Abstract
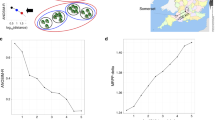
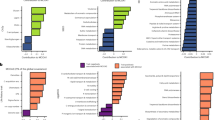
Understanding the processes that are driving variation of natural microbial communities across space or time is a major challenge for ecologists. Environmental conditions strongly shape the metabolic function of microbial communities; however, other processes such as biotic interactions, random demographic drift or dispersal limitation may also influence community dynamics. The relative importance of these processes and their effects on community function remain largely unknown. To address this uncertainty, here we examined bacterial and archaeal communities in replicate ‘miniature’ aquatic ecosystems contained within the foliage of wild bromeliads. We used marker gene sequencing to infer the taxonomic composition within nine metabolic functional groups, and shotgun environmental DNA sequencing to estimate the relative abundances of these groups. We found that all of the bromeliads exhibited remarkably similar functional community structures, but that the taxonomic composition within individual functional groups was highly variable. Furthermore, using statistical analyses, we found that non-neutral processes, including environmental filtering and potentially biotic interactions, at least partly shaped the composition within functional groups and were more important than spatial dispersal limitation and demographic drift. Hence both the functional structure and taxonomic composition within functional groups of natural microbial communities may be shaped by non-neutral and roughly separate processes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Falkowski, P. G., Fenchel, T. & Delong, E. F. The microbial engines that drive Earth’s biogeochemical cycles. Science 320, 1034–1039 (2008).
Sunagawa, S. et al. Structure and function of the global ocean microbiome. Science 348, 1261359 (2015).
Zaikova, E. et al. Microbial community dynamics in a seasonally anoxic fjord: Saanich Inlet, British Columbia. Environ. Microbiol. 12, 172–191 (2010).
Powell, J. R. et al. Deterministic processes vary during community assembly for ecologically dissimilar taxa. Nat. Commun. 6, 8444 (2015).
Strom, S. L. Microbial ecology of ocean biogeochemistry: a community perspective. Science 320, 1043–1045 (2008).
Lima-Mendez, G. et al. Determinants of community structure in the global plankton interactome. Science 348,1262073 (2015).
Ofijeru, I. D. et al. Combined niche and neutral effects in a microbial wastewater treatment community. Proc. Natl Acad. Sci. USA 107, 15345–15350 (2010).
Burke, C., Steinberg, P., Rusch, D., Kjelleberg, S. & Thomas, T. Bacterial community assembly based on functional genes rather than species. Proc. Natl Acad. Sci. USA 108, 14288–14293 (2011).
Martiny, J. B. H. et al. Microbial biogeography: putting microorganisms on the map. Nat. Rev. Microbiol. 4, 102–112 (2006).
Raes, J., Letunic, I., Yamada, T., Jensen, L. J. & Bork, P. Toward molecular trait-based ecology through integration of biogeochemical, geographical and metagenomic data. Mol. Syst. Biol. 7, 473 (2011).
Louca, S., Parfrey, L. W. & Doebeli, M. Decoupling function and taxonomy in the global ocean microbiome. Science 353, 1272–1277 (2016).
Nelson, M. B., Martiny, A. C. & Martiny, J. B. H. Global biogeography of microbial nitrogen-cycling traits in soil. Proc. Natl Acad. Sci. USA 113, 8033–8040 (2016).
Fukami, T., Martijn Bezemer, T., Mortimer, S. R. & Putten, W. H. Species divergence and trait convergence in experimental plant community assembly. Ecol. Lett. 8, 1283–1290 (2005).
Helsen, K., Hermy, M. & Honnay, O. Trait but not species convergence during plant community assembly in restored semi-natural grasslands. Oikos 121, 2121–2130 (2012).
Whitman, W. B., Coleman, D. C. & Wiebe, W. J. Prokaryotes: the unseen majority. Proc. Natl Acad. Sci. USA 95, 6578–6583 (1998).
Lande, R., Engen, S. & Saether, B. Stochastic Population Dynamics in Ecology and Conservation (Oxford Univ. Press, 2003).
Fernandez, A. et al. How stable is stable? Function versus community composition. Appl. Environ. Microbiol. 65, 3697–3704 (1999).
Vanwonterghem, I. et al. Deterministic processes guide long-term synchronised population dynamics in replicate anaerobic digesters. ISME J. 8, 2015–2028 (2014).
Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Prosser, J. I. Dispersing misconceptions and identifying opportunities for the use of ‘omics’ in soil microbialecology. Nat. Rev. Microbiol. 13, 439–446 (2015).
Goffredi, S. K., Kantor, A. H. & Woodside, W. T. Aquatic microbial habitats within a neotropical rainforest: bromeliads and pH-associated trends in bacterial diversity and composition. Microb. Ecol. 61, 529–542 (2011).
Farjalla, V. F. et al. Ecological determinism increases with organism size. Ecology 93, 1752–1759 (2012).
Srivastava, D. S. et al. Are natural microcosms useful model systems for ecology? Trends Ecol. Evol. 19, 379–384 (2004).
Martinson, G. O. et al. Methane emissions from tank bromeliads in neotropical forests. Nat. Geosci. 3, 766–769 (2010).
Goffredi, S. K., Jang, G., Woodside, W. T. & Ussler, W. Bromeliad catchments as habitats for methanogenesis in tropical rainforest canopies. Front. Microbiol. 2,256 (2011).
Marino, N. A. C., Srivastava, D. S. & Farjalla, V. F. Aquatic macroinvertebrate community composition in tank-bromeliads is determined by bromeliad species and its constrained characteristics. Insect Cons. Div. 6, 372–380 (2013).
Yarza, P. et al. Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences. Nat. Rev. Microbiol. 12, 635–645 (2014).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Canfield, D. E. & Thamdrup, B. Towards a consistent classification scheme for geochemical environments, or, why we wish the term ‘suboxic’ would go away. Geobiology 7, 385–392 (2009).
Atwood, T. B. et al. Predator-induced reduction of freshwater carbon dioxide emissions. Nat. Geosci. 6, 191–194 (2013).
Chase, J. M. & Myers, J. A. Disentangling the importance of ecological niches from stochastic processes across scales. Phil. Trans. R. Soc. Lond. B 366, 2351–2363 (2011).
Gotelli, N. J. Null model analysis of species co-occurrence patterns. Ecology 81, 2606–2621 (2000).
Ulrich, W. & Gotelli, N. J. Null model analysis of species associations using abundance data. Ecology 91, 3384–3397 (2010).
Ulrich, W. Species co-occurrences and neutral models: reassessing J. M. Diamond’s assembly rules. Oikos 107, 603–609 (2004).
Horner-Devine, M. C. & Bohannan, B. J. M. Phylogenetic clustering and overdispersion in bacterial communities. Ecology 87, S100–S108 (2006).
Pausas, J. G. & Verdu, M. The jungle of methods for evaluating phenotypic and phylogenetic structure of communities. Bioscience 60, 614–625 (2010).
Sloan, W. T. et al. Quantifying the roles of immigration and chance in shaping prokaryote community structure. Environ. Microbiol. 8, 732–740 (2006).
Sloan, W. T., Woodcock, S., Lunn, M., Head, I. M. & Curtis, T. P. Modeling taxa-abundance distributions in microbial communities using environmental sequence data. Microb. Ecol. 53, 443–455 (2007).
Legendre, P. & Legendre, L. Developments in Environmental Modelling 2nd edn (Elsevier, 1998).
Legendre, P. Studying beta diversity: ecological variation partitioning by multiple regression and canonicalanalysis. J. Plant Ecol. 1, 3–8 (2008).
Suttle, C. A. Marine viruses—major players in the global ecosystem. Nat. Rev. Microbiol. 5, 801–812 (2007).
Shapiro, O. H. & Kushmaro, A. Bacteriophage ecology in environmental biotechnology processes. Curr. Opin. Biotechnol. 22, 449–455 (2011).
Carrias, J.-F. et al. Two coexisting tank bromeliads host distinct algal communities on a tropical inselberg. Plant Biol. 16, 997–1004 (2014).
Langenheder, S., Lindstrom, E. S. & Tranvik, L. J. Weak coupling between community composition and functioning of aquatic bacteria. Limnol. Oceanogr. 50, 957–967 (2005).
Strickland, M. S., Lauber, C., Fierer, N. & Bradford, M. A. Testing the functional significance of microbial community composition. Ecology 90, 441–451 (2009).
Fierer, N. et al. Cross-biome metagenomic analyses of soil microbial communities and their functional attributes. Proc. Natl Acad. Sci. USA 109, 21390–21395 (2012).
Vanwonterghem, I., Jensen, P. D., Rabaey, K. & Tyson, G. W. Genome-centric resolution of microbial diversity, metabolism and interactions in anaerobic digestion. Environ. Microbiol. 18, 3144–3158 (2016).
Reed, D. C. et al. Predicting the response of the deep-ocean microbiome to geochemical perturbations by hydrothermal vents. ISME J. 9, 1857–1869 (2015).
Nazareno, A. G. & Laurance, W. F. Brazil’s drought: beware deforestation. Science 347, 1427–1427 (2015).
Golterman, H. Clymo, R. & Ohnstad, M. Methods for Physical and Chemical Analysis of Freshwaters 2nd edn, 169 (IBP Handbook No. 8, Blackwell 1978).
Zagatto, E., Jacintho, A., Mortatti, J. & Bergamin, F. H. An improved flow injection determination of nitrite in waters by using intermittent flows. Anal. Chim. Acta 120, 399–403 (1980).
Fofonoff, N. P. & Millard-Junior, R. Algorithms for Computation of Fundamental Properties of Seawater . UNESCO technical paper in marine science 44 (UNESCO, 1983).
Andrade-Eiroa, A., Canle, M. & Cerda, V. Environmental applications of excitation-emission spectrofluorimetry: an in-depth review II. Appl. Spec. Rev. 48, 77–141 (2013).
Murphy, K. R., Stedmon, C. A., Graeber, D. & Bro, R. Fluorescence spectroscopy and multi-way techniques. PARAFAC. Anal. Meth. 5, 6557–6566 (2013).
Stedmon, C. A., Markager, S. & Bro, R. Tracing dissolved organic matter in aquatic environments using a new approach to fluorescence spectroscopy. Mar. Chem. 82, 239–254 (2003).
Stedmon, C. A. & Bro, R. Characterizing dissolved organic matter fluorescence with parallel factor analysis: a tutorial. Limnol. Oceanogr. Meth. 6, 572–579 (2008).
Murphy, K. R., Stedmon, C. A., Wenig, P. & Bro, R. Openfluor—an online spectral library of auto-fluorescence by organic compounds in the environment. Anal. Meth. 6, 658–661 (2014).
Jørgensen, L. et al. Global trends in the fluorescence characteristics and distribution of marine dissolved organic matter. Mar. Chem. 126, 139–148 (2011).
Murphy, K. R. et al. Organic matter fluorescence in municipal water recycling schemes: Toward a unified PARAFAC model. Environ. Sci. Technol. 45, 2909–2916 (2011).
Osburn, C. L., Wigdahl, C. R., Fritz, S. C. & Saros, J. E. Dissolved organic matter composition and photoreactivity in prairie lakes of the U.S. Great Plains. Limnol. Oceanogr. 56, 2371–2390 (2011).
Yamashita, Y., Boyer, J. N. & Jaffe, R. Evaluating the distribution of terrestrial dissolved organic matter in a complex coastal ecosystem using fluorescence spectroscopy. Cont. Shelf Res. 66, 136–144 (2013).
Caporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 6, 1621–1624 (2012).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Li, W., Fu, L., Niu, B., Wu, S. & Wooley, J. Ultrafast clustering algorithms for metagenomic sequence analysis. Brief. Bioinform. 13, 656–668 (2012).
Gevers, D. et al. Re-evaluating prokaryotic species. Nat. Rev. Microbiol. 3, 733–739 (2005).
Martiny, A. C., Tai, A. P., Veneziano, D., Primeau, F. & Chisholm, S. W. Taxonomic resolution, ecotypes and the biogeography of Prochlorococcus. Environ. Microbiol. 11, 823–832 (2009).
Koeppel, A. F. & Wu, M. Species matter: the role of competition in the assembly of congeneric bacteria. ISME J. 8, 531–540 (2014).
Keswani, J. & Whitman, W. B. Relationship of 16S rRNA sequence similarity to DNA hybridization in prokaryotes. Int. J. Syst. Evol. Microbiol. 51, 667–678 (2001).
Stackebrandt, E. & Ebers, J. Taxonomic parameters revisited: tarnished gold standards. Microbiol. Today 33, 152 (2006).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Pruesse, E. et al. SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 35, 7188–7196 (2007).
Caporaso, J. G. et al. PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26, 266–267 (2010).
Price, M. N., Dehal, P. S. & Arkin, A. P. Fasttree: computing large minimum evolution trees with profiles instead of a distance matrix. Mol. Biol. Evol. 26, 1641–1650 (2009).
Kubo, K. et al. Archaea of the Miscellaneous Crenarchaeotal Group are abundant, diverse and widespread in marine sediments. ISME J. 6, 1949–1965 (2012).
Zhang, J., Kobert, K., Flouri, T. & Stamatakis, A. PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30, 614–620 (2014).
Hahn, A., Hanson, N., Kim, D., Konwar, K. & Hallam, S. Assembly independent functional annotation of short-read data using SOFA: short-ORF functional annotation. In IEEE Conf. Comput. Intel. Bioinformatics Comput. Biol. 1–6 (IEEE, 2015).
Konwar, K. M. et al. MetaPathways v2.5: quantitative functional, taxonomic and usability improvements. Bioinformatics 31, 3345–3347 (2015).
Tatusova, T., Ciufo, S., Fedorov, B., O’Neill, K. & Tolstoy, I. RefSeq microbial genomes database: new representation and annotation strategy. Nucleic Acids Res. 42, D553–D559 (2014).
Wolda, H. Similarity indices, sample size and diversity. Oecologia 50, 296–302 (1981).
Hester, E. R., Barott, K. L., Nulton, J., Vermeij, M. J. & Rohwer, F. L. Stable and sporadic symbiotic communities of coral and algal holobionts. ISME J. 10, 1157–1169 (2015).
Chase, J. M., Kraft, N. J. B., Smith, K. G., Vellend, M. & Inouye, B. D. Using null models to disentangle variation in community dissimilarity from variation in α-diversity. Ecosphere 2, 1–11 (2011).
Connor, E. F. & Simberloff, D. The assembly of species communities: chance or competition? Ecology 60, 1132–1140 (1979).
Strona, G., Nappo, D., Boccacci, F., Fattorini, S. & San-Miguel-Ayanz, J. A fast and unbiased procedure to randomize ecological binary matrices with fixed row and column totals. Nat. Commun. 5, 4114 (2014).
Hausdorf, B. & Hennig, C. Null model tests of clustering of species, negative co-occurrence patterns and nestedness in meta-communities. Oikos 116, 818–828 (2007).
Chao, A., Jost, L., Chiang, S., Jiang, Y.-H. & Chazdon, R. L. A two-stage probabilistic approach to multiple-community similarity indices. Biometrics 64, 1178–1186 (2008).
Chave, J., Chust, G. & Thebaud, C. in Scaling Biodiversity (eds Storch, D. et al. ) 150–167 (Cambridge Univ. Press, 2007).
Hubbell, S. P. The Unified Neutral Theory of Biodiversity and Biogeography Vol. 32 (Princeton Univ. Press, 2001).
Ochman, H. & Wilson, A. Evolution in bacteria: evidence for a universal substitution rate in cellular genomes. J. Mol. Evol. 26, 74–86 (1987).
Kuo, C.-H. & Ochman, H. Inferring clocks when lacking rocks: the variable rates of molecular evolution in bacteria. Biol. Direct 4, 35 (2009).
Eliason, S. R. Maximum Likelihood Estimation: Logic and Practice (SAGE, 1993).
Burns, A. R. et al. Contribution of neutral processes to the assembly of gut microbial communities in thezebrafish over host development. ISME J. 10, 655–664 (2015).
Venkataraman, A. et al. Application of a neutral community model to assess structuring of the human lung microbiome. mBio 6, e02284-14 (2015).
Holyoak, M., Leibold, M. & Holt, R. Metacommunities: Spatial Dynamics and Ecological Communities (Univ. Chicago Press, 2005).
McCullagh, P. & Nelder, J. Generalized Linear Models. Chapman & Hall/CRC Monographs on Statistics & Applied Probability 2nd edn (Taylor & Francis, 1989).
Seabold, J. & Perktold, J. Statsmodels: econometric and statistical modeling with Python. In Proc. 9th Python Sci. Conf. (eds Jones, E. & Millman, J.) 57–61 (SciPy, 2010).
Shao, J. Linear model selection by cross-validation. J. Am. Stat. Assoc. 88, 486–494 (1993).
P.S.A.S. SAS/STAT 9.1 User’s Guide: The REG Procedure (SAS Institute, 2008).
Borcard, D., Legendre, P. & Drapeau, P. Partialling out the spatial component of ecological variation. Ecology 73, 1045–1055 (1992).
Field, A., Miles, J. & Field, Z. Discovering Statistics Using R (SAGE, 2012).
Acknowledgements
We thank M. Chen for help with the molecular work. We thank A. L. Gonzalez and A. MacDonald for discussions and comments on our paper. We thank T. Benevides for helping with the absorption measurements. S.L. acknowledges the financial support of the Department of Mathematics, University of British Columbia. S.L. and M.D. acknowledge the support of Natural Sciences and Engineering Research Council (NSERC). V.F.F. is grateful to the Brazilian Council for Research, Development and Innovation (CNPq) for research funds (Pesquisador Visitante Especial, PVE, Research Grant 400454/2014-9) and productivity grants. S.M.S.J. acknowledges the post-graduate scholarship provided by Fundação de Amparo à Pesquisa do Estado do Rio de Janeiro (FAPERJ). J.S.L. acknowledges the financial support of Coordenacao de Aperfeicoamento de Pessoal de Ensino Superior (CAPES). We thank M. P. F. Barros, A. R. Soares, J. L. Nepomuceno and their research groups of the Nucleus of Ecology and Socio-Environmental Development of Macae (NUPEM/UFRJ) for proving field and laboratory assistance during the samplings.
Author information
Authors and Affiliations
Contributions
V.F.F., S.L., S.M.S.J., A.P.F.P. and J.S.L. performed the field work. V.F.F. and S.M.S.J. performed the chemical measurements in the laboratory. S.L. performed the molecular work in the laboratory, the DNA sequence analysis and the statistical analyses. S.L., M.D., V.F.F., D.S.S. and L.W.P. interpreted the statistical findings. S.L. wrote a first draft of the manuscript, and all authors contributed to the final preparation of the manuscript. M.D. and V.F.F. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Supplementary information
Supplementary Figures 1–15, Supplementary Tables 1–10 (PDF 12771 kb)
Supplementary Data
Functional annotations of prokaryotic taxa (TXT 18 kb)
Rights and permissions
About this article
Cite this article
Louca, S., Jacques, S., Pires, A. et al. High taxonomic variability despite stable functional structure across microbial communities. Nat Ecol Evol 1, 0015 (2017). https://doi.org/10.1038/s41559-016-0015
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41559-016-0015
This article is cited by
-
Comparative analysis on root exudate and rhizosphere soil bacterial assembly between tomatoes and peppers infected by Ralstonia
Chemical and Biological Technologies in Agriculture (2024)
-
Plant–plant and plant–soil interactions under drought and the presence of invasive buffelgrass (Cenchrus ciliaris)
Biological Invasions (2024)
-
Evolutionary ecology of microbial populations inhabiting deep sea sediments associated with cold seeps
Nature Communications (2023)
-
Functional conservation of microbial communities determines composition predictability in anaerobic digestion
The ISME Journal (2023)
-
Distinct taxonomic and ecological functions of microbiome in sediments of different depth in Bohai Sea and Yellow Sea
Journal of Oceanology and Limnology (2023)