Abstract
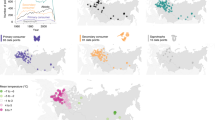
Climate change affects not just where species are found, but also when species’ key life-history events occur—their phenology. Measuring such changes in timing is often hampered by a reliance on biased survey data: surveys identify that an event has taken place (for example, the flower is in bloom), but not when that event happened (for example, the flower bloomed yesterday). Here, we show that this problem can be circumvented using statistical estimators, which can provide accurate and unbiased estimates from sparsely sampled observations. We demonstrate that such methods can resolve an ongoing debate about the relative timings of the onset and cessation of flowering, and allow us to place modern observations reliably within the context of the vast wealth of historical data that reside in herbaria, museum collections, and written records. We also analyse large-scale citizen science data from the United States National Phenology Network and reveal not just earlier but also potentially more variable flowering in recent years. Evidence for greater variability through time is important because increases in variation are characteristic of systems approaching a state change.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
09 February 2019
In the version of this Article originally published, the rate of change plotted in Figure 2 was incorrect because of a coding error. The corrected figure is shown below. In the original Figure 2 legend, the onset of flowering slope was given as ‘0.99, 95% CI: 0.90–1.08’, the cessation of flowering slope was given as ‘1.02, 95% CI: 0.91–1.13’, and the r2adjusted for each model was given as greater than 74%’. The correct values are ‘1.04, 95% CI: 0.97–1.12’, ‘1.25, 95% CI: 0.87–1.62’, and ‘60%’, respectively. The main text and the conclusion that the slopes of these relationships are statistically indistinguishable from 1.00 are unchanged. These errors have now been corrected in the PDF and HTML versions of the article. The authors are grateful to A. Iler, who drew attention to this issue.
References
IPCC Climate Change 2014: Synthesis Report (eds Core Writing Team, Pachauri, R. K. & Meyer L. A.) (Cambridge Univ. Press, Cambridge, 2015).
Menzel, A. et al. European phenological response to climate change matches the warming pattern. Glob. Change Biol. 12, 1969–1976 (2006).
Parmesan, C. et al. Ecological and evolutionary responses to recent climate change. Annu. Rev. Ecol. Evol. Syst. 37, 637–669 (2006).
Wolkovich, E. M. et al. Warming experiments underpredict plant phenological responses to climate change. Nature 485, 494–497 (2012).
Silvertown, J. A new dawn for citizen science. Trends Ecol. Evol. 24, 467–471 (2009).
Pyke, G. H. & Ehrlich, P. R. Biological collections and ecological/environmental research: a review, some observations and a look to the future. Biol. Rev. 85, 247–266 (2010).
Robbirt, K. M., Davy, A. J., Hutchings, M. J. & Roberts, D. L. Validation of biological collections as a source of phenological data for use in climate change studies: a case study with the orchid ophrys sphegodes. J. Ecol. 99, 235–241 (2011).
Baird, R. C. Leveraging the fullest potential of scientific collections through digitisation. Biodivers. Informatics 7, 130–136 (2010).
Beaman, R. S. & Cellinese, N. Mass digitization of scientific collections: new opportunities to transform the use of biological specimens and underwrite biodiversity science. ZooKeys 209, 7–17 (2012).
Balke, M. et al. Biodiversity into your hands: a call for a virtual global natural history ‘metacollection’. Front. Zool. 10, 55 (2013).
Willis, C. G. et al. Crowdcurio: an online crowdsourcing platform to facilitate climate change studies using herbarium specimens. New Phytol. 215, 479–488 (2017).
Meyer, C., Weigelt, P. & Kreft, H. Multidimensional biases, gaps and uncertainties in global plant occurrence information. Ecol. Lett. 19, 992–1006 (2016).
Roberts, D. L. & Solow, A. R. Flightless birds: when did the dodo become extinct? Nature 426, 245–245 (2003).
Ruggles, R. & Brodie, H. An empirical approach to economic intelligence in World War II. J. Am. Stat. Assoc. 42, 72–91 (1947).
Cooke, P. Optimal linear estimation of bounds of random variables. Biometrika 67, 257–258 (1980).
Weissman, I. Confidence intervals for the threshold parameter. Commun. Stat. Theory Methods 10, 549–557 (1981).
Inouye, D. W. Effects of climate change on phenology, frost damage, and floral abundance of montane wildflowers. Ecology 89, 353–362 (2008).
Aldridge, G., Inouye, D. W., Forrest, J. R., Barr, W. A. & Miller-Rushing, A. J. Emergence of a mid-season period of low floral resources in a montane meadow ecosystem associated with climate change. J. Ecol. 99, 905–913 (2011).
CaraDonna, P. J., Iler, A. M. & Inouye, D. W. Shifts in flowering phenology reshape a subalpine plant community. Proc. Natl Acad. Sci. USA 111, 4916–4921 (2014).
Visser, M. E. & Both, C. Shifts in phenology due to global climate change: the need for a yardstick. Proc. R. Soc. B 272, 2561–2569 (2005).
Miller-Rushing, A. J., Inouye, D. W. & Primack, R. B. How well do first flowering dates measure plant responses to climate change? The effects of population size and sampling frequency. J. Ecol. 96, 1289–1296 (2008).
Davis, C. C., Willis, C. G., Connolly, B., Kelly, C. & Ellison, A. M. Herbarium records are reliable sources of phenological change driven by climate and provide novel insights into species phenological cueing mechanisms. Am. J. Bot. 102, 1599–1609 (2015).
Papworth, S., Rist, J., Coad, L. & Milner-Gulland, E. Evidence for shifting baseline syndrome in conservation. Conserv. Lett. 2, 93–100 (2009).
Cook, B. I. et al. Sensitivity of spring phenology to warming across temporal and spatial climate gradients in two independent databases. Ecosystems 15, 1283–1294 (2012).
Gelman, A., Carlin, J. B., Stern, H. S. & Rubin, D. B. Bayesian Data Analysis 3rd edn (CRC Press, Boca Raton, 2014).
Scheffer, M., Carpenter, S., Foley, J. A., Folke, C. & Walker, B. Catastrophic shifts in ecosystems. Nature 413, 591–596 (2001).
Scheffer, M. et al. Early-warning signals for critical transitions. Nature 461, 53–59 (2009).
Körner, C. & Basler, D. Phenology under global warming. Science 327, 1461–1462 (2010).
Cook, B. I., Wolkovich, E. M. & Parmesan, C. Divergent responses to spring and winter warming drive community level flowering trends. Proc. Natl Acad. Sci. USA 109, 9000–9005 (2012).
Brooks, S. J., Self, A., Toloni, F. & Sparks, T. Natural history museum collections provide information on phenological change in british butterflies since the late-nineteenth century. Int. J. Biometeorol. 58, 1749–1758 (2014).
R Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, 2016); https://www.R-project.org/
Clements, C. F. et al. Experimentally testing the accuracy of an extinction estimator: Solow’s optimal linear estimation model. J. Anim. Ecol. 82, 345–354 (2013).
Therneau, T. deming: Deming, Thiel-Sen and Passing-Bablock Regression. R package version 1.0-1 (R Foundation for Statistical Computing, Vienna, 2014); https://CRAN.R-project.org/package=deming.
Miller-Rushing, A. J. & Primack, R. B. Global warming and flowering times in Thoreau’s Concord: a community perspective. Ecology 89, 332–341 (2008).
Willis, C. G., Ruhfel, B., Primack, R. B., Miller-Rushing, A. J. & Davis, C. C. Phylogenetic patterns of species loss in Thoreau’s woods are driven by climate change. Proc. Natl Acad. Sci. USA 105, 17029–17033 (2008).
Harris, I., Jones, P., Osborn, T. & Lister, D. Updated high-resolution grids of monthly climatic observations–the CRU TS3. 10 dataset. Int. J. Climatol. 34, 623–642 (2014).
Carpenter, B. et al. Stan: a probabilistic programming language. J. Stat. Softw. 76, 1–32 (2017).
Ellwood, E. R., Temple, S. A., Primack, R. B., Bradley, N. L. & Davis, C. C. Record-breaking early flowering in the eastern United States. PLoS ONE 8, e53788 (2013).
Gelman, A. & Pardoe, I. Bayesian measures of explained variance and pooling in multilevel (hierarchical) models. Technometrics 48, 241–251 (2006).
Acknowledgements
We thank the volunteers of the NPN for data collection. W.D.P. and T.J.D. were funded by Fonds de Recherche Nature et Technologies grant number 168004. The Colorado data collection was supported by National Science Foundation grants DEB 75-15422, DEB 78-07784, BSR 81-08387, DEB 94-08382, IBN 98-14509, DEB 02-38331 and DEB 09-22080 to D.W.I. We are grateful to D. Roberts and A. Solow for helping to calculate the s.e. of estimates of timing, and E. L. Wolkovich and B. G. Waring for comments on the paper. We also thank the photographers whose images we used in Fig. 3.
Author information
Authors and Affiliations
Contributions
W.D.P., T.J.D. and C.C.D. conceived of the study. W.D.P. analysed the data. All authors wrote the manuscript and interpreted the results.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–4, Supplementary Tables 1–8, Supplementary Reference
R code
R code to calculate limits. This file was missing when the Article was fisrt published.
Rights and permissions
About this article
Cite this article
Pearse, W.D., Davis, C.C., Inouye, D.W. et al. A statistical estimator for determining the limits of contemporary and historic phenology. Nat Ecol Evol 1, 1876–1882 (2017). https://doi.org/10.1038/s41559-017-0350-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-017-0350-0
This article is cited by
-
A Bayesian Treatment of the German Tank Problem
The Mathematical Intelligencer (2024)
-
Ten best practices for effective phenological research
International Journal of Biometeorology (2023)
-
Arctic warming-induced cold damage to East Asian terrestrial ecosystems
Communications Earth & Environment (2022)
-
Phenological advance in the South African Namaqualand Daisy First and Peak Bloom: 1935–2018
International Journal of Biometeorology (2022)
-
A comparison of herbarium and citizen science phenology datasets for detecting response of flowering time to climate change in Denmark
International Journal of Biometeorology (2022)