Abstract
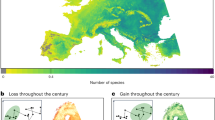
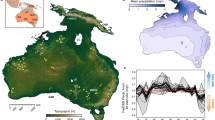
Human-mediated transport beyond biogeographic barriers has led to the introduction and establishment of alien species in new regions worldwide. However, we lack a global picture of established alien species richness for multiple taxonomic groups. Here, we assess global patterns and potential drivers of established alien species richness across eight taxonomic groups (amphibians, ants, birds, freshwater fishes, mammals, vascular plants, reptiles and spiders) for 186 islands and 423 mainland regions. Hotspots of established alien species richness are predominantly island and coastal mainland regions. Regions with greater gross domestic product per capita, human population density, and area have higher established alien richness, with strongest effects emerging for islands. Ants and reptiles, birds and mammals, and vascular plants and spiders form pairs of taxonomic groups with the highest spatial congruence in established alien richness, but drivers explaining richness differ between the taxa in each pair. Across all taxonomic groups, our results highlight the need to prioritize prevention of further alien species introductions to island and coastal mainland regions globally.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
di Castri, F. in Biological Invasions: A Global Perspective (eds Drake, J. A. ) 1–30 (Wiley, 1989).
Simberloff, D. et al. Impacts of biological invasions: what’s what and the way forward. Trends Ecol. Evol. 28, 58–66 (2013).
Lewis, S. L. & Maslin, M. A. Defining the Anthropocene. Nature 519, 171–180 (2015).
Blackburn, T. M. et al. A proposed unified framework for biological invasions. Trends Ecol. Evol. 26, 333–339 (2011).
Seebens, H. et al. Global trade will accelerate plant invasions in emerging economies under climate change. Glob. Chang. Biol. 21, 4128–4140 (2015).
Poessel, S. A. Beard, K. H., Callahan, C. M., Ferreira, R. B. & Stevenson, E. T. Biotic acceptance in introduced amphibians and reptiles in Europe and North America. Glob. Ecol. Biogeogr. 22, 192–201 (2013).
Essl, F. et al. Socioeconomic legacy yields and invasion debt. Proc. Natl Acad. Sci. USA 108, 203–207 (2011).
Jeschke, J. M . & Genovesi, P. Do biodiversity and human impact influence the introduction or establishment of alien mammals? Oikos 120, 57–64 (2011).
Blackburn, T. M., Cassey, P. & Lockwood, J. L. The island biogeography of exotic bird species. Glob. Ecol. Biogeogr. 17, 246–251 (2008).
Capinha, C., Essl, F., Seebens, H., Moser, D. & Pereira, H. M. The dispersal of alien species redefines biogeography in the Anthropocene. Science 348, 1248–1251 (2015).
Essl, F., Dullinger, S., Moser, D., Steinbauer, K. & Mang, T. Macroecology of global bryophyte invasions at different invasion stages. Ecography 38, 488–498 (2015).
van Kleunen, M. et al. Global exchange and accumulation of non-native plants. Nature 525, 100–103 (2015).
Dyer, E. E. et al. The global distribution and drivers of alien bird species richness. PLoS Biol. 15, e2000942 (2017).
Froese, R. & Pauly, D. (eds) FishBase v. 09/2015 (2015); http://www.fishbase.org
Guénard, B., Weiser, M. D., Gomez, K., Narula, N. & Economo, E. P. The Global Ant Biodiversity Informatics (GABI) database: synthesizing data on ant species geographic distribution. Myrmecol. News 24, 83–89 (2017).
World Spider Catalog Version 17.0 (Natural History Museum Bern, 2015) http://wsc.nmbe.ch
Brummit, R. K. World Geographical Scheme for Recording Plant Distributions 2nd edn. (Hunt Institute for Botanical Documentation, 2001).
Meyer, C., Kreft, H., Guralnick, R. & Jetz, W. Global priorities for an effective information basis of biodiversity distributions. Nat. Commun. 6, 8221 (2015).
Meyer, C., Weigelt, P. & Kreft, H. Multidimensional biases, gaps and uncertainties in global plant occurrence information. Ecol. Lett. 19, 992–1006 (2016).
Guénard, B., Weiser, M. D. & Dunn, R. R. Global models of ant diversity suggest regions where new discoveries are most likely are under disproportionate deforestation threat. Proc. Natl Acad. Sci. USA 109, 7368–7373 (2012).
Pyšek, P. et al. Disentangling the role of environmental and human pressures on biological invasions across Europe. Proc. Natl Acad. Sci. USA 107, 12157–12162 (2010).
Gaston, K. J. Global patterns of biodiversity. Nature 405, 220–227 (2000).
Lambdon, P. W. et al. Alien flora of Europe: species diversity, temporal trends, geographical patterns and research needs. Preslia 80, 101–149 (2008).
Denslow, J. S. Weeds in paradise: thoughts on the invasibility of tropical islands. Ann. Missouri Bot. Gard. 90, 119–127 (2003).
Lonsdale, W. M. Global patterns of plant invasions and the concept of invasibility. Ecology 80, 1522–1536 (1999).
Lockwood, J. L., Cassey, P. & Blackburn, T. M. The more you introduce the more you get: the role of colonization pressure and propagule pressure in invasion ecology. Divers. Distrib. 15, 904–910 (2009).
Gallardo, B. & Aldridge, D. C. The “dirty-dozen”: socio-economic factors amplify the invasion potential of 12 high-risk aquatic invasive species in Great Britain and Ireland. J. Appl. Ecol. 50, 757–766 (2013).
Gotzek, D. et al. Global invasion history of the tropical fire ant: a stowaway on the first global trade routes. Mol. Ecol. 24, 374–388 (2015).
Nentwig, W. Introduction, establishment rate, pathways and impact of spiders alien to Europe. Biol. Invas. 17, 2757–2778 (2015).
Genovesi, P ., Bacher, S ., Kobelt, M ., Pascal, M . & Scalera, R. in Handbook of Alien Species in Europe (eds DAISIE ) 119–128 (Springer, 2009).
Qian, H. & Ricklefs, R. E. Global concordance in diversity patterns of vascular plants and terrestrial vertebrates. Ecol. Lett. 11, 547–553 (2008).
Lever, C. in Encyclopedia of Biological Invasions (eds Simberloff, D. & Rejmánek, M. ) 1–4 (Univ. California Press, 2011).
Felemban, H. M. On the exotic birds imported into Jeddah, Saudi Arabia. Zool. Middle East 8, 15–16 (1993).
Dyer, E. E ., Redding, D. W . & Blackburn, T. M. The global avian invasions atlas, a database of alien bird distributions worldwide. Sci. Data 4, 170041 (2017).
Espinosa-Perez, H. & Ramirez, M. Exotic and invasive fishes in Mexico. Check List 11, 1627 (2015).
McDowall, R. M. The Reed Field Guide to New Zealand Freshwater Fishes (Reed, 2000 ).
NIWA Atlas of NZ Freshwater Fishes (accessed 24 May 2017) https://www.niwa.co.nz/freshwater-and-estuaries/nzffd/NIWA-fish-atlas
Ellender, B. R. & Weyl, O. L. F. A review of current knowledge, risk and ecological impacts associated with non-native freshwater fish introductions in South Africa. Aquat. Invasions 9, 117–132 (2014).
Skelton, P. H. A Complete Guide to the Freshwater Fishes of Southern Africa (Southern Book Publishers, 2001).
Invasive Species of Japan Database (Environmental Risk Research Center, National Institute for Environmental Studies, Japan, accessed 18 May 2016) https://www.nies.go.jp/biodiversity/invasive/index_en.html
Brazil Invasive Alien Species Database (Inter-American Biodiversity Information Network, accessed 17 May 2016) http://i3n.institutohorus.org.br/www
Ghosh, T., Powell, R. L., Elvidge, C. D., Baugh, K. E., Sutton, P. C. & Anderson, S. Shedding light on the global distribution of economic activity. Open Geogr. J. 3, 148–160 (2010).
Weigelt, P. & Kreft, H. Quantifying island isolation – insights from global patterns of insular plant species richness. Ecography 36, 417–429 (2013).
Pinheiro, J., Bates, D., DebRoy, S., Sarkar, D. & R Core Team . nlme: Linear and nonlinear mixed effects models. R Package Version 3.1-128 (2016).
Bjornstad, O.N. ncf: Spatial nonparametric covariance functions. R Package Version 1.1-7 (2016).
R Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2016).
Acknowledgements
This research benefited from support from the European Commission (COST Action TD1209). The Deutsche Forschungsgemeinschaft supported H.S. (DFG, grant SE 1891/2-1), M.v.K. (KL 1866/9-1) and M.W. (FZT 118), the Austrian Science Foundation supported F.E., B.L. and D.M. (FWF, grant I2086-B16). P.P. and J.P. were supported by the Academy of Sciences of the Czech Republic (no. RVO 67985939), Praemium Academiae award to P.P. and Czech Science Foundation (project no. 14-36079G). C. Capinha was supported by a postdoctoral grant from the Portuguese Foundation for Science and Technology (FCT/MCTES) and POPH/FSE (EC) grant SFRH/BPD/84422/2012. E.G.-B. was supported by the Spanish Ministry of Economy and Competitiveness (projects CGL2013-43822-R and CGL2015-69311-REDT). C.M. was supported by the Volkswagen Foundation through a Freigeist Fellowship.
Author information
Authors and Affiliations
Contributions
The GloNAF core team (M.v.K., P.P., W.D., F.E., J.P., M.W., H.K. and P.W.), T.M.B., H.S. and B.L. conceived the idea; W.D. coordinated data collation, and designed and led the analyses and writing with major inputs from F.E., D.M. M.v.K., P.P., H.K., M.W., J.P. and P.W., and further inputs from all other authors. Data were contributed by the GloNAF database for vascular plants, E.P.E. and B.G. for ants, C. Capinha, F.E., H.S. and P.G.-D. for amphibians and reptiles, T.M.B. and E.E.D. for birds, C. Casal, E.G.-B., P.F. and N.E.M. for fishes, P.C. and S.L.S for mammals, and W.N. for spiders. D.M. collected and calculated data on region area and sampling effort. C.M. contributed data on completeness of native species richness inventories.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Three Supplementary Figures; six Supplementary Tables (PDF 945 kb)
Rights and permissions
About this article
Cite this article
Dawson, W., Moser, D., van Kleunen, M. et al. Global hotspots and correlates of alien species richness across taxonomic groups. Nat Ecol Evol 1, 0186 (2017). https://doi.org/10.1038/s41559-017-0186
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41559-017-0186
This article is cited by
-
Multiple invasion routes have led to the pervasive introduction of earthworms in North America
Nature Ecology & Evolution (2024)
-
Mutualisms weaken the latitudinal diversity gradient among oceanic islands
Nature (2024)
-
Risk of introduction and establishment of alien vertebrate species in transboundary neighboring areas
Nature Communications (2024)
-
Invasive and native plants show different root responses to feedback-mediated soil heterogeneity
Plant and Soil (2024)
-
Effect of introduction pathways on the invasion success of non-native plants along environmental gradients
Biological Invasions (2024)