa DNA for Illumina shotgun sequencing was derived from a single adult female (Pfo_Bar-1) from a clonal line established from a fish collected in 1996 in the Rio Purification near Barretal, Tamaulipas, Mexico. Total sequence genome coverage on the Illumina HiSeq 2000 instrument was ~95× with tiered library insert sizes of pre-determined sequence coverage for each (45× fragments, 45 × 3 kb, 5 × 8 kb and 0.05 × 40 kb). All sequences were assembled using ALLPATHS57 with default parameters. Assembly connectivity was further improved with the external scaffolding tool SSPACE58 and final scaffold correction was achieved with mate pair (3 kb) discordance analysis using REAPER59. The total assembled bases comprise 748 Mb.
For a single male P. latipinna from north of Tampico (Pla_Tam) we generated 34× sequence coverage of paired 100 bp reads (20×) and 3 kb paired reads (14×) and for P. mexicana (single female from Laguna Champaxan, Pme_Cha) 30 × sequence coverage of paired 100 bp reads (22×) and 3 kb reads (8×). Both species-specific sequence sets were aligned to the PoeFor_5.1.2 reference to generate sequence contigs as previously described23. Additional scaffolding was achieved with the use of SSPACE58. The assemblies comprise 18,161 and 18,275 total scaffolds with N50 contig and scaffold lengths of 33 kb and 280 kb, respectively, for P. latipinna, and 40 kb and 270 kb, respectively, for P. mexicana.
For all P. formosa population samples we generated sequence on the HiSeq2500 instrument (100 bp read lengths) whereas the P. latipinna and P. mexicana isolates were sequenced on Illumina ×10 instruments (125 bp read lengths). All reads were pre-processed by removing duplicate reads with Picard v.1.113 (http://picard.sourceforge.net), and only properly paired reads were aligned with the appropriate reference using BWA-MEM60.
Gene annotation
Automated gene predictions followed previous methods for Ensembl61 and NCBI62 pipelines, including masking of repeats before ab initio gene predictions, for evidence-supported gene model building. However, the P. latipinna and P. mexicana genomes were only annotated for gene content with the NCBI pipeline62. In both annotation processes gene models were supported or novel based on RNA sequencing (RNA-seq) data from P. formosa, P. mexicana and P. latipinna independent tissues (see NCBI annotation report: https://www.ncbi.nlm.nih.gov/genome/annotation_euk/Poecilia_formosa/101/) for the full list of tissues. For the Ensembl P. formosa gene build only the RNA-seq data from liver, brain, skin ovary, gills and embryonic tissues were used. The final Ensembl P. formosa gene set comprises models based on orthologous proteins from the vertebrate division of UniProtKB, longest translations of some stickleback gene models from Ensembl 73, as well as models from RNA-seq data. RNA-seq data were used to further improve gene model accuracy by alignment to nascent gene models to delineate boundaries of untranslated regions as well as to identify genes not found through interspecific similarity evidence from other species. Our measures of gene representation for the aligned core eukaryotic genes (n = 458) using CEGMA63 showed >92% were complete at 90% of their estimated length in all three Poecilia species.
SNP and heterozygosity analysis
Resequenced genomes were aligned against the P. formosa reference to extract SNPs with SAMtools and VCFtools using stringent criteria. Fixed heterozygous sites informing about ancestry were defined by the rule that the heterozygous genotype should be observed in all isolates and in case of missing genotypes at least 5 P. formosa isolates are heterozygotes with the remaining isolates missing genotype calls. Genome-wide expected heterozygosity (theta) was estimated using a maximum-likelihood method64 based on sites with a minimum of 4× sequence coverage after taking into account sequencing error and random sampling of two homologous chromosomes in a diploid organism.
The within-species per-site nucleotide diversity was calculated for sequenced genomes of each species using VCFtools65 and was averaged over all the SNP sites. Tajima’s D values were calculated for each 50 kb non-overlapping genomic window using genome-wide SNP data sets in the software VCFtools, and subsequently a genome-wide average was calculated.
To validate a previous study that claimed backcrossing has occurred we used INTROGRESS as in ref.29 to analyse the introgression of genotypes between divergent, hybridizing lineages, including estimating genomic clines from multi-locus genotype data and testing for deviations from neutral expectations.
DNA sequences described in population sequencing for P. formosa, P. latipinna and P. mexicana from 5 individuals were each aligned with the homologous reference of each at an average input sequence coverage of 11× using BWA-MEM. Total SNPs were called by GATK HaplotypeCaller and GenotypGVCFs66. All SNPs were used to report expected LoFs classified as single-base substitutions that disrupt splice acceptor, splice donor, stop loss or gain codons using the VAAST software67. To filter variants that were shared across all samples in either of the sexual parents we removed LoFs that were shared in all. We reasoned the likelihood of fishes having LoFs fixed among different populations is very rare. However, we did allow LoFs to be shared across all in the P. formosa samples. As they are clonal it is feasible that the LoFs could remain fixed over many generations.
Mitochondrial genome analyses
Reference mitochondrial genomes of all three species were produced from long-PCR products sequencing (Supplementary Note 4). Long PCRs were performed with primers designed from the complete mitochondrial genomes of Hypoatherina tsurugae (NC_004386.1), Gambusia holbrooki (NC_004388.1), Melanoteania lacustris (NC_004385.1) and Colalabis (NC_003183.1). Fragment 1 (10.7 kb): TRPFishFor: 5′_AGACCAAGGGCCTTCAAAGCC_3′, 15995Rev.Fish: 5′_CTTTGGGAGCTAGGGGTGAGAGTT_3′, Fragment 2 (7 kb): L15995: 5′_AACTCTCACCCCTAGCTCCCAAAG_3′, TRPFishRev: 5′_GGCTTTGAAGGCCCTTGGTCT_3′, Fragment 3 (7.3 kb): H16100R: 5′_ATGTAGGGTTACAYTACTTTAAAT_3′, ATP6fPoec-long: 5′_AACTATCWATTAACATAGGTCTTGCWGGCGCT_3′ using Takara Taq under the following conditions: 94 °C 90 s, 94 °C 15 s, 49 to 63 °C 30 s followed by 68 °C 6.45 min for each step, 72 °C 12 min.
PCR products were mechanically hydro-sheered and cloned into shotgun libraries before Sanger sequencing. Sequences were first assembled into contigs using the phred/phrap assembler (www.phrap.org/phredphrapconsed.html) and then contigs further merged using the cap3 assembler (http://seq.cs.iastate.edu/cap3.html). Remaining gaps were closed by additional PCRs. Whenever aligned shotgun sequences indicated potentially conflicting sequence data, fragments were re-amplified and re-sequenced by specific direct PCRs. The mitochondrial genomes from NGS data were assembled with the Geneious program version 8.1.7 (http://www.geneious.com/) (map to reference algorithm, medium/low sensitivity, up to 5 iterations) using the P. reticulata mitochondrial genome (Genbank Accession number AB898687.1) as outgroup reference. Consensus sequences were edited with Bioedit (http://www.mbio.ncsu.edu/BioEdit/bioedit.html) and analyzed using MEGA (genetic distances) (http://www.megasoftware.net/) and PopART (mitochondrial haplotype minimum spanning network) (http://popart.otago.ac.nz).
Repeat analyses
Repeat (TEs and non-TEs) libraries of P. formosa, P. mexicana and P. latipinna were established using an automatic annotation with RepeatModeler1.0 http://www.repeatmasker.org, combined with manual search of known TE proteins and phylogenetic classification. For comparison we used the genomes of platyfish (Xiphophorus maculatus)68 and guppy (P. reticulata) (http://www.ncbi.nlm.nih.gov/assembly/GCA_000633615.2). TE superfamily classification is based on a universal classification69. TE contents in genome assemblies were estimated using RepeatMasker 3.3.0 (http://www.repeatmasker.org). The relative age of TE copies in poeciliid genomes was calculated through Kimura distance analyses (http://www.repeatmasker.org/genomicDatasets/RMGenomicDatasets.html). The proportion of sites with transitions (p) and transversions (q) were transformed as follows: (K = −½ ln(1 – 2p – q) – ¼ ln(1 – 2q)). Transcriptome assembly of the Amazon molly was masked to evaluate the proportion of transcribed TE superfamilies.
For phylogenetic analyses, nucleotide sequences were first translated using Augustus70. TE proteins (reverse transcriptase or transposase) were then aligned using MUSCLE71 in PhyML package72 and phylogenetic trees were reconstructed with the maximum likelihood method using approximate likelihood ratio test (aLRT) non-parametric branch supports.
Gypsy insertions longer than 3 kb (based on RepeatMasker annotation) were manually analysed to identify specific insertions. Insertions were extracted with flanking regions (+/−1 kb) and aligned against P. mexicana and P. latipinna genome assemblies via BLAST. P. formosa insertions were considered specific when they shared no flanking regions with the parental species. A similar method was used to investigate the presence of solo-LTR.
Gene selection
Orthologous genes of P. formosa, P. latipinna, P. mexicana and P. reticulata were identified using Inparanoid http://inparanoid.sbc.su.se/cgi-bin/index.cgi (default settings). For each gene both protein and cDNA sequences were aligned using clustal-omega (v.1.2.1) http://www.clustal.org/omega/ (option: -outfmt fasta). Non-conserved blocks from the alignments were removed by Gblocks (v.0.91b) http://molevol.cmima.csic.es/castresana/Gblocks.html (options: -b4 10 -b5 n –b3 5). The resulting sequence alignments were converted into a codon alignment using pal2nal (v.v14) http://www.bork.embl.de/pal2nal/. Codon alignments were converted into phylip format using clustal-omega (option: -outfmt phy). Trees were built using Phylip (v.3.696, http://evolution.genetics.washington.edu/phylip.html) with P. reticulata as outgroup. For phylogenetic analyses by maximum likelihood the ‘Environment for Tree Exploration’ (ETE) toolkit http://etetoolkit.org/ was used. For detection of positive selection, we calculated six different models for null hypothesis or alternative hypothesis. Comparison 1 included two branch specific models: One model with a fixed ϖ = 1 value (b_neut) and a second with the marked branch being allowed to evolve independently (b_free). Comparison 2 included two branch-site specific models, model bsA1 (neutral) versus model bsA (positive selection) to identify sites under positive selection on a specific branch. Comparison 3 included two site specific models, model M1 (neutral) and M2 (positive selection). Candidate genes for positive selection were required to be significant in all three comparisons. Genes were considered to be under relaxed selection if ϖ was significantly different between foreground and background branches (b_free vs. M0) and ϖ for the foreground branch did not significantly differ from 1 in a comparison between the models b_free and b_neut. For all comparisons a LRT was performed.
Immune genes
Fasta files were searched using the usearch algorithm (https://www.drive5.com/usearch/) with user-specific similarity thresholds of 80% for MHC genes and 90% for all innate immunity genes. Genes were assembled using Geneious 8.1.7 (http://www.geneious.com) and edited using Bioedit. For the MHC analyses only consensus sequences including an open reading frame of at least 200 bp were included and different MHC alleles were defined using amino acid differences. The innate immunity genes (6 different Toll-like receptors, 1 region of IFNPhi4, 2 Interleukins, Interferon regulating factor 2, complement components C2 and C3, CLEC, CD59 and TRAF6) were assembled using the P. formosa reference gene provided by Ensembl. MEGA was used to calculate genetic distances and phylogenetic tree. Neighbour-joining, maximum likelihood, and minimum evolution produced the same topology in tree reconstruction.
Pairwise relative genetic distance was calculated using the program MEGA v.6.06. For each gene mean values for individuals or species were calculated and compared using ANOVA. ANOVA and Scheffé Posthoc test were done with STATISTICA v.13 (Dell Inc.). Selection analysis (dN–dS; N, nonsynonymous; S, synonymous base exchange) were done using MEGA v.6.06.
Allele-specific gene expression analysis
Six P. formosa RNA-Seq sequencing files were downloaded (liver: SRR629501, SRR629518; skin: SRR629511, SRR629503; gills: SRR629508, SRR629510) from SRA and were filtered P. latipinna (GCF_001443285.1) and P. mexicana (GCF_001443325.1) reference RNA sequences were downloaded from NCBI. To assign ancestral alleles of P. formosa genes, sequence homology between ancestral alleles and P. formosa genes were identified using Blastn73. When multiple representation of homology was observed, the ancestral allele that generated the longest sequence alignment was kept to represent one ancestral allele of a P. formosa gene. Of the 25338 coding genes in the P. formosa genome that have a genome feature as ‘mRNA’, 22118 genes can be assigned to both P. latipinna and P. mexicana alleles. Short sequencing files generated from P. formosa RNA-seq reads were mapped to the ancestral allele reference sequences that are generated by combining sequences of both ancestral alleles using Bowtie274 Customized Perl script was used to retrieve and quantify the short reads that only aligned to one of the ancestral alleles73.
Differential expression between parental alleles was tested using edgeR http://bioconductor.org/packages/release/bioc/html/edgeR.html (P. latipinna alleles were used as control). Log2(P. mex / P. lat) was used to label the relativize expression of both ancestral alleles. The false discovery rate (FDR) <0.05 was used to determine whether a gene shows allelic expression bias towards one ancestral allele.
Overall P. formosa gene expression was assessed by mapping the RNA-seq sequence reads to the P. formosa genome (GCF_000485575.1) using tophat2 (http://ccb.jhu.edu/software/tophat/index.shtml), and read counts quantified using featureCounts75. A gene was determined to be expressed if at least one sample of the two biological replicates reached a library size-normalized read count (that is, count per million reads) of 0.5.
Detection of segmental duplications, copy number variations
Regions of genomic duplications were estimated across the genomes of 19 P. formosa individuals, 5 P. latipinna individuals and 4 P. mexicana individuals, following an approach based on differences in depth of coverage. We used the RepeatMasker (www.repeatmasker.org) output from NCBI for the Poecilia_formosa-5.1.2 assembly and generated output from Tandem Repeat Finder 4.07b, using default parameters. Genomic regions identified by either approach were hard-masked to remove most of the repetitive sequence present in the assembly (Supplementary Table 30). We further sought to identify and mask potential hidden repeats. Scaffolds and contigs were partitioned into 36 bps kmers, with adjacent kmers overlapping by 5 bps. These kmers were mapped to the repeat masked version of the assembly using mrsFast 3.3.0, to account for multi-mappings. Over-represented kmers defined as those with more than 42 mappings (accounting for a cumulative proportion of 90% of the mappings) in the assembly (Supplementary Fig. 8) were additionally hard-masked.
We evaluated the overall sequencing performance on the raw reads and demarcated the regions of the reads that displayed the best qualities. We initially mapped the reads to an unmasked version of the assembly using BWA60 and removed PCR duplicates using PicardTools (http://picard.sourceforge.net/). Non-duplicated reads were then clipped into two consecutive fragments of 36 bps. The resulting reads were then mapped to the prepared kmer masked version of the assembly using mrFast 2.5.0.0. mrCaNaVaR 0.51 was applied to infer the copy number in 1 kb non-overlapping windows of unmasked sequence, that is, the real window size may exceed 1 kb because it includes any repeat or gap. Notably, as reads will not map to the genomic coordinates masked in the assembly, a spurious drop off in read depth estimates might appear at the edges of masked regions, which could underestimate the copy number inferences in later steps. To prevent this, the 36 bps flanking any masked region or gap were also masked and thus not included in the window definition. Genome-wide read depth distribution was calculated by iteratively excluding windows with extreme read depth values relative to the normal distribution and the remaining windows defined as control regions. The mean read depth in these control regions was considered to correspond to a copy number equal to two and used to convert the read depth value in each window into a GC-corrected absolute copy number (see Supplementary Figs. 9 and 10 for distribution in copy number values in control regions and per sample duplication content for all P. formosa samples, Supplementary Figs.11 and 12 for the parental P. latipinna and P. mexicana samples).
We defined segmental duplications (SDs) as at least five consecutive windows of non-overlapping non-masked sequences with copy number values higher than the mean plus three standard deviations, allowing one of the windows to have a copy number value that is larger than the mean plus two standard deviations. Cutoffs were defined on a per sample basis. Furthermore, regions with an absolute copy number above 100 in any sample were excluded. Gaps were removed from the called intervals in downstream analysis.
GO enrichment analyses on the set of duplicated genes were performed using R and the topGO package on a per species basis. First, the Ensembl IDs of the duplicates genes were mapped to the corresponding GO terms using the biomaRt package. GO enrichment was studied for the GO ontologies ‘Biological Process’ and ‘Molecular Function’. We used Fishers exact test to detect enrichments, and corrected the raw P values with the Benjamini–Hochberg method.
Paternal introgression
To determine whether paternal introgression events had occurred in any of the analysed samples, we assessed the existence of allele imbalance at heterozygous sites across all scaffolds larger than 100 Kb. Sequencing reads were aligned to the P. formosa reference assembly using BWA60 v.0.7.4 with default parameters and PCR duplicates were removed using PicardTools (http://picard.sourceforge.net/). SNPs were called using Freebayes (v.0.9.14) with the following parameters: standard-filters, no-population-priors, report-genotype-likelihood-max. We retained for subsequent analysis mapped, biallelic heterozygous SNPs with a minimum QUAL > = 30 and a minimum DP > = 5. We also required at least 20% of the reads supporting each SNP derived from the minor allele. In the absence of introgression allele frequencies should be approximately 50% of the sites. In the case of paternal introgression a distorted frequency is expected skewed towards alleles that are present in the paternal species. We identified putative paternal introgression events as bimodal distributions of the reference allele frequency that result from the presence of an extra copy inherited from the male host. We confirmed that the inferred copy number throughout the identified scaffolds was higher in the samples where the paternal introgression was detected. To trace back the imbalanced allele to the corresponding paternal species, sequencing data derived from five P. mexicana and five P. latipinna fishes was used to call SNPs using the approach described above. After filtering, fixed homozygous sites with opposing genotypes in the two species were identified by the following criteria: either all P. mexicana samples were homozygous for the reference allele and all P. latipinna samples were homozygous for the alternative allele, or the other way around. These fixed homozygous sites were then intersected with heterozygous variants called in each of the 19 P. formosa samples and for each site the percentage of reads carrying the allele present in P. mexicana was calculated. In the absence of paternal introgression, this frequency should be ~50%. If there is an introgressed scaffold, the imbalanced allele at heterozygous sites should match the allele present in the corresponding paternal species, and thus this frequency would deviate from 50%.
To model the relief from genomic decay allowed by paternal introgression the equations described in a previous study12 were used with parameter estimates changed as stated.
Life Science Reporting Summary
Further information on experimental design is available in the Life Sciences Reporting Summary.
Data availability
All assemblies are available at Genbank under the following accession numbers: GCF_000485575.1 (Poecilia_formosa-5.1.2) BioProject PRJNA89109, GCA_001443325.1 (Poecilia_mexicana-1.0) BioProject PRJNA196869, and GCA_001443285.1 (Poecilia_latipinna-1.0), BioProject PRJNA196862. Accession numbers for population sample genome reads are given in Supplementary Table 3.
References
Charlesworth, B. & Charlesworth, D. Rapid fixation of deleterious alleles can be caused by Muller’s ratchet. Genet. Res. 70, 63–73 (1997).
Muller, H. J. The relation of recombination to mutational advance. Mutat. Res. Fund. Mol. M. 1, 2–9 (1964).
Lynch, M., Conery, J. & Burger, R. Mutational meltdowns in sexual populations. Evolution 49, 1067–1080 (1995).
Lynch, M. & Gabriel, W. Mutation load and the survival of small populations. Evolution 44, 1725–1737 (1990).
Bell, G. The Masterpiece of Nature: The Evolution and Genetics of Sexuality (Univ. California Press, Berkeley, 1982).
Van Valen, L. A new evolutionary law. Evolut. Theory 1, 1–30 (1973).
McDonald, M. J., Rice, D. P. & Desai, M. M. Sex speeds adaptation by altering the dynamics of molecular evolution. Nature 531, 233–236 (2016).
Maynard Smith, J. The Evolution of Sex (Cambridge Univ. Press, London, 1978).
Lively, C. M. & Morran, L. T. The ecology of sexual reproduction. J. Evol. Biol. 27, 1292–1303 (2014).
Tucker, A. E., Ackerman, M. S., Eads, B. D., Xu, S. & Lynch, M. Population-genomic insights into the evolutionary origin and fate of obligately asexual Daphnia pulex. Proc. Natl. Acad. Sci. USA 110, 15740–15745 (2013).
Lampert, K. P. & Schartl, M. The origin and evolution of a unisexual hybrid: Poecilia formosa. Phil. Trans. R. Soc. B 363, 2901–2909 (2008).
Loewe, L. & Lamatsch, D. K. Quantifying the threat of extinction from Muller’s ratchet in the diploid Amazon molly (Poecilia formosa). BMC Evol. Biol. 8, 88 (2008).
Quattro, J. M., Avise, J. C. & Vrijenhoek, R. C. An ancient clonal lineage in the fish genus Poeciliopsis (Atheriniformes: Poeciliidae). Proc. Natl Acad. Sci. USA 89, 348–352 (1992).
Stöck, M., Lampert, K. P., Möller, D., Schlupp, I. & Schartl, M. Monophyletic origin of multiple clonal lineages in an asexual fish (Poecilia formosa). Mol. Ecol. 19, 5204–5215 (2010).
Speijer, D., Lukes, J. & Elias, M. Sex is a ubiquitous, ancient, and inherent attribute of eukaryotic life. Proc. Natl. Acad. Sci. USA 112, 8827–8834 (2015).
Schurko, A. M., Neiman, M. & Logsdon, J. M.Jr. Signs of sex: what we know and how we know it. Trends Ecol. Evol. 24, 208–217 (2009).
Flot, J. F. et al. Genomic evidence for ameiotic evolution in the bdelloid rotifer Adineta vaga. Nature 500, 453–457 (2013).
Xu, S. et al. Hybridization and the origin of contagious asexuality in Daphnia pulex. Mol. Biol. Evol. 32, 3215–3225 (2015).
Hubbs, C. L. & Hubbs, L. C. Apparent parthenogenesis in nature, in a form of fish of hybrid origin. Science 76, 628–630 (1932).
Avise, J. C. Evolutionary perspectives on clonal reproduction in vertebrate animals. Proc. Natl. Acad. Sci. USA 112, 8867–8873 (2015).
Schartl, M., Wilde, B., Schlupp, I. & Parzefall, J. Evolutionary origin of a parthenoform, the Amazon molly Poecilia formosa, on the basis of a molecular genealogy. Evolution 49, 827–835 (1995).
Schlupp, I. The evolutionary ecology of gynogenesis. Annu. Rev. Ecol. Evol. Syst. 36, 399–417 (2005).
Shen, Y. et al. X. couchianus and X. hellerii genome models provide genomic variation insight among Xiphophorus species. BMC Genom. 17, 37 (2016).
Hickey, D. A. Selfish DNA: a sexually-transmitted nuclear parasite. Genetics 101, 519–531 (1982).
Arkhipova, I. & Meselson, M. Deleterious transposable elements and the extinction of asexuals. Bioessays 27, 76–85 (2005).
Bast, J. et al. No accumulation of transposable elements in asexual arthropods. Mol. Biol. Evol. 33, 697–706 (2016).
Hirase, S., Ozaki, H. & Iwasaki, W. Parallel selection on gene copy number variations through evolution of three-spined stickleback genomes. BMC Genom. 15, 735 (2014).
Miller, M. P., Unal, E., Brar, G. A. SpringerAmpamp; Amon, A. Meiosis I chromosome segregation is established through regulation of microtubule-kinetochore interactions. Elife 1, e00117 (2012).
Alberici da Barbiano, L., Gompert, Z., Aspbury, A. S., Gabor, C. R. & Nice, C. C. Population genomics reveals a possible history of backcrossing and recombination in the gynogenetic fish Poecilia formosa. Proc. Natl. Acad. Sci. USA 110, 13797–13802 (2013).
Xu, S., Omilian, A. R. & Cristescu, M. E. High rate of large-scale hemizygous deletions in asexually propagating Daphnia: implications for the evolution of sex. Mol. Biol. Evol. 28, 335–342 (2011).
Miller, D. E. et al. A whole-chromosome analysis of meiotic recombination in Drosophila melanogaster. G3 2, 249–260 (2012).
Williams, A. L. et al. Non-crossover gene conversions show strong GC bias and unexpected clustering in humans. Elife 4, e04637 (2015).
Nanda, I. et al. Stable inheritance of host species-derived microchromosomes in the gynogenetic fish Poecilia formosa. Genetics 177, 917–926 (2007).
Schartl, M. et al. Incorporation of subgenomic amounts of DNA as compensation for mutational load in a gynogenetic fish. Nature 373, 68–71 (1995).
Vrijenhoek, R. C. Unisexual fish: model systems for studying ecology and evolution. Annu. Rev. Ecol. 25, 71–96 (1994).
Litman, G. W., Rast, J. P. & Fugmann, S. D. The origins of vertebrate adaptive immunity. Nat. Rev. Immunol. 10, 543–553 (2010).
Apanius, V., Penn, D., Slev, P. R., Ruff, L. R. & Potts, W. K. The nature of selection on the major histocompatibility complex. Crit. Rev. Immunol. 17, 179–224 (1997).
Lampert, K. P., Fischer, P. & Schartl, M. Major histocompatilibility complex variability in the clonal Amazon molly, Poecilia formosa: is copy number less important than genotype? Mol. Ecol. 18, 1124–1136 (2009).
Schaschl, H., Tobler, M., Plath, M., Penn, D. J. & Schlupp, I. Polymorphic MHC loci in an asexual fish, the amazon molly (Poecilia formosa: Poeciliidae). Mol. Ecol. 17, 5220–5230 (2008).
Fumey, J., Hinaux, H., Noirot, C., Rétaux, S. & Casane, D. Evidence of Late Pleistocene origin of Astyanax mexicanus cavefish. Preprint at https://www.biorxiv.org/content/early/2017/10/27/094748 (2016).
Debortoli, N. et al. Genetic exchange among bdelloid rotifers is more likely due to horizontal gene transfer than to meiotic sex. Curr. Biol. 26, 723–732 (2016).
Bogart, J. P., Bartoszek, J., Noble, D. W. & Bi, K. Sex in unisexual salamanders: discovery of a new sperm donor with ancient affinities. Heredity 103, 483–493 (2009).
Mandegar, M. A. & Otto, S. P. Mitotic recombination counteracts the benefits of genetic segregation. Proc. R. Soc. B 274, 1301–1307 (2007).
Hotz, H., Semlitsch, R. D., Gutmann, E., Guex, G. D. & Beerli, P. Spontaneous heterosis in larval life-history traits of hemiclonal frog hybrids. Proc. Natl. Acad. Sci. USA 96, 2171–2176 (1999).
Vrijenhoek, R. C. in Population Biology and Evolution (eds Wöhrmann, K. & Loeschke, V.) 217–231 (Springer, Heidelberg, 1984).
Howard, R. S. & Lively, C. M. Parasitism, mutation accumulation and the maintenance of sex. Nature 367, 554–557 (1994).
Tobler, M. & Schlupp, I. Parasites in sexual and asexual mollies (Poecilia, Poeciliidae, Teleostei): a case for the Red Queen? Biol. Lett. 1, 166–168 (2005).
Vrijenhoek, R. C. & Parker, E. D. in Lost Sex (eds Schön, I. et al.) 99–131 (Springer, Dordrecht, 2009).
Mallet, J. Hybridization as an invasion of the genome. Trends Ecol. Evol. 20, 229–237 (2005).
Lampert, K. P. et al. Automictic reproduction in interspecific hybrids of poeciliid fish. Curr. Biol. 17, 1948–1953 (2007).
Abbott, R. et al. Hybridization and speciation. J. Evol. Biol. 26, 229–246 (2013).
Maheshwari, S. & Barbash, D. A. The genetics of hybrid incompatibilities. Annu. Rev. Genet. 45, 331–355 (2011).
Vidovic, D. & Matzinger, P. Unresponsiveness to a foreign antigen can be caused by self-tolerance. Nature 336, 222–225 (1988).
Wegner, K. M., Kalbe, M., Kurtz, J., Reusch, T. B. H. & Milinski, M. Parasite selection for immunogenetic optimality. Science 301, 1343 (2003).
Poletaev, A. B., Churilov, L. P., Stroev, Y. I. & Agapov, M. M. Immunophysiologiy versus immunopahtology: natural autoimmunity in human health and disease. Pathophysiology 19, 221–231 (2012).
Tobler, M., Wahli, T. & Schlupp, I. Comparison of parasite communities in native and introduced populations of sexual and asexual mollies of the genus Poecilia. J. Fish. Biol. 67, 1072–1082 (2005).
Gnerre, S. et al. High-quality draft assemblies of mammalian genomes from massively parallel sequence data. Proc. Natl. Acad. Sci. USA 108, 1513–1518 (2011).
Boetzer, M. & Pirovano, W. SSPACE-LongRead: scaffolding bacterial draft genomes using long read sequence information. BMC Bioinform. 15, 211 (2014).
Hunt, M. et al. REAPR: a universal tool for genome assembly evaluation. Genome Biol. 14, R47 (2013).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Flicek, P. et al. Ensembl 2014. Nucleic Acids Res. 42, D749–D755 (2014).
O’Leary, N. A. et al. Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 44, D733–D745 (2016).
Parra, G., Bradnam, K. & Korf, I. CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23, 1061–1067 (2007).
Haubold, B., Pfaffelhuber, P. & Lynch, M. mlRho - a program for estimating the population mutation and recombination rates from shotgun-sequenced diploid genomes. Mol. Ecol. 19, 277–284 (2010).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Kennedy, B. et al. Using VAAST to identify disease-associated variants in next-generation sequencing data. Curr. Protoc. Hum. Genet. 81, 11–25 (2014).
Schartl, M. et al. The genome of the platyfish, Xiphophorus maculatus, provides insights into evolutionary adaptation and several complex traits. Nat. Genet. 45, 567–572 (2013).
Bao, W., Kojima, K. K. & Kohany, O. Repbase Update, a database of repetitive elements in eukaryotic genomes. Mob. DNA 6, 11 (2015).
Stanke, M., Steinkamp, R., Waack, S. & Morgenstern, B. AUGUSTUS: a web server for gene finding in eukaryotes. Nucleic Acids Res. 32, W309–W312 (2004).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010).
Lu, Y. et al. Molecular genetic response of Xiphophorus maculatus-X. couchianus interspecies hybrid skin to UVB exposure. Comp. Biochem. Physiol. C 178, 86–92 (2015).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Acknowledgements
We thank E. G. Ávila for assistance and help in the field, I. Schlupp for P. formosa samples from Texas and discussions, M. Niklaus-Ruiz for help in preparation of the manuscript, and R. Agrawala for consultation on the assisted assembly aspects of this project. This work was supported by grants to W.C.W. (NIH: 2R24OD011198-04A1), M.W.H. (NSF DBI-1564611), M.S. (German Research Foundation DFG projects Scha408/10-1 and Scha408/12-1), M.St. (Heisenberg-Fellowship STO 493/2-2 of the German Science Foundation/DFG), T.M.B. (MINECO BFU2014-55090-P (FEDER), U01 MH106874 grant, Howard Hughes International Early Career, Obra Social ‘La Caixa’ and Secretaria d’Universitats i Recerca del Departament d’Economia i Coneixement de la Generalitat de Catalunya) and R.B.W. (NIH: R24OD011120). The genome annotation work carried out by NCBI was supported by the Intramural Research Program of the NIH, National Library of Medicine. The genome annotation work by Ensembl was supported by funding from the Wellcome Trust (WT108749/Z/15/Z and WT098051), the National Institutes of Health (R24 RR032658-01) and the European Molecular Biology Laboratory.
Author information
Authors and Affiliations
Contributions
W.C.W. and M.S. initiated and managed the genome project. P.M., C.T., and L.H. built the assembly, B.A., D.N.M. generated the Ensembl gene annotation, K.P. led the NCBI gene annotation, F.F., M.M. for annotation of gene variants, G.W.C.T and M.W.H. did the gene family analysis, D.C. and J.N.V. performed the repeat and TE analyses, M.St., K.L., and J.W. performed the mtDNA analyses, K.L. performed the immune gene analyses, S.K., Z.W., and M.S. performed the analysis of male specific, TE silencing machinery and meiosis genes, S.X., M.L. performed and reviewed the analysis of heterozygosity and gene conversion, C.M.G. participated in the population analyses, collection and sample identification, R.G.P., L.F.K. and T.M.B. did the introgression and segmental duplication analysis, L.L. did the modeling of genomic decay, Y.L. and R.B.W. performed the ASE analyses, S.K. and M.S. performed the selection analysis, all authors contributed to data interpretation, W.C.W. and M.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Notes 1–7, Supplementary Figures 1–17, Supplementary Tables 1–6, 10–12, 16–24, 26–34.
Supplementary Table 7
Positively selected genes in P. formosa.
Supplementary Table 8
Duplicated genes in P. formosa.
Supplementary Table 9
Gene family expansion and retraction in P. formosa.
Supplementary Table 13
Poecilia variants.
Supplementary Table 14
Genes under relaxed selection in P. formosa, mexicana and latipinna.
Supplementary Table 15
Genes positive selection in P. formosa, mexicana, latipinna.
Supplementary Table 25
GO-term enrichment of biological processes in introgressed genes.
Supplementary Table 35
Duplicated genes in P. formosa.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Warren, W.C., García-Pérez, R., Xu, S. et al. Clonal polymorphism and high heterozygosity in the celibate genome of the Amazon molly. Nat Ecol Evol 2, 669–679 (2018). https://doi.org/10.1038/s41559-018-0473-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-018-0473-y
This article is cited by
-
Genetic and phenotypic diversification in a widespread fish, the Sailfin Molly (Poecilia latipinna)
BMC Ecology and Evolution (2024)
-
Predator exposure early in life shapes behavioral development and individual variation in a clonal fish
Scientific Reports (2024)
-
Evolutionary mechanisms and practical significance of reproductive success and clonal diversity in unisexual vertebrate polyploids
Science China Life Sciences (2024)
-
Reproductive individuality of clonal fish raised in near-identical environments and its link to early-life behavioral individuality
Nature Communications (2023)
-
Population genetics of clonally transmissible cancers
Nature Ecology & Evolution (2022)