Abstract
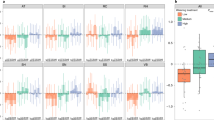
Plant–soil feedbacks (PSF) are important interactions that may influence range dynamics in a changing world. What remains largely unknown is the generality of plant–soil biotic interactions across populations and the potential role of specific soil biota, both of which are key for understanding how PSF might change future communities and ecosystems. We combined landscape-level field observations and experimental soil treatments to test whether a dominant tree alters soil environments to impact its own performance and range shifts towards higher elevations. We show: (1) soil conditioning by trees varies with elevation, (2) soil biota relate to PSF, (3) under simulated conditions, biotic PSF constrain range shifts at lower elevations but allow for expansions at higher elevations, and (4) differences in soil conditioning predict feedback outcomes in specific range-shift scenarios. These results suggest that variable plant–soil biotic interactions may influence the migration and fragmentation of tree species, and that models incorporating soil parameters will more accurately predict future species distributions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Coudon, C., Gegout, J. C., Piedallu, C. & Rameau, J. C. Soil nutritional factors improve models of plant species distribution: an illustration with Acer campestre (L.) in France. J. Biogeogr. 33, 1750–1763 (2006).
Beauregard, F. & de Blois, S. Beyond a climate-centric view of plant distribution: edaphic variables add value to distribution models. PLoS ONE 9, e92642 (2014).
van der Putten, W. H. Climate change, aboveground–belowground interactions, and species’ range shifts. Annu. Rev. Ecol. Evol. S. 43, 365–383 (2012).
van der Putten, W. H. et al. Plant–soil feedbacks: the past, the present and future challenges. J. Ecol. 101, 265–276 (2013).
Bailey, J. K. et al. Indirect genetic effects: an evolutionary mechanism linking feedbacks, genotypic diversity and coadaptation in a climate change context. Funct. Ecol. 28, 87–95 (2014).
Van Nuland, M. E. et al. Plant–soil feedbacks: connecting ecosystem ecology and evolution. Funct. Ecol. 30, 1032–1042 (2016).
Bever, J. D., Westover, K. M. & Antonovics, J. Incorporating the soil community into plant population dynamics: the utility of the feedback approach. J. Ecol. 85, 561–573 (1997).
Morriën, E. & van der Putten, W. H. Soil microbial community structure of range-expanding plant species differs from co-occurring natives. J. Ecol. 101, 1093–1102 (2013).
Engelkes, T. et al. Successful range-expanding plants experience less above-ground and below-ground enemy impact. Nature 456, 946–948 (2008).
Van Grunsven, R. H. A., van der Putten, W. H., Bezemer T. M. & Veenendaal, E. M. Plant–soil feedback of native and range-expanding plant species is insensitive to temperature. Oecologia 162, 1059–1069 (2010).
McCarthy-Neumann, S. & Ibáñez, I. Tree range expansion may be enhanced by escape from negative plant–soil feedbacks. Ecology 93, 2637–2649 (2012).
Gundale, M. J. et al. Interactions with soil biota shift from negative to positive when a tree species is moved outside its native range. New Phytol. 202, 415–421 (2014).
Jump, A. S., Mátyás, C. & Peñuelas, J. The altitude-for-latitude disparity in the range retractions of woody species. Trends Ecol. Evol. 24, 694–701 (2009).
Schweitzer, J. A. et al. Are there evolutionary consequences of plant–soil feedbacks along soil gradients? Funct. Ecol. 28, 55–64 (2014).
terHorst, C. P. & Zee, P. C. Eco-evolutionary dynamics in plant–soil feedbacks. Funct. Ecol. 30, 1062–1072 (2016).
Chen, I. C., Hill, J. K., Ohlemüller, R., Roy, D. B. & Thomas, C. D. Rapid range shifts of species associated with high levels of climate warming. Science 333, 1024–1026 (2011).
Bridle, J. R. & Vines, T. H. Limits to evolution at range margins: when and why does adaptation fail? Trends Ecol. Evol. 22, 140–147 (2007).
Angert, A. L. The niche, limits to species’ distributions, and spatiotemporal variation in demography across the elevation ranges of two monkeyflowers. Proc. Natl Acad. Sci. USA 106, 19693–19698 (2009).
Ettema, C. H. & Wardle, D. A. (2002). Spatial soil ecology. Trends Ecol. Evol. 17, 177–183 (2002).
Yang, Y. et al. The microbial gene diversity along an elevation gradient of the Tibetan grassland. ISME J. 8, 430–440 (2014).
Wagg, C., Husband, B. C., Green, D. S., Massicotte, H. B. & Peterson, R. L. Soil microbial communities from an elevational cline differ in their effect on conifer seedling growth. Plant Soil 340, 491–504 (2010).
Classen, A. T. et al. Direct and indirect effects of climate change on soil microbial and soil microbial–plant interactions: what lies ahead? Ecosphere 6, 1–21 (2015).
Sedlacek, J. F., Bossdorf, O., Cortés, A. J., Wheeler, J. A. & van Kleunen, M. What role do plant–soil interactions play in the habitat suitability and potential range expansion of the alpine dwarf shrub Salix herbacea? Basic Appl. Ecol. 15, 305–315 (2014).
Bardgett, R. D. & Wardle, D. A. Aboveground–Belowground Linkages: Biotic Interactions, Ecosystem Processes, and Global Change (Oxford Univ. Press, 2010).
Fierer, N. & Jackson, R. The diversity and biogeography of soil bacterial communities. Proc. Natl Acad. Sci. USA 103, 626–631 (2006).
Woolbright, S. A, Whitham, T. G ., Gehring, C. A ., Allan, G. J & Bailey, J. K. Climate relicts and their associated communities as natural ecology and evolution laboratories. Trends Ecol. Evol. 29, 406–416 (2014).
Kardol, P., De Deyn, G. B., Laliberte, E., Mariotte, P. & Hawkes. C. V. Biotic plant–soil feedbacks across temporal scales. J. Ecol. 101, 309–315 (2013).
Sundqvist, M. K., Sanders, N. J. & Wardle, D. A. Community and ecosystem responses to elevational gradients: processes, mechanisms, and insights for global change. Annu. Rev. Ecol. Evol. S. 44, 261–280 (2013).
Braatne, J. H., Rood, S. B. & Heilman P. E. in Biology of Populus and its Implications for Management and Conservation (eds Stattler, R. F. et al.) 57–85 (NRC Research, 1996).
Capon, S. J. et al. Riparian ecosystems in the 21st century: hotspots for climate change adaptation? Ecosystems 16, 359–381 (2013).
Fischer, D. G. et al. Plant genetic effects on soils under climate change. Plant Soil 379, 1–19 (2013).
Hargreaves, A. L., Samis, K. E. & Eckert, C. G. Are species’ range limits simply niche limits writ large? A review of transplant experiments beyond the range. Am. Nat. 183, 157–173 (2014).
Ordonez, A. & Williams, J. W. Climatic and biotic velocities for woody taxa distributions over the last 16 000 years in eastern North America. Ecol. Lett. 16, 773–781 (2013).
Evans, L. M. et al. Geographical barriers and climate influence demographic history in narrowleaf cottonwoods. Heredity 114, 387–396 (2015).
Holeski, L. M., Zinkgraf, M. S., Couture, J. J., Whitham, T. G. & Lindroth, R. L. Transgenerational effects of herbivory in a group of long-lived tree species: maternal damage reduces offspring allocation to resistance traits, but not growth. J. Ecol. 101, 1062–1073 (2013).
Madritch, M. D., Greene, S. L. & Lindroth, R. L. Genetic mosaics of ecosystem functioning across aspen-dominated landscapes. Oecologia 160, 119–127 (2009).
Gehring, C. A., Mueller, R. C. & Whitham, T. G. Environmental and genetic effects on the formation of ectomycorrhizal and arbuscular mycorrhizal associations in cottonwoods. Oecologia 149, 158–164 (2006).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Krohn, A. et al. Optimization of 16S amplicon analysis using mock communities: implications for estimating community diversity. Preprint at http://doi.org/10.7287/peerj.preprints.2196v2 (2016).
Bever, J. D. et al. Rooting theories of plant community ecology in microbial interactions. Trends Ecol. Evol. 25, 468–478 (2010).
Brinkman, P. E., van der Putten, W. H., Bakker, E. J. & Verhoeven, K. Plant–soil feedback: experimental approaches, statistical analyses and ecological interpretations. J. Ecol. 98, 1063–1073 (2010).
Sykorova, Z., Ineichen, K., Wiemken, A. & Redecker, D. The cultivation bias: different communities of arbuscular mycorrhizal fungi detected in roots from the field, from bait plants transplanted to the field, and from a greenhouse trap experiment. Mycorrhiza 18, 1–14 (2007).
Zinke, P. J. The pattern of influence of individual forest trees on soil properties. Ecology 43, 130–133 (1962).
McCarthy-Neumann, S. & Kobe, R. K. Conspecific and heterospecific plant–soil feedbacks influence survivorship and growth of temperate tree seedlings. J. Ecol. 98, 408–418 (2010).
Blanquart, F., Kaltz, O., Nuismer, S. L. & Gandon, S. A practical guide to measuring local adaptation. Ecol. Lett. 16, 1195–1205 (2013).
Ke, P. J., Miki, T. & Ding, T. S. The soil microbial community predicts the importance of plant traits in plant–soil feedback. New Phytol. 206, 329–341 (2015).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2013); http://www.r-project.org
Acknowledgements
This material is based on work supported by the National Science Foundation Graduate Research Fellowship under Grant No. DGE-0929298. Funding for the project was also provided from the University of Tennessee. We thank A. Krohn at the Environmental Genetics and Genomics Laboratory for sequencing and bioinformatics assistance, as well as A. Classen and N. Sanders for providing helpful comments on the manuscript. Special thanks to P. Patterson at Northern Arizona University as well as I. Ware, K. McFarland, P. Meidl, C. Daws, E. Johnson, R. Wooliver, L. Mueller, A. Pfennigwerth and R. Zenni for field, greenhouse and lab support.
Author information
Authors and Affiliations
Contributions
M.E.V.N., J.K.B. and J.A.S. participated in the study design. M.E.V.N. performed the field work, data collection and statistical analyses. All authors discussed the results. M.E.V.N. wrote the initial manuscript draft, with significant edits from J.K.B. and J.A.S.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Discussion; Supplementary Methods; Supplementary Tables 1–10; Supplementary Figures 1–7 (PDF 1728 kb)
Rights and permissions
About this article
Cite this article
Van Nuland, M., Bailey, J. & Schweitzer, J. Divergent plant–soil feedbacks could alter future elevation ranges and ecosystem dynamics. Nat Ecol Evol 1, 0150 (2017). https://doi.org/10.1038/s41559-017-0150
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41559-017-0150
This article is cited by
-
Climate-driven convergent evolution in riparian ecosystems on sky islands
Scientific Reports (2023)
-
Grass species identity shapes communities of root and leaf fungi more than elevation
ISME Communications (2022)
-
Stacked distribution models predict climate-driven loss of variation in leaf phenology at continental scales
Communications Biology (2022)
-
Climate-driven divergence in plant-microbiome interactions generates range-wide variation in bud break phenology
Communications Biology (2021)
-
Conspecific and heterospecific plant–soil biota interactions of Lonicera japonica in its native and introduced range: implications for invasion success
Plant Ecology (2021)