Abstract
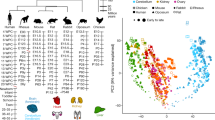
Despite morphological diversification of chordates over 550 million years of evolution, their shared basic anatomical pattern (or ‘bodyplan’) remains conserved by unknown mechanisms. The developmental hourglass model attributes this to phylum-wide conserved, constrained organogenesis stages that pattern the bodyplan (the phylotype hypothesis); however, there has been no quantitative testing of this idea with a phylum-wide comparison of species. Here, based on data from early-to-late embryonic transcriptomes collected from eight chordates, we suggest that the phylotype hypothesis would be better applied to vertebrates than chordates. Furthermore, we found that vertebrates’ conserved mid-embryonic developmental programmes are intensively recruited to other developmental processes, and the degree of the recruitment positively correlates with their evolutionary conservation and essentiality for normal development. Thus, we propose that the intensively recruited genetic system during vertebrates’ organogenesis period imposed constraints on its diversification through pleiotropic constraints, which ultimately led to the common anatomical pattern observed in vertebrates.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Erwin, D. H. et al. The Cambrian conundrum: early divergence and later ecological success in the early history of animals. Science 334, 1091–1097 (2011).
Wallace, A. The Origin of Animal Body Plans: A Study in Evolutionary Developmental Biology (Cambridge Univ. Press, Cambridge, 2000).
Galis, F. & Metz, J. A. Testing the vulnerability of the phylotypic stage: on modularity and evolutionary conservation. J. Exp. Zool. 291, 195–204 (2001).
Irie, N. & Kuratani, S. The developmental hourglass model: a predictor of the basic body plan? Development 141, 4649–4655 (2014).
Piasecka, B., Lichocki, P., Moretti, S., Bergmann, S. & Robinson-Rechavi, M. The hourglass and the early conservation models—co-existing patterns of developmental constraints in vertebrates. PLoS Genet. 9, e1003476 (2013).
Kalinka, A. T. & Tomancak, P. The evolution of early animal embryos: conservation or divergence? Trends Ecol. Evol. 27, 385–393 (2012).
Duboule, D. Temporal colinearity and the phylotypic progression: a basis for the stability of a vertebrate Bauplan and the evolution of morphologies through heterochrony. Dev. Suppl. 135–142 (1994).
Sander, K. Specification of the basic body pattern in insect embryogenesis. Adv. Insect Physiol. 12, 125–238 (1976).
Raff, A. The Shape of Life: Genes, Development, and the Evolution of Animal Form (Univ. of Chicago Press, Chicago, 1996).
Richardson, M. K., Minelli, A., Coates, M. & Hanken, J. Phylotypic stage theory. Trends Ecol. Evol. 13, 158 (1998).
Hazkani-Covo, E., Wool, D. & Graur, D. In search of the vertebrate phylotypic stage: a molecular examination of the developmental hourglass model and von Baer’s third law. J. Exp. Zool. B Mol. 304, 150–158 (2005).
Irie, N. & Sehara-Fujisawa, A. The vertebrate phylotypic stage and an early bilaterian-related stage in mouse embryogenesis defined by genomic information. BMC Biol. 5, 1 (2007).
Domazet-Loso, T. & Tautz, D. A phylogenetically based transcriptome age index mirrors ontogenetic divergence patterns. Nature 468, 815–818 (2010).
Kalinka, A. T. et al. Gene expression divergence recapitulates the developmental hourglass model. Nature 468, 811–814 (2010).
Irie, N. & Kuratani, S. Comparative transcriptome analysis reveals vertebrate phylotypic period during organogenesis. Nat. Commun. 2, 248 (2011).
Quint, M. et al. A transcriptomic hourglass in plant embryogenesis. Nature 490, 98–101 (2012).
Levin, M., Hashimshony, T., Wagner, F. & Yanai, I. Developmental milestones punctuate gene expression in the Caenorhabditis embryo. Dev. Cell 22, 1101–1108 (2012).
Wang, Z. et al. The draft genomes of soft-shell turtle and green sea turtle yield insights into the development and evolution of the turtle-specific body plan. Nat. Genet. 45, 701–706 (2013).
Cheng, X., Hui, J. H., Lee, Y. Y., Wan Law, P. T. & Kwan, H. S. A “developmental hourglass” in fungi. Mol. Biol. Evol. 32, 1556–1566 (2015).
Xu, F. et al. High expression of new genes in trochophore enlightening the ontogeny and evolution of trochozoans. Sci. Rep. 6, 34664 (2016).
Levin, M. et al. The mid-developmental transition and the evolution of animal body plans. Nature 531, 637–641 (2016).
Irie, N. Remaining questions related to the hourglass model in vertebrate evolution. Curr. Opin. Genet. Dev. 45, 103–107 (2017).
Chiba, S., Sasaki, A., Nakayama, A., Takamura, K. & Satoh, N. Development of Ciona intestinalis juveniles (through 2nd ascidian stage). Zool. Sci. 21, 285–298 (2004).
Hotta, K. et al. A web-based interactive developmental table for the ascidian Ciona intestinalis, including 3D real-image embryo reconstructions: I. From fertilized egg to hatching larva. Dev. Dynam. 236, 1790–1805 (2007).
Holland, L. Z. Genomics, evolution and development of amphioxus and tunicates: the Goldilocks principle. J. Exp. Zool. B 324, 342–352 (2015).
Janvier, P. Facts and fancies about early fossil chordates and vertebrates. Nature 520, 483–489 (2015).
Albalat, R. & Canestro, C. Evolution by gene loss. Nat. Rev. Genet. 17, 379–391 (2016).
Yanai, I., Peshkin, L., Jorgensen, P. & Kirschner, M. W. Mapping gene expression in two Xenopus species: evolutionary constraints and developmental flexibility. Dev. Cell 20, 483–496 (2011).
Benton, J. M. Vertebrate Palaeontology. 4th edn (Wiley-Blackwell, Hoboken, NJ, 2014).
Roux, J., Rosikiewicz, M. & Robinson-Rechavi, M. What to compare and how: comparative transcriptomics for Evo-Devo. J. Exp. Zool. B 324, 372–382 (2015).
Jeffery, J. E., Bininda-Emonds, O. R., Coates, M. I. & Richardson, M. K. Analyzing evolutionary patterns in amniote embryonic development. Evol. Dev. 4, 292–302 (2002).
Galis, F. Why do almost all mammals have seven cervical vertebrae? Developmental constraints, Hox genes, and cancer. J. Exp. Zool. 285, 19–26 (1999).
Duboule, D. & Wilkins, A. S. The evolution of ‘bricolage’. Trends Genet. 14, 54–59 (1998).
Pavlicev, M. & Cheverud, J. M. Constraints evolve: context dependency of gene effects allows evolution of pleiotropy. Annu. Rev. Ecol. Evol. Syst. 46, 413–434 (2015).
Wagner, G. P. & Zhang, J. The pleiotropic structure of the genotype-phenotype map: the evolvability of complex organisms. Nat. Rev. Genet. 12, 204–213 (2011).
Riedl, R. Order in Living Organisms: a Systems Analysis of Evolution (Wiley, Hoboken, NJ, 1978).
Hong, J. W., Hendrix, D. A. & Levine, M. S. Shadow enhancers as a source of evolutionary novelty. Science 321, 1314 (2008).
Perry, M. W., Boettiger, A. N. & Levine, M. Multiple enhancers ensure precision of gap gene-expression patterns in the Drosophila embryo. Proc. Natl Acad. Sci. USA 108, 13570–13575 (2011).
Papakostas, S. et al. Gene pleiotropy constrains gene expression changes in fish adapted to different thermal conditions. Nat. Commun. 5, 4071 (2014).
Zalts, H. & Yanai, I. Developmental constraints shape the evolution of the nematode mid-developmental transition. Nat. Ecol. Evol. 1, 0113 (2017).
Seki, R. et al. Functional roles of Aves class-specific cis-regulatory elements on macroevolution of bird-specific features. Nat. Commun. 8, 14229 (2017).
Lindblad-Toh, K. et al. A high-resolution map of human evolutionary constraint using 29 mammals. Nature 478, 476–482 (2011).
Carroll, S. B., Grenier, J. K. & Weatherbee, S. D. From DNA to Diversity: Molecular Genetics and the Evolution of Animal Design 9th edn (Blackwell, Malden, MA, 2005
Pavlicev, M. & Wagner, G. P. A model of developmental evolution: selection, pleiotropy and compensation. Trends Ecol. Evol. 27, 316–322 (2012).
Fisher, R. A. The Genetical Theory of Natural Selection (Clarendon, Oxford, 1930).
Orr, H. A. Adaptation and the cost of complexity. Evolution 54, 13–20 (2000).
Satoh, N., Rokhsar, D. & Nishikawa, T. Chordate evolution and the three-phylum system. Proc. R. Soc. B 281, 20141729 (2014).
Sasagawa, Y. et al. Quartz-Seq: a highly reproducible and sensitive single-cell RNA sequencing method, reveals non-genetic gene-expression heterogeneity. Genome Biol. 14, R31 (2013).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
Li, L., Stoeckert, C. J. Jr. & Roos, D. S. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res. 13, 2178–2189 (2003).
Lechner, M. et al. Proteinortho: detection of (co-)orthologs in large-scale analysis. BMC Bioinformatics 12, 124 (2011).
Chen, F., Mackey, A. J., Vermunt, J. K. & Roos, D. S. Assessing performance of orthology detection strategies applied to eukaryotic genomes. PLoS ONE 2, e383 (2007).
Tena, J. J. et al. Comparative epigenomics in distantly related teleost species identifies conserved cis-regulatory nodes active during the vertebrate phylotypic period. Genome Res. 24, 1075–1085 (2014).
Hejnol, A. & Dunn, C. W. Animal evolution: are phyla real? Curr. Biol. 26, R424–R426 (2016).
Dunn, C. W., Zapata, F., Munro, C., Siebert, S. & Hejnol, A. Pairwise comparisons across species are problematic when analyzing funcional genomic data. Preprint at http://www.biorxiv.org/content/early/2017/02/09/107177 (2017).
Li, J. J., Huang, H., Bickel, P. J. & Brenner, S. E. Comparison of D. melanogaster and C. elegans developmental stages, tissues, and cells by modENCODE RNA-seq data. Genome Res. 24, 1086–1101 (2014).
Gene Ontology Consortium. Gene Ontology Consortium: going forward. Nucl. Acids Res. 43, D1049–D1056 (2015).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Georgi, B., Voight, B. F. & Bucan, M. From mouse to human: evolutionary genomics analysis of human orthologs of essential genes. PLoS Genet. 9, e1003484 (2013).
Acknowledgements
The EXPANDE (EXpression Profiling AloNg Development and Evolution) project and N.I. was supported in part by Grants in Aid from the Ministry of Education, Culture, Sports, Science and Technology, Japan (15H05603, 22128003, 24570243, 3902 and 17H06387) and the Platform for Dynamic Approaches to Living System from the Ministry of Education, Culture, Sports, Science and Technology, Japan. T.G.K. was supported in part by KAKENHI 16H04724. The research performed by J.-K.Y. was supported by grants from Academia Sinica and the Ministry of Science and Technology, Taiwan (AS-98-CDA-L06, 102-2311-B-001-011-MY3 and 104-2923-B-001-002-MY3). We thank K. Yamanaka for help collecting the embryos (mouse, chicken, turtle, frog and zebrafish), extracting RNAs and sample preparation using Quartz-Seq for early embryos from mice. We thank C. Tanegashima, K. Itomi, O. Nishimura and S. Kuraku for help constructing libraries, sequencing samples and quality checking some of the RNA-seq data. We thank F. Castellan for critically reading the manuscript. We thank Y. Uchida for providing high-resolution images of zebrafish embryos. We thank G. Renaud and J. Kelso for providing the robust demultiplexing method for analysing the ascidian sequencing data.
Author information
Authors and Affiliations
Consortia
Contributions
N.I., P.K. and S.K. conceived the study. S.F., K.S., T.-M.L., J.-K.Y., T.G.K., Y.S. and N.I. collected the samples. H.H., S.G., S.F., Y.S., T.G.K. and N.I. conducted the experiments needed for RNA-seq. F.L., S.L., G.Z. and H.H. made new gene sets in X. laevis and B. floridae. H.H., S.G., M.U., P.K., M.I. and N.I. analysed the data. N.I., J.-K.Y., M.U., T.G.K. and S.K. edited the paper. N.I. and P.K. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
A list of the members of the EXPANDE Consortium and their and affiliations appears in the Supplementary Information.
Electronic supplementary material
Supplementary Information
Supplementary Methods, Supplementary Notes, Supplementary Figures 1–13, Supplementary Tables 1–16
Supplementary Data
FPKM-based normalized gene expression profiles at each developmental stage for each species
Rights and permissions
About this article
Cite this article
Hu, H., Uesaka, M., Guo, S. et al. Constrained vertebrate evolution by pleiotropic genes. Nat Ecol Evol 1, 1722–1730 (2017). https://doi.org/10.1038/s41559-017-0318-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-017-0318-0
This article is cited by
-
Stability in gene expression and body-plan development leads to evolutionary conservation
EvoDevo (2023)
-
Weak gene–gene interaction facilitates the evolution of gene expression plasticity
BMC Biology (2023)
-
Potential contribution of intrinsic developmental stability toward body plan conservation
BMC Biology (2022)
-
Identification and characterization of male reproduction-related genes in pig (Sus scrofa) using transcriptome analysis
BMC Genomics (2020)
-
Inter-embryo gene expression variability recapitulates the hourglass pattern of evo-devo
BMC Biology (2020)