Abstract
Plasticity enhances species fitness and survival under climate change. Ocean acidification poses a potential threat to copepods, a major zooplankton group that serves as a key link between the lower and higher trophic levels in the marine environment, yet the mechanisms underlying different adaptive responses remain poorly understood. Here we show that although elevated CO2 can exert negative effects on reproduction of Paracyclopina nana, multigenerational plasticity can enable recovery after three generations. By integrating the methylome and transcriptome with the draft genome and undertaking DNA methylation treatments, we demonstrate the vital role of epigenetic modifications in ocean acidification responses and identify regions associated with reproductive resilience. Our results demonstrate that DNA methylation might play an important role in enhancing species fitness of copepods and that failing to consider phenotypic plasticity could lead to overestimation of species’ vulnerabilities.
Similar content being viewed by others
Main
The adaptability of marine species and ecosystems to elevated dissolved CO2 levels is a key driver of ecosystem resilience. Accumulating laboratory and field evidence show that the impending problem of ocean acidification (OA) can substantially and negatively affect marine species that build skeletons and shells of biogenic calcium carbonate (for example, oysters, corals, echinoderms and foraminifera) in ways that could lead to species extinction1. Recent studies have further demonstrated that OA can also disrupt important ecological processes (for example, competition, predator–prey relationships, habitat selection and navigation) of some non-calcified invertebrates and reef fish2,3. In particular, adverse reproductive outcomes at elevated CO2 have been found in a variety of primary consumers, including crustaceans and zooplankton4 (Supplementary Table 1), implying that elevated CO2-induced OA may ultimately affect the sustainability of species and trophodynamics of marine ecosystems.
Whether marine animals are able to adapt to OA by adjusting their reproductive strategies, let alone the underlying mechanisms involved, remains largely unknown. Phenotypic plasticity is an adaptive strategy that enables marine organisms to cope with elevated CO25. Plasticity in energy allocation, behavioural, morphological or physiological traits has been validated as a key survival factor in adaptation to environmental changes6. For example, copepods can spare more energy for reproduction at the expense of other functions to cope with the stress posed by elevated CO27,8 (regulation of physiological homeostasis in response to elevated CO2). In addition to the energetic trade-off between acidification tolerance traits and plasticity, recent ecological and evolutionary studies on phenotypic plasticity have focused on gene expression plasticity in response to environmental changes9,10. In particular, environmentally induced epigenetic variation is probably a major molecular mechanism for enhancing phenotypic plasticity. Indeed, previous studies have demonstrated that epigenetic changes allow calcifying marine invertebrates (for example, corals and oysters) to adapt to OA11,12,13,14. However, epigenetic regulation related to the intergenerational acclimation of non-calcifiers (for example, crustaceans) to elevated CO2 remains largely unexplored15,16. Furthermore, little is known about epigenetic plasticity in copepods, despite the presence/absence of high variations in genome-wide cytosine and guanine dinucleotide (CpG) methylation across arthropods17. Given the rapid adaptation reported in multiple copepod taxa18,19, understanding the epigenetic plasticity associated with copepod phenotypes is particularly important for predicting the effects of OA on zooplankton communities.
Copepods not only play an important role in regulating the global biogeochemical cycle and trophodynamics by contributing to the cycling of elements in the ocean20 but also serve as the key link between the lower and higher trophic levels21 and support fishery production22,23. Using the small cyclopoid copepod Paracyclopina nana as a model, we investigated intergenerational changes in DNA methylation patterns of copepods in response to elevated CO2, focusing on intergenerational plasticity. On the basis of the intergenerational effects observed in copepods exposed to elevated CO219,24, we further hypothesized that epigenetic plasticity could enable copepods to better cope with OA. Although the exoskeletons of copepods composed of polysaccharide chitin are unrelated to OA, negative impacts of elevated CO2 on growth and reproduction of some copepods have been reported24,25. This suggests that OA may affect copepod sustainability, which may cascade into adverse effects on ocean trophodynamics over a prolonged period. In this paper, we reveal that specific epigenetic changes induced by multigenerational OA exposure are important plasticity characteristics that enable small copepods to cope with elevated CO2.
Experimental paradigm
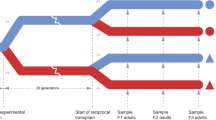
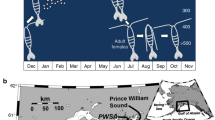
The copepod Paracyclopina sp. thrives in brackish water with large pH fluctuations26 (Fig. 1a). The East Sea (Sea of Japan) is a strong anthropogenic CO2 sink and is much more susceptible to OA than other oceans27 (Fig. 1b). In addition, intense biogeochemical transformations (for example, upwelling of deeper acidified seawater) in brackish water often lead to pH values reaching the pH projected for other oceans for the year 210028. Thus, brackish-water species such as P. nana will face the challenge presented by OA associated with climate change in the future. In this Article, we hypothesize that OA can induce epigenetic changes (genomic DNA methylation) associated with transgenerational reproductive resilience in P. nana, which could improve the species’s ability to cope with long-term OA. To minimize both genetic variations and possible past pH disturbance, we selected females from a line of P. nana inbred since 2000 (Methods). We then hatched copepod nauplii for another three generations at pH 8.0. The final offspring generation was raised at pH 8.0 (F0N), pH 7.7 or pH 7.3 (F0M). In the next generations, the pH treatment groups were divided into two batches. One batch was continuously maintained under elevated CO2 for another two generations (multigenerational groups, F1M, F2M), and the other batch was returned to the control condition and reared for another two generations (transgenerational groups, F1T, F2T) (Fig. 1c and Methods).
a, Image of an ovigerous (egg-bearing) P. nana female. b, Map showing P. nana collection location. c, Experimental schematic diagram. The three pH levels represent present-day control conditions (pH 8.0), a near-future scenario (pH 7.7) and extremely low-pH conditions expected to be reached by the year 2300 according to representative concentration pathway (RCP) 8.5 (pH 7.3). See Methods for detailed experimental design. Basemap in b created using Google Earth (https://goo.gl/maps/MV6LtgDFf1z66KYw6; map data: Google, TMap Mobility). Credit (DNA icon): Flaticon.com.
Reproductive plasticity during multigenerational exposure
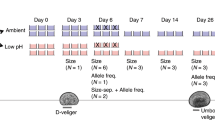
To understand the responses and intergenerational plasticity of P. nana to OA, we examined the effects of OA on in vivo endpoints related to pH tolerance. Throughout the experiment, neither male nor female developmental time nor body length was affected by OA stress in the pH 7.3 or pH 7.7 groups (Fig. 2a,b; P < 0.05). Similarly, elevated CO2 did not affect the transgenerational exposure groups (F0T, F1T and F2T) (Extended Data Fig. 1; P < 0.05). These results are similar to those reported for the calanoid copepod Calanus sp., which is not sensitive to elevated CO2 conditions21. By contrast, significant decreases in fecundity and sex ratio were observed in both the F0M and F1M groups (P < 0.05; Fig. 2c,d). These results suggest that multigenerational exposure to elevated CO2 could threaten the population dynamics of this copepod. However, both parameters recovered to the control level (pH 8.0) in the F2M group (P < 0.05), suggesting that multigenerational exposure can alleviate the adverse effects of OA. We found that F2M’s reductions in offspring fecundity diminished even on exposure to a pH beyond natural variability (pH 7.3). Notably, F2M reproductive resilience was also found in the next generations (F3M, F4M and F5M) (Supplementary Fig. 2). In other words, the effects of elevated CO2 disappeared after three generations of OA exposure, even though the reproduction of P. nana was highly sensitive to elevated CO2 initially. These findings were further supported by laboratory microscale experiments using field-collected copepods (Supplementary Fig. 3 and Supplementary Discussion 2). Taken together, our results implied that multigenerational plasticity could enable copepods to better cope with long-lasting elevated CO2.
a, Developmental time was not significantly affected by elevated CO2 (n = 10–13 per treatment). N–C, nauplius to copepodid. N–A, nauplius to adult. b, Body length was not significantly affected by elevated CO2 (n = 3–5 per treatment). c, Fecundity was significantly decreased by elevated CO2, but the negative effect was alleviated by the F2 generation (F0_7.7: P = 0.01, F0_7.3: P = 0.02, F1_7.3: P = 0.043, n = 3–5 per treatment). d, Sex ratio (female/total) was significantly decreased by elevated CO2, but the negative effects were alleviated in the third generation (F0_7.3: P = 0.016, F1_7.3: P = 0.016, n = 10–13 per treatment). Significant differences were analysed using Student’s paired t test followed by Tukey’s test. Asterisks denote statistically significant differences from the control group. Data are presented as the mean ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001.
Genome-wide and OA-specific DNA methylomes
As reported in previous studies29,30, DNA methylation might be an adaptive strategy that allows some arthropod species to respond quickly to environmental stress. We focused on how copepods can achieve evolutionary success in acidified oceans by assessing methylome changes. The P. nana de novo genome was sequenced and assembled (NCBI accession number JAGUCH000000000) to facilitate the additional assembly and annotation of methylome (multigenerational (F1M, F2M) and transgenerational (F1T, F2T) groups) and transcriptome datasets obtained from the F0 to F2 inbred lines at the control pH and OA stress conditions (Supplementary Fig. 1, Supplementary Tables 2–4, Supplementary Discussion 1 and Supplementary Dataset 1). To identify the correlation between the reproductive resilience observed in copepods exposed to elevated CO2 over three consecutive generations and relevant DNA methylation variations, we have tracked epigenetic changes of female copepods from multigenerational groups (F0N_8.0: 9.7%; F0M_7.3: 9.8%; F1M_7.3: 10.2%; and F2M_7.3: 10.8% of all CpG; ~30–35×, mean coverage 32.9×; Extended Data Fig. 2, Supplementary Dataset 2a–c and Supplementary Discussion 3). Thus, we focused on genes relating to fertility of female copepods, which are key to reproductive success (Methods). Considering that most arthropods have lower global DNA methylation and lost transposable element (TE) methylation, it is likely that the methylation pattern in P. nana has evolved independently in arthropod lineages, as observed in centipedes and mealy bugs31. Notably, most DNA methylation in P. nana occurred in both intergenic and gene regions, contrasting with previous observations that DNA methylation in invertebrates generally occurs predominantly in gene bodies rather than intergenic regions13,32,33,34,35.
Genotype-independent adjustments of methylation levels
On the basis of the evidence that methylation patterns are significantly affected by genome structure36, we further investigated whether P. nana’s unique DNA methylation patterns depend on the presence of adaptive genetic variant. We used a circos plot to obtain a genome-wide view of P. nana’s differentially methylated regions (DMRs) (Fig. 3a). We expected the DNA methylation variation to be determined by genetic changes if the epigenotype were genotype dependent. Our data indicated, however, that intergenerational DNA methylation changes were not affected by genetic changes (Mantel test: r = −0.4286, P = 0.583; Kendall tau rank correlation coefficient; F0N: τ = −0.016, P = 0.005; F0M: τ = −0.016, P = 0.007; F1M: τ = −0.015, P = 0.009; F2M: τ = −0.014, P = 0.015; Supplementary Fig. 4). We also tested whether elevated CO2 could change methylated cytosine levels in the CpG islands. Epigenome-wide methylated CpG screening showed no correlation with genetic diversity as estimated by single-nucleotide polymorphisms (SNPs). This is further supported by a fixation index (FST) comparison with OA-induced DNA methylation changes (Fig. 3b and Supplementary Fig. 5). Differences in methylation levels between parents and offspring (F0–F1 and F1–F2) were greater as the number of genic SNPs increased, but there was no correlation between the epigenetic inheritance factor and genic FST (r = 0.02 and −0.02) or exonic FST (r = 0.02 and −0.04). Thus, the genome-wide DNA methylation variants were somewhat correlated with the genotype, but the association between gene-body methylation (GBM) and genotype at particular loci is weak. This result indicated that some portion of genetically independent DNA methylation variation may contribute to phenotypic plasticity in P. nana. In addition, linear mixed-effect models support our results that the contribution of epigenomic variation to phenotypic plasticity in multigenerational traits is irrelevant to adaptive genetic variation (intra-class correlation (ICC) = 0.013, ICC2 = 99.9%; 4.948 (F0M; P < 2 × 10–16), 5.144 (F1M; P < 2 × 10–16) and 5.192 (F2M; P < 2 × 10–16); Supplementary Table 5). The observed genetically independent epigenetic variation was further evident in the interplay between DNA methylation and TEs (11.7% of the annotated TEs; Extended Data Fig. 3a,b, Supplementary Dataset 2b and Supplementary Discussion 3).
a, Circos plot showing Paracyclopina nana genome-wide DNA methylation profile (Supplementary Methods). b, Epigenetic variants associated with multigenerational resilience are independent of genetic variants. Genes with higher methylation in the offspring than in the parents are indicated in red, whereas blue indicates the opposite. The epigenetic inheritance factor (along the y axis) is contrasted with genotypic factors (along the x axis). The Pearson correlation coefficients (r) show that intergenerational epigenetic variation correlates weakly with adaptive genotype factors (0 < |r| < 0.2). mCG denotes a methylation at CpG sites. vsF0N indicates DMRs normalized to the F0N group. Δmeth denotes F1M–F0M and F2M–F1M, respectively.
Correlation between DNA methylation and reproductive fitness
We exposed the copepods to the DNA methylation inhibitor 5-azacytidine (5-Aza) for three generations to elucidate a clear cause–effect relationship between the observed DNA methylation changes and the acquired multigenerational reproductive resilience. We hypothesized that fecundity would not recover in F2M if DNA methylation was inhibited. To minimize the effects of 5-Aza on the whole-body epigenome profile, we performed acute 5-Aza toxicity tests at concentrations of 0.5 mg l–1 and 1 mg l–1 (Fig. 4a and Supplementary Fig. 6). As expected, fecundity did not recover in F2M with 5-Aza treatment. In groups treated with 1 mg l–1 5-Aza, we observed a constant decrease in fertility at pH 8.0 (control), indicating long-term toxic effects. Interestingly, we did not observe multigenerational buffering in the 0.5 mg l–1 5-Aza group, suggesting that DNA methylation correlates strongly with reproductive fitness. It should also be noted that 5-Aza-induced methyltransferase-inhibiting effects reduced fertility in the F1 pH 7.7 group. This group fully recovered within a generation of exposure to elevated CO2, but fecundity was decreased after 5-Aza exposure. These results suggest that epigenetic mechanisms could enhance the reproductive resilience of copepods in response to environmental changes. To further investigate the correlation between F2M phenotypic plasticity and 5-Aza exposure, we assessed global DNA methylation using immunohistochemistry with an anti-5-methylcytosine (anti-5-mC) antibody (Fig. 4b and Extended Data Fig. 4). Quantitative immunohistochemistry staining revealed DNA was less methylated in the 5-Aza F2M group, indicating that 5-Aza prevented stable DNA methylation inheritance. This result indicates that epigenetic information inherited from parental generations affects copepods’ future responsiveness to elevated CO2 for multiple generations.
a, In the F2M group continuously exposed to 5-Aza, fecundity does not recover. Significant differences were analysed using one-way ANOVA followed by Tukey’s test. Asterisks denote statistical significance relative to the control groups (pH 8.0). Data are presented as the mean ± s.e.m. (*P < 0.05; **P < 0.01; ***P < 0.001). (F0: P = 0.003 (Cont_7.7), P < 0.001 (Cont_7.3), P = 0.035 (0.5 mg l–1_7.3), P = 0.023 (1 mg l–1_7.7), P = 0.011 (1 mg l–1_7.3); F1: P = 0.036 (Cont_7.7), P = 0.01 (Cont_7.3), P = 0.012 (0.5 mg l–1_7.7), P = 0.007 (0.5 mg l–1_7.3), P = 0.005 (1 mg l–1_7.7), P = 0.005 (1 mg l–1_7.3); F2: P = 0.001 (0.5 mg l–1_7.7), P = 0.001 (0.5 mg l–1_7.3), P = 0.018 (1 mg l–1_7.7), P = 0.017 (1 mg l–1_7.3); n = 3 per treatment). b, Methylated DNA localization within Paracyclopina nana in the absence or presence of 0.5 mg l–1 5-Aza. The small white square details the extent and pattern of DNA methylation. Compared with the untreated groups, the groups treated with 5-Aza showed no significant changes in DNA methylation levels.
Effect of elevated CO2 on epigenotypes
Given our in vivo results, we investigated whether environmentally induced DNA methylation changes can mediate multigenerational plasticity in response to parental OA stress. On average, all samples’ genome-wide methylation levels were high in both genic and intergenic regions, indicating that the overall methylation pattern did not change in response to environmental cues (exon: 32.46% versus 32.29, t-test P < 2.2 × 10–16; intron: 27.70% versus 27.58%, t-test P = 4.075 × 10–12; intergenic: 57.04% versus 56.91%, t-test P < 2.2 × 10–16; Fig. 5a). We then focused our study on changes in DNA methylation at genic region (for example, GBM), as GBM is the most common form of CpG methylation in invertebrate genomes13,14,33,34, and it is an important adaptive mechanism that allows transcriptional flexibility in response to environmental stress37. Consistent with this, our data indicated that only GBM had a significant correlation with gene expression, unlike DNA methylation of other genomic regions (for example, promoter regions), implying the relationship between GBM and gene expression is clear in copepods but the correlation is still weak, as shown that the link between these two factors is complex in some invertebrate species12,37 (Supplementary Fig. 4). The number of DMRs (t tests corrected P < 0.05) was much higher in F2M (337 genes) than in F0M and F1M (286 and 326 genes, respectively) (Supplementary Dataset 3a–c). Interestingly, GBM changes associated with multigenerational pH stress exposure resistance were opposite in hyper- and hypomethylated genes (F0M: P = 4.466 × 10–6; F1M: P = 0.8753; F2M: P = 0.01463; hypermethylated genes: 175, 163 and 146 in F0M, F1M and F2M, respectively; hypomethaylted genes: 111, 163 and 192 in F0M, F1M and F2M, respectively; Fig. 5b top). In addition, DNA methylation changes correlated positively from F0 to F2, and the linear regression results indicate a significant association between maternal and offspring DNA methylation (F0 versus F1: P = 2.076 × 10–15; F1 versus F2: P = 4.581 × 10–6; F0 versus F2: P = 0.6417; Fig. 5b bottom). Finally, we performed comparative analysis in multigenerational (F0M, F1M and F2M) and transgenerational (F1T and F2T) groups to distinguish epigenetic changes from environmentally driven patterns (Extended Data Fig. 5a,b and Supplementary Dataset 4). Thus, our data suggest that multigenerational OA exposure is associated with an overall increase in the number of DMGs and that environmental changes experienced by parental generations lead to changes in DNA methylation landscapes that are transmitted to multiple subsequent generations (Supplementary Dataset 5). Specifically, we further evaluated DMGs for the F2M generation’s reproductive fitness. We hypothesized that DMGs commonly found in the three groups (F0M, F1M and F2M) as well as those found only in F2M would be closely related to reproductive resilience (Fig. 5c,d, Extended Data Fig. 6, Supplementary Datasets 6 and 7a–c and Supplementary Discussion 4). The differential methylation of these genes consistently observed in the F2M groups was related to their resistance to elevated CO2. Taken together, our data suggested that epigenetic responses acquired from the parental generation and independent DMG changes in the F2M group mediate phenotypic plasticity in P. nana.
a, Increase in methylation levels was observed at similar rates in the intergenic and genic regions. b, Log2 fold changes in GBMs in response to elevated CO2 plotted against whole-genome bisulfite sequencing for each gene. The density plots of significantly hyper- (red) and hypo- (blue) methylated genes (top). Regression analysis between GBMs over three generations compared with the F0N (bottom). c, Venn diagram of hyper- and hypomethylated genes shared among F0M, F1M and F2M. d, Heatmap representation of the canonical pathways significantly affected by elevated CO2 for three generations, as shown by IPA.
Interplay between DMGs and DEGs in copepod’s plasticity
We used Ingenuity Pathway Analysis (IPA), a web-based bioinformatic tool that visually expresses biological functions, pathways and molecular networks, to examine system-wide multigenerational OA effects based on the combined differentially methylated gene (DMG) and differentially expressed gene (DEG) sets (Supplementary Dataset 8a–c). On the basis of the fact that hyper and hypomethylated genes within the gene-body region are indicators of gene expression38, we found that 187 of the three generations’ 504 significant DMGs overlapped with DEGs (Supplementary Dataset 9). Considering the weak correlation between DNA methylation and gene expression in our preceding analyses, our results showed an unexpectedly high number of combined DEGs and DMGs (DEMGs), and the dataset consisted of several significantly enriched reproduction-related pathways, including the oocyte meiosis, phagosome and mTOR signalling pathways (paired t test: DEMG versus DEG, P = 0.005; DEMG versus DMG, P = 0.012; Supplementary Dataset 10 and Supplementary Fig. 7). Interestingly, despite DEG stabilization, our data show that the F2M generation had the highest number of both DEGs and DMGs (Extended Data Fig. 7 and Supplementary Dataset 9). Similarly, changes in DNA methylation and the regulations of related genes are also essential for the rapid environmental adaptation of invasive species39. This suggests that maternal conditions can affect DNA methylation patterns in the offspring, and the DNA methylation modulation associated with functional gene expression may serve as an important driver of copepods’ plasticity. Consistent with methylation’s role as an expression regulator, our analysis provides evidence that GBM strongly increases with gene expression (Extended Data Fig. 8a). Furthermore, our study demonstrated that GBM is involved in the inhibition of transcriptional noise that can produce abnormal proteins, as commonly observed in mammals and other marine invertebrates (for example, corals and oysters) (Extended Data Fig. 8b). Network analyses revealed significant interaction and crosstalk between F2 DEMGs related to nervous system functions, including genes associated with cholinergic and glutamatergic synaptic proteins, the calcium signalling pathway, mitogen-activated protein kinase and cellular stress (Fig. 6a and Supplementary Dataset 11). Notably, we found that multigenerational exposure to elevated CO2 leads to several DMGs and DEGs involved in the metabolic process, although most differentially expressed transcripts were found at lower levels. Similar to results reported in other invertebrates, our results suggest that long-term acclimation to chronic pH stress substantially increases the energy required to maintain cellular homeostasis40,41. Therefore, environmental stress memory enabled the F2M generation to respond more efficiently to OA stress than previous generations. This postulation is supported by the observed upregulation of multiple metabolic pathways in the F2M generation compared with F0M, including triacylglycerol biosynthesis, the tricarboxylic acid cycle and glycolysis (Supplementary Fig. 8). Indeed, these DEMG results were supported by the feeding behaviour (Supplementary Fig. 9). The feeding performance in the F2M group was much higher than seen in the F2N and F2T groups. These results demonstrated that under multigenerational exposure to elevated CO2, P. nana exhibits plasticity while maintaining stable reproductive capacity through increased energy intake42. Our IPA analysis also demonstrated that these gene clusters are rich in signature pathways, including calcium, gonadotropin (GnRH) and sirtuin signalling (Fig. 6b and Supplementary Discussion 5). In arthropods, corazonin receptors (Crz) are highly conserved neurohormones belonging to the GnRH superfamily that are involved in a wide spectrum of metabolic activities, stress reactions, reproduction regulation and female fertility43. Thus, our genetic network analysis highlights the close interaction between gene expression and epigenetics in response to transcriptional needs caused by multigenerational plasticity. Moreover, to understand the possible role of other epigenetic mechanisms in the multigenerational plasticity, transcriptional changes of histone modification genes were further analysed (Supplementary Fig. 10 and Supplementary Discussion 6). These results suggest DNA methylation and histone modification partially regulate multigenerational resilience in reproductive capacity. Taken together, these complementary DEMGs suggest (1) that epigenetic changes may contribute to resilience in response to pH stress and (2) that intracellular Ca2+ signalling cascade regulation by observed epigenetic memory is key for microscopic organism plasticity to constant environmental stress conditions.
a, Network diagram representing the relevant interaction DEMGs: organism injury and abnormalities, endocrine system disorders, neurological disease, reproductive system disease and metabolic disease. The hexagon indicates the lack of correlation between DNA methylation and gene expression changes in the F0M generation. The colour intensity indicates the degree of DEMG upregulation (red) and downregulation (blue), with unaffected genes in grey (P < 0.05). CP, canonical pathway. Analysis and illustration conducted using IPA software. b, Schematic diagram of the observed DEMGs (Supplementary Discussion 5).
Conclusions
The methylation pattern found in the P. nana genome is different from that in other invertebrates, and epigenetic changes induced by elevated CO2 were related to phenotypic plasticity. These changes may offer resilience to the offspring in response to elevated CO2. The CO2-induced changes in DNA methylation patterns were clearly evident, including genes that play key roles in reproduction-related pathways. These adaptive features of plasticity offer stability to the zooplankton community and therefore to higher trophic levels or feedback loops44. The results of our study on epigenetic plasticity and multigenerational population fitness demonstrated the adaptability of marine organisms to OA. Previous studies on the adaptive mechanisms of copepods have focused primarily on population size and genetic diversity19. However, failure to consider phenotypic plasticity could result in overestimation of the future population’s vulnerability, particularly the decline in diversity caused by environmental disturbances. In fact, in contrast to previous meta-analyses that classified copepods as vulnerable to acidification5, our results indicated that P. nana can acclimate to multigenerational exposure to elevated CO2 through epigenetic plasticity in the absence of genetic diversity. Thus, our findings suggest that epigenetic plasticity could potentially compensate for copepod population losses as a rapid response to changing environmental conditions. Taken together, understanding the responses and adaptability of copepods to OA not only is important for assessing the impact on biodiversity, biogeochemical cycles and trophodynamics in the marine environment, but also provides critical insights into marine ecology.
Methods
Experimental set-up
To minimize both genetic variation and possible past pH disturbance, we selected females from a line of P. nana inbred since the year 2000. We then hatched copepod nauplii for another three generations at pH 8.0. The final offspring generation was raised at pH 8.0 (F0N), pH 7.7 or pH 7.3 (F0M). Nauplii grown in the pH 8.0 control condition were raised under identical conditions for another two generations (F1N and F2N). The nauplii from the OA-exposed groups (pH 7.3 and 7.7) were divided into two groups. One group was returned to the control condition (pH 8.0) and reared for another two generations (transgenerational groups F1T and F2T), and the other group was reared in the low-pH conditions (pH 7.3 and 7.7) for the two subsequent generations (multigenerational group F1M and F2M). To measure elevated CO2 effects, copepods were exposed to three pH conditions (Supplementary Table 6), and several in vivo endpoints were determined. A pH of 8.0 was selected as the control, pH 7.7 was projected as the end-of-the-century \(p_{{\mathrm{CO}}_2}\) level and pH 7.3 was chosen as an extreme ocean acidification scenario to maximize the elevated CO2 effects (as predicted for 2300 under RCP 8.5)45,46. Over the past year, the pH in the brackish water of Lake Hwajinpo ranged from 7.4 to 8.9 (National Institute of Environmental Research in South Korea). The pH trend of brackish lagoon can be obscured by the annual volatility of the pH amplitude. RCP 8.5 modelling predicts that the already very low blackish water area (≤pH 7.4) will drop by up to 0.1 pH units due to the strong buffering capacity of seawater (RCP 8.5 scenario47). The challenge presented by OA involves lifetime exposure to low pH (for example, throughout the life cycle covering reproductive, developmental and growth phases) rather than short-term stress. Thus, epigenetic, molecular and phenotypic changes were studied on exposure to pH 7.3. In addition, to understand copepods’ response to lower pH related to the interannual variability of pH amplitudes, further in vivo experiments of the pH 7.0 group were performed (Supplementary Dataset 12). Stable pH conditions were established in each treatment by bubbling CO2 in seawater coupled with an independent real-time pH negative feedback system (iks aquastar; iks Computer Systeme GmbH). The pH was measured using a pH/ATC electrode (#300729.1, Denver Instruments) connected to a pH–ISE meter (Model 225, Denver Instruments). All P. nana copepods used in the experiment had been grown in an ambient pH of 8.0 for 3 years before this \(p_{{\mathrm{CO}}_2}\) acclimation treatment. In this experiment, P. nana acclimated to the ambient pH condition were transferred to 20 ml vials with the three different pH conditions (pH 8.0, 7.7 and 7.3) for the in vivo experiments and to 120 ml glass jars for the transcriptome and whole-genome bisulfite sequencing (WGBS) analyses. To identify the correlation between the observed reproductive resilience in response to elevated CO2 over three consecutive generations and relevant DNA methylation variations, WGBS was performed after confirming the expected reproductive phenotypes in vivo from additional replicate batches. The same procedure was applied for the transcriptome analysis. Note that, despite the fact that all the experiments were conducted in triplicate, the number of biological samples was 10–15 (specimens per treatment group) × 3 (batches) = 30–45 (total specimens), with a minimum of nine replicates for determination of the same in vivo endpoint (three biological replicates in each of three batches were processed as independently performed experiments). Furthermore, the reproductive endpoints of five copepods were re-analysed for each of the three replicates for WGBS and transcriptomic analysis (n = 15 copepods per group). Each container had its water changed daily. To avoid pH variation from gas exchange, all containers (tanks, beakers, jars and vials) were tightly capped and sealed with parafilm. Total alkalinity was measured in triplicate via Gran-titration48 to determine the pH corresponding to the predicted \(p_{{\mathrm{CO}}_2}\) level. After titration, carbonate system dynamics were calculated using the CO2 SYS programme on the basis of the \(p_{{\mathrm{CO}}_2}\) and total alkalinity measurements49,50. Other environmental conditions (dissolved oxygen, salinity and temperature) were strictly monitored and controlled to minimize differences between the pH treatment groups.
In vivo endpoints
To investigate the copepods’ multigenerational plasticity under pH stress, we performed multiple in vivo experiments over several copepod generations (F0–F2). We evaluated developmental time, body length (female), fecundity and sex ratio endpoints at each pH level (pH 8.0, 7.7 and 7.3). The in vivo endpoints were measured by dividing the copepods into three groups: normal (copepods reared under ambient CO2 conditions whose parents were also reared under ambient CO2 conditions), transgenerational (copepods reared under elevated CO2 conditions whose parents were reared under ambient CO2 conditions) and chronic exposure (copepods reared under elevated CO2 conditions whose parents were also reared under the same level of elevated CO2 conditions). To eliminate possible genetic variation due to life history and epigenome, the nauplii used in these experiments were obtained from an inbred line of females with minimal genetic variation, and all in vivo endpoints were obtained from the same batch (10–15 individuals per replicate and a minimum of three replicate containers per treatment).
To examine potential developmental retardation, ovigerous females acclimated to pH 8.0 pre-exposure were collected and incubated in artificial seawater for 2 hours. Newborn nauplii were collected and transferred to 20 ml vials containing seawater at different pH values. Using a stereomicroscope (SZX-ILLK200, Olympus Corporation), we monitored each copepod group’s developmental stages daily until nauplii became adults (2 weeks). Once all nauplii had matured, we continued to measure ovigerous female body length and fecundity. To measure copepod body length, we selected ten male and ten female individuals from the adults used in the experiment (n ≥ 30). We measured cephalothorax length (the first segment from the top of the head to the posterior end) as a body size indicator. To observe the OA effect on fecundity over three generations, ten ovigerous females selected from the remaining adults were transferred to new individual 20 ml vials at each selected pH. The newly born nauplii were counted every 24 hours. Because the sex ratio was not determined until the copepods matured, newborn nauplii were exposed to elevated CO2 (7.7 and 7.3) until maturation. After they metamorphosed into adults, each group’s sex ratio was determined on the basis of antennule appearance—a male characteristic—under the stereomicroscope51. During the experiments, P. nana were cultured in the experimental conditioned seawater with the microalga T. suecica as a dietary source (approximately 6 × 104 cells ml–1) at room temperature (25 °C). All experiments were conducted in triplicate (biological replicates). The same experimental procedure was applied to the F1 and F2 generations.
To measure the feeding behaviour of copepods with multigenerational OA exposure, we measured the food concentration in the gut content of the copepods using the process described in ref. 52 with minor modifications. Briefly, feeding behaviour is represented as the rate of increase in gut content by monitoring chlorophyll and phaeopigment content when copepods with empty guts are given food. Depending on body size, 3–15 copepods that had been starved for 8 hours were placed in each 1 l beaker (three replicates in control, transgenerational and multigenerational groups). The same amount of phytoplankton was then added to each beaker and gently mixed to ensure the equal distribution of cells. The copepods were taken from one of the beakers at 5–10 min intervals for 120 min, and their intestinal contents were analysed. Next, the copepods were homogenized in 90% aqueous acetone after being rinsed with filtered seawater, and the fluorescent intensity was measured before and after acidification with 10% HCl using Ultrospec 2100 Pro UV/Visible spectrophotometer (Amersham Biosciences). The chlorophyll and phaeopigment content of each copepod were calculated as the slopes of a linear regression on the weight of pigment per time (the rate of gut filling) divided by the weight of pigment per volume (the food concentration)53.
where K is the calibration constant, f0 and fa are the fluorescence intensity before and after acidification, n is the number of copepods and R is the acidification ratio.
WGBS
To investigate molecular mechanisms associated with reproductive resilience, we prepared four biological replicates per treatment (each replicate containing approximately 30 adult female copepods). It is noted that the maternal parent in some species can provide a unique microenvironment (for example, brooding) for its offspring, thereby contributing to >50% heritable variance54, and the correlation between maternal and offspring phenotypes is typically greater than that between paternal and offspring phenotypes55. To highlight the role of maternal effects in relation to reproductive success, we characterized the epigenome and transcriptome of female copepods. We analysed the information on methylated nucleotides using WGBS. The quality and quantity of the genomic DNA extracted from animals in each treatment (F0N, F0M, F1M and F2M) were verified via agarose gel electrophoresis. The quality-controlled genomic DNA was used to construct bisulfite conversion libraries using the Accel Methyl-Seq DNA library. All libraries’ quality and quantity were assessed using an Agilent Technologies 2100 Bioanalyzer. The qualified libraries were sequenced using an Illumina Novaseq 6000 (Illumina) to produce paired-end data in the fastq format.
To investigate whether the observed epigenetic effects can be transmitted across multiple generations and whether they can be representative of the epigenetic changes in response to the treatments with elevated CO2, we performed further WGBS experiments on subsequent generations (F3N, F3M and F4M) and the transgenerational groups (F1T and F2T) (Supplementary Dataset 13). To obtain information on epigenetic changes in multiple generations, F3M’s and F4M’s GBM data were analysed using discriminant analysis of principal components (DAPC) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways (Supplementary 5). Compared multigenerational groups with transgenerational groups, linear regression analysis and hierarchical clustering analysis were performed (Supplementary Dataset 4) (detailed methods description follows). Although it is noted that removal of batch effect may conceal some genuine biological differences56, removal of batch effect can increase the power of overall pattern analysis. As such, batch effects were removed by ComBat method correction of sva package in R57 in this study.
Methylation site identification
A total of 694,213,062 WGBS reads were obtained across 16 samples. All raw WGBS data were trimmed using cutadapt v.1.8.1 to remove adaptors58, low-quality (–q 20) and short (–m 30) reads. The trimmed data were mapped to the reference de novo genome with adjusted parameters (–score_min L,0,−0.6) and deduplicated, after which the methylation score and coverage per nucleotide were obtained by Bismark v.0.22.359. Using the resulting Bismark coverage and summary file, mapping information, methylation percentage and other information were extracted using a custom code. The methylated site tables were annotated using custom Python code with gff3 files from the de novo genome annotation and repeat element analyses. On the basis of the genome annotation, all DNA methylation sites were classified as gene bodies (exon and introns), promoters (up to <1 kb from genes), repeat elements or intergenic regions (regions excluding those defined as gene body or promoter).
Global methylation analysis
The global methylation of different samples was analysed via a principal component analysis to assess the relationship between replicates and samples as well as to identify abnormal values among replicate samples. Normality testing proceeded using the Shapiro–Wilk test. To assess sample differences accurately, we analysed the correlation between samples using the Kendall ranked coefficients approach. In this analysis, abnormal replicates were excluded from all samples. We calculated the methylations’ total genomic context for their positions (exon, intron, intergenic or promoters) using gff3 annotated tables that merged all sample methylation data.
Assessing the relationship between genotype and epigenotype
To exclude SNP effects from our methylation analysis and assess sample genotype methylation, we mapped SNPs and called on the whole genome and gene bodies. SNPs with methylation information were called using Bis-SNP v.1.0.1 on the basis of the reference genome, and deduplicated BAM files were created via Bismark methylation sequence mapping60. In this way, variations in observed methylation patterns were not confused with C–T DNA polymorphisms. From the Bis-SNP result tables, we extracted site information as heterogeneous (0/1), homozygous non-reference (1/1 or larger number than 1/1) or homozygous reference (0/0)60. We filtered the data with a 10–100 SNP position coverage for every sample. The relationship between number of SNPs and number of methylations or inter-origin absolute difference methylation levels of gene bodies was analysed and visualized using custom Python code.
Genetic heterogeneity was measured using QST statistics, which is appropriate for large measured populations (>10), small mean FST and strong selection61. For QST computation, we first calculated the FST value on the basis of the Hudson estimator62, after which QST estimates were calculated using the formula QST = 2 FST /(1+ FST)63.
Differential methylation analysis
Positional methylation levels ranged from 0 (no methylation) to 1 (all methylation) for individual methylation calling coverage files. For gene-level analysis, each position was annotated as an exon, intron or intergenic region in the gff3 annotation file, and the median methylation level for each gene position was calculated.
To identify differential methylation in a gene, we set F0N as the control condition for generalized linear model calculation in R with the MethylKit package64. The design model for the generalized linear model was glm(methylated, non-methylated ~ generation * position, family = ‘binomial’). In this model, ‘methylated, non-methylated’ means that the response variable represents the number of methylated and non-methylated reads, and ‘generation’ and ‘position’ are variables for pH, generation conditions and genomic coordinates. We calculated the final methylation value by aggregating the replicates for each condition. Genes with <5 methylated positions were filtered out, and genes with P < 0.05 were considered statistically significant.
Differential gene expression analysis
RNA was extracted in triplicate (three biological replicates for each treatment, 30 individuals per replicate) using the same conditions in which the methylation samples were obtained. RNA was extracted from the homogenized samples using TRIzol reagent (Molecular Research Center). Following the manufacturer’s instructions, the RNA pellet was collected from the TRIzol suspension sample using isopropanol precipitation and 75% ethanol in diethyl pyrocarbonate (DEPC) water followed by DEPC water. We converted the RNA samples to complementary DNA and constructed the sequencing libraries. The library products were used for sequencing on an Illumina Novaseq 6000. The paired-end raw RNA-seq data were trimmed by Trimmomatic v.0.38 to remove adaptors, low-quality and short reads before downstream analyses. To quantify the transcription levels and identify differential gene expression, we analysed the clean RNA-seq data using the Tuxedo protocol65. We mapped the trimmed RNA-seq reads data to the corresponding reference genome using HISAT v.2.2.1 and then sorted and converted the SAM output to the aligned BAM format file using SAMtools v.1.966. Next, we used the alignment files to assemble the transcripts and estimate expression abundances as coverage, FPKM and TPM using StringTie v.2.1.4. Finally, we used the Ballgown R package to calculate the differential expression between two or more experiments from the StringTie results. DEGs were identified as genes with P < 0.05 and an absolute log fold change (|logFC|) > 1.
Transcription validation
The cleaned RNA-seq data for each sample were mapped to the reference assembly using HISAT2 v.2.2.1. With the annotation file (gff or bed format) in hand, each gene and exon’s mean coverage was calculated from the mapping file using BedTools v.2.1.0 and a Python script. To investigate the gene-body methylation effect on transcription, these coverage data were calculated using the gene methylation data. To evaluate the genes’ and exons’ transcriptional noise, we obtained the coverage per normalized exon on the basis of six replicates and computed the average coverage ratio from exons 2 to 6 versus exon 1 in unmethylated, methylated and densely methylated (>80% methylation) groups.
DAPC
To investigate epigenetic and transcriptional variation on multigenerational OA exposure, we conducted a DAPC analysis, a multivariate statistical approach used to identify group differences67. In the DAPC, we transformed the data using a principal component analysis and divided the groups into two clusters. Variation between groups and within groups along with the criteria required to maximize between-group variation were used to discriminate the clusters.
Differential methylation and gene expression comparison
We performed an integrated analysis of DMRs and DEGs under three conditions (F0_DMG-F0_DEG, F1_DMG-F1_DEG and F2_DMG-F2_DEG) to investigate DNA methylation and gene expression association. The gene-body methylation ratio and gene expression fold change were compared for each gene under the same conditions and were visualized using scatterplots. To acquire detailed gene and pathway information for the methylated genes and their expression, we used gene ontology functional annotation. Because P. nana is a non-model organism and the gene ontology terms have many limitations, we also used an NCBI BLAST analysis with the Swiss-prot database to obtain exact results that met the specific e-value parameter (e-value <1 × 10–10). We obtained Uniprot accession numbers for the genes from DAVID v.6.8 for the functional annotation68.
To reveal the DMG and DEG relationship, we performed correlation and pathway analysis using IPA (Ingenuity Systems) web-based software for calculating and visualizing the biological functions of genes and molecules. We collected the combined DEMG set before the core analysis. The gene and biological function network was expanded from the DEMG data on the basis of gene expression, methylation data and Ingenuity knowledge base. The network and canonical pathways with P < 0.05 were considered significant.
DNA methylation change validation using 5-Aza exposure experiment
To assess the relationship between DNA methylation and reproductive resilience, we inhibited DNA methylation using 5-Aza (Sigma-Aldrich) and conducted fecundity tests. First, we evaluated the no-observed-effect concentration and half lethal concentration (LC50) at 48 and 96 hours to assess 5-Aza acute toxicity. No food was supplied during the experiments. Half of the medium was renewed 48 hours after exposure. Mortality was measured using a stereomicroscope (SZX-ILLK200, Olympus Corporation) every 24 hours. A copepod was considered dead when it showed no sign of movement. Finally, no-observed-effect concentration, LC10 and LC50 values were calculated using a Probit analysis (ToxRat v.2.09, GmbH, 2005). Fecundity was measured in P. nana exposed to 0, 0.5 and 1.0 mg l–1 5-Aza.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Raw data and de novo assembly of P. nana genome can be found in NCBI BioProject PRJNA714788. The RNA sequence data used in this study are accessible from NCBI under BioProject number PRJNA714790, and whole-genome bisulfite sequencing (WGBS) data are available from PRJNA714791. Individual SRA accessions of transcriptome and WGBS data are listed in Supplementary Datasets 2 and 13. Source data are provided with this paper.
Code availability
Analysis of methylation data used the scripts at https://github.com/lyijin/working_with_dna_meth (ref. 41). Other analytical and plotting scripts are available at https://doi.org/10.5281/zenodo.6830436 (v.1.0.0)69.
References
Cerrano, C. et al. Red coral extinction risk enhanced by ocean acidification. Sci. Rep. 3, 1457 (2013).
Munday, P. L. et al. Ocean acidification impairs olfactory discrimination and homing ability of a marine fish. Proc. Natl Acad. Sci. USA 106, 1848–1852 (2009).
Lord, J. P., Harper, E. M. & Barry, J. P. Ocean acidification may alter predator–prey relationships and weaken nonlethal interactions between gastropods and crabs. Mar. Ecol. Prog. Ser. 616, 83–94 (2019).
Gibbin, E. M., N’Siala, G. M., Chakravarti, L. J., Jarrold, M. D. & Calosi, P. The evolution of phenotypic plasticity under global change. Sci. Rep. 7, 17253 (2017).
Vagas, C. A. et al. Species-specific responses to ocean acidification should account for local adaptation and adaptive plasticity. Nat. Ecol. Evol. 1, 0084 (2017).
Nunney, L. Adapting to a changing environment: modeling the interaction of directional selection and plasticity. J. Hered. 107, 15–24 (2016).
Vehmaa, A., Brutemark, A. & Engström-Öst, J. Maternal effects may act as an adaptation mechanism for copepods facing pH and temperature changes. PLoS ONE 7, e48538 (2012).
Isari, J., Zervoudaki, S., Saiz, E., Pelejero, C. & Peters, J. Copepod vital rates under CO2-induced acidification: a calanoid species and a cyclopoid species under short-term exposures. J. Plankton Res. 37, 912–922 (2015).
Kenkel, C. D. & Matz, M. V. Gene expression plasticity as a mechanism of coral adaptation to a variable environment. Nat. Ecol. Evol. 1, 0014 (2016).
Mallard, F., Nolte, V. & Schlötterer, S. The evolution of phenotypic plasticity in response to temperature stress. Genome Biol. Evol. 12, 2429–2440 (2020).
Torda, G. et al. Rapid adaptive responses to climate change in corals. Nat. Clim. Change 7, 627–636 (2017).
Chandra Rajan, K., Meng, Y., Yu, Z., Roberts, S. B. & Vengatesen, T. Oyster biomineralization under ocean acidification: from genes to shell. Glob. Change Biol. 27, 3779–3797 (2021).
Liew, Y. J. et al. Epigenome-associated phenotypic acclimatization to ocean acidification in a reef-building coral. Sci. Adv. 4, eaar8028 (2018).
Lim, Y. K. et al. DNA methylation changes in response to ocean acidification at the time of larval metamorphosis in the edible oyster, Crassostrea hongkongensis. Mar. Environ. Res. 163, 105214 (2021).
Schunter, C. et al. Molecular signatures of transgenerational response to ocean acidification in a species of reef fish. Nat. Clim. Change 6, 1014–1018 (2016).
Tsang, H. H., Welch, M. J., Munday, P. L., Ravasi, T. & Schunter, C. Proteomic response to ocean acidification in the brain of juvenile coral reef fish. Front. Mar. Sci. 7, 605 (2020).
Thomas, G. W. C. et al. Gene content evolution in the arthropods. Genome Biol. 21, 15 (2020).
Stern, D. B. & Lee, C. E. Evolutionary origins of genomic adaptations in an invasive copepod. Nat. Ecol. Evol. 4, 1084–1094 (2020).
Dam, HansG. A. et al. Rapid, but limited, zooplankton adaptation to simultaneous warming and acidification. Nat. Clim. Change 11, 780–786 (2021).
Cripps, G., Lindeque, P. & Flynn, K. J. Have we been underestimating the effects of ocean acidification in zooplankton? Glob. Change Biol. 20, 3377–3385 (2014).
Turner, J. T. The importance of small pelagic planktonic copepods and their role in pelagic marine food webs. Zool. Stud. 43, 255–266 (2004).
Lee, K.-W., Park, H. G., Lee, S.-M. & Kang, H.-K. Effects of diets on the growth of the brackish water cyclopoid copepod Paracyclopina nana Smirnov. Aquaculture 256, 346–353 (2006).
Selden, P. A., Huys, R., Stephenson, M. H., Heward, A. P. & Taylor, P. N. Crustaceans from bitumen clast in carboniferous glacial diamictite extend fossil record of copepods. Nat. Commun. 1, 50 (2010).
Thor, P. & Dupont, S. Transgenerational effects alleviate severe fecundity loss during ocean acidification in a ubiquitous planktonic copepod. Glob. Change Biol. 21, 2261–2271 (2015).
Wang, M., Jeong, C.-B., Lee, Y. H. & Lee, J.-S. Effects of ocean acidification on copepods. Aquat. Toxicol. 196, 17–24 (2018).
Santangelo, J. M., Rocha, A. D. M., Bozelli, R. L., Carneiro, L. S. & Esteves, F. D. A. Zooplankton responses to sandbar opening in a tropical eutrophic coastal lagoon. Estuar. Coast. Shelf Sci. 71, 657–668 (2007).
Park, G.-H. et al. Large accumulation of anthropogenic CO2 in the East (Japan) Sea and its significant impact on carbonate chemistry. Glob. Biogeochem. Cycles 20, GB4013 (2006).
Hofmann, G. E. et al. High-frequency dynamics of ocean pH: a multi-ecosystem comparison. PLoS ONE 6, e28983 (2011).
Jeremias, G. et al. Transgenerational inheritance of DNA hypomethylation in Daphnia magna in response to salinity stress. Environ. Sci. Technol. 52, 10114–10123 (2018).
de la Peña, M. V., Piskobulu, V., Murgatroyd, C. & Hager, R. DNA methylation patterns respond to thermal stress in the viviparous cockroach Diploptera punctata. Epigenetics 16, 313–326 (2021).
Lewis, S. H. et al. Widespread conservation and lineage-specific diversification of genome-wide DNA methylation patterns across arthropods. PLoS Genet. 16, e1008864 (2020).
Lim, Y. K., Dang, X. & Thiyagarajan, V. Transgenerational responses to seawater pH in the edible oyster, with implications for the mariculture of the species under future ocean acidification. Sci. Tot. Environ. 782, 146704 (2021).
Dixon, G., Liao, Y., Bay, L. K. & Matz, M. V. Role of gene body methylation in acclimatization and adaptation in a basal metazoan. Proc. Natl Acad. Sci. USA 115, 13342–13346 (2018).
Liew, Y. J. et al. Intergenerational epigenetic inheritance in reef-building corals. Nat. Clim. Change 10, 254–259 (2020).
Zemach, A., McDaniel, I. E., Silva, P. & Zilberman, D. Genome-wide evolutionary analysis of eukaryotic DNA methylation. Science 328, 916–919 (2010).
Cavalli, G. & Heard, E. Advances in epigenetics link genetics to the environment and disease. Nature 571, 489–499 (2019).
Li, Y. et al. DNA methylation regulates transcriptional homeostasis of algal endosymbiosis in the coral model Aiptasia. Sci. Adv. 4, eaat2142 (2018).
Arechederra, M. et al. Hypermethylation of gene body CpG islands predicts high dosage of functional oncogenes in liver cancer. Nat. Commun. 9, 3164 (2018).
Huang, X. et al. Rapid response to changing environments during biological invasions: DNA methylation perspectives. Mol. Ecol. 26, 6621–6633 (2017).
Pan, T.-C. F., Applebaum, S. L. & Manahan, D. T. Experimental ocean acidification alters the allocation of metabolic energy. Proc. Natl Acad. Sci. USA 112, 4696–4701 (2015).
Evans, T. G., Pespeni, M. H., Hofmann, G. E., Palumbi, S. R. & Sanford, E. Transcriptomic responses to seawater acidification among sea urchin populations inhabiting a natural pH mosaic. Mol. Ecol. 26, 2257–2275 (2017).
Pedersen, S. A. et al. Multigenerational exposure to ocean acidification during food limitation reveals consequences for copepod scope for growth and vital rates. Environ. Sci. Technol. 48, 12275–12284 (2014).
Tsuchiya, R. et al. Maternal GABAergic and GnRH/corazonin pathway modulates egg diapause phenotype of the silkworm Bombyx mori. Proc. Natl Acad. Sci. USA 118, e2020028118 (2021).
Taucher, J. et al. Influence of ocean acidification on plankton community structure during a winter-to-summer succession: an imaging approach indicates that copepods can benefit from elevated CO2 via indirect food web effects. PLoS ONE 12, e0169797 (2017).
Bopp, L. R. et al. Multiple stressors of ocean ecosystems in the 21st century: projections with CMIP5 models. Biogeosciences 10, 6225–6245 (2013).
Hartin, C. A., Bond-Lamberty, B., Patel, P. & Mundra, A. Ocean acidification over the next three centuries using a simple global climate carbon-cycle model: projections and sensitivities. Biogeosciences 13, 4329–4342 (2016).
Shukla, P. R. et al. (eds) Special Report on Climate Change and Land (IPCC, 2019).
Gran, G. Determination of the equivalence point in potentiometric titrations. Part II. Analyst 77, 661–671 (1952).
Bagur, R. & Hajnóczky, G. Intracellular Ca2+ sensing: its role in calcium homeostasis and signaling. Mol. Cell 66, 780–788 (2017).
Lewis, E. & Wallace, D. W. R. Program Developed for CO2 System Calculations ORNL/CDIAC-105 (DOE, 1998).
Itô, T. The biology of a harpacticoid copepod, Tigriopus japonicus Mori. J. Fac. Sci. Hokkaido Univ. Ser. 6 17, 474–499 (1970).
Dagg, M. J. A method for the determination of copepod feeding rates during short time intervals. Mar. Biol. 75, 63–67 (1983).
Strickland, J. D. H. & Parsons, T. R. A Practical Handbook of Seawater Analysis (Fisheries Research Board of Canada, 1968).
Moore, C. L. in Evolution and Learning: The Baldwin Effect Reconsidered (eds Weber, B. H. & Depew, D. J.) 115–139 (MIT Press, 2003).
Moore, C. L. in Advances in the Study of Behavior (eds Slater, P. J. B. et al.) 47–118 (Academic Press, 1995).
Goh, W. W. B., Wang, W. & Wong, L. Why batch effects matter in omics data, and how to avoid them. Trends Biotechnol. 35, 498–507 (2017).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for bisulfite-seq applications. Bioinformatics 27, 1571–1572 (2011).
Liu, Y., Siegmund, K. D., Laird, P. W. & Berman, B. P. Bis-SNP: combined DNA methylation and SNP calling for bisulfite-seq data. BMC Genome Biol. 13, R61 (2012).
Whitlock, M. C. & Guillaume, F. Testing for spatially divergent selection: comparing QST to FST. Genetics 183, 1055–1063 (2009).
Bhatia, G., Patterson, N., Sankararaman, S. & Price, A. L. Estimating and interpreting FST: the impact of rare variants. Genome Res. 23, 1514–1521 (2013).
Leinonen, T., McCairns, R. J. S., O’Hara, R. B. & Merilä, J. QST–FST comparisons: evolutionary and ecological insights from genomic heterogeneity. Nat. Rev. Genet. 14, 170–190 (2013).
Akalin, A. et al. methylKit: a comprehensive R package for the analysis of genome-wide DNA methylation profiles. BMC Genome Biol. 13, R87 (2012).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Jombart, T., Devillard, S. & Balloux, F. Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet. 11, 94 (2010).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Lee, Y.H. et al. lsoda/Pnana_Methylation: code for epigenetic plasticity enables copepods to cope with ocean acidification (epigenetics). Zenodo https://zenodo.org/record/6830436 (2022).
Acknowledgements
This work was supported by grants from the Collaborative Genome Program of the Korean Institute of Marine Science and Technology Promotion funded by the Ministry of Oceans and Fisheries (no. 20180430) and from MarineBiotics Project (20210469) of the Ministry of Ocean and Fisheries, Korea to J.-S.L., by a grant from the National Research Foundation of Korea (2021R1A2C100943912) to H.G.P. and by a grant from the National Research Foundation of Korea (2020R11A1A01074607) to Y.H.L.
Author information
Authors and Affiliations
Contributions
Y.H.L., M.-S.K. and J.-S.L. conceived of and designed the project. H.G.P. collected field samples and provided resources. Y.H.L. and M.-S.K. performed experiments and conducted bioinformatics and statistical analyses. M.W., H.G.P., R.K.B. and R.S.-S.W. provided input during data analysis and data interpretation. Y.H.L., M.-S.K. and J.-S.L. analysed the data and wrote the manuscript. J.-S.L. and Y.H.L. supervised the overall project. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Climate Change thanks Xin Lin, Manuel Aranda, Vengatesen Thiyagarajan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The effect of transgenerational exposure on pH stress is not significant for the copepod Paracyclopina nana.
a. Developmental time was not significantly affected by elevated CO2 conditions (n=10-15 per treatment). b. Body length was not significantly affected by elevated CO2 conditions (n=3 per treatment). c. Number of brooding females was significantly lower in response to pH stress compared to the control in F0, but no significant effects of pH stress were detected in the transgenerational exposure groups (F1T and F2T) (F0_7.7: P = 0.01, F0_7.3: P = 0.02, n=3 per treatment). d. The sex ratio returned to control levels in F1T and F2T generations (F0_7.3: P = 0.016, n=10-15 per treatment). e. Body length measurement (n=10, respectively). Significant differences were analyzed using Student’s paired t-test followed by Tukey’s test. Asterisks denote statistical significance relative to the control group (pH 8.0). Data are presented as the mean ± s.e.m. (*P < 0.05, **P < 0.01, and ***P < 0.001).
Extended Data Fig. 2
Epigenetic landscape. a. Number of differentially methylated regions (DMRs) between treatments in three methylation contexts (CpG, CHH, and CHG). ‘Unique’ indicates the number of unique DMRs in each methylation context. ‘Hyper’ and ‘hypo’ indicate higher and lower methylation, respectively, in the elevated CO2 treatment groups compared to the control group. b. Metaplots representing DNA methylation patterns. Density plot analysis of all samples revealed a bimodal DNA methylation distribution. The methylation levels of exons, introns, and intergenic regions were equally high. c. Methylated position distribution across a standardized gene model with 1-kb flanking regions. The red line represents the transcription start and end sites. The black line indicates the widths of the exons and introns’ average normalized lengths (Exon1: 362 bp, Exon2: 402 bp, Exon3: 369 bp, Exon-3: 332 bp, Exon-2: 360 bp, Exon-1: 474 bp, Intron1: 466 bp, Intron2: 362 bp, Intron-2: 292 bp, Intron-1: 332 bp).
Extended Data Fig. 3 DNA methylation proportion of transposable elements (TEs) in elevated CO2 treatment groups versus control groups.
a. Proportion of differential methylated genes in all TEs and gene body. b. Violin plots showing changes in DNA methylation in all TEs, Gypsy, and Pao under elevated CO2 conditions. Proportion of DNA methylation in TEs varies across multigenerational exposure to elevated CO2 concentrations. CpG methylation levels of all TEs in F0M change significantly compared with the control group (F0N) (F0M hyper vs. control P = 4.589e-06, F0M hypo vs. control P = 0.01681) (Left). GYPSY TEs (P = 1.229e-08) and PAO TEs (P>2.2e-16) had increased DNA methylation levels under elevated CO2 conditions (Right) (F2M hyper vs. control P = 0.030, F2M hypo P = 0.047). Significant differences were analyzed using Student’s paired t-test followed by Tukey’s test (*P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001).
Extended Data Fig. 4 Effects of 5-azacytidine (5-Aza) on DNA methylation.
5-Aza treatment blurred DNA methylation as found using mouse anti-5-mC antibody labeling and a fluorescein-5-isothiocyanate anti-goat secondary antibody. This indicated DNA demethylation heterogeneity. Furthermore, the increased F0M to F2M immunoreactivity emphasizes that F2M obtained a permanent genome-wide DNA methylation imprint.
Extended Data Fig. 5 Comparative analysis between the multigenerational and transgenerational groups (F1T and F2T) to distinguish epigenetic changes acquired from the environment.
a. Linear regression analysis of the relationship between the multigenerational and transgenerational groups. The correlation between multigenerational and transgenerational groups was weak, indicating that overall epigenetic pattern is distinguished by the environment. This result probably reflects specific epigenetic alterations in the DNA methylome of the multigenerational groups. b. Functional enrichment analysis of differentially methylated regions normalized to the F0N groups. The hierarchical clustering analysis revealed that differentially methylated genes (DMGs) of the F2M group was more similar to the transgenerational groups than the other multigenerational groups exposed to the same level of CO2, suggesting that the differences observed in resilience-specific methylation resulting from multigenerational exposure to elevated CO2 may not be merely due to chronic exposure to elevated CO2 (P < 0.05).
Extended Data Fig. 6 Identification and functional prediction of differentially methylated genes (DMGs).
a. Gene ontology (GO) enrichment analysis of the DMGs. The x-axis indicates the hypo- and hypermethylated genes, and the y-axis shows the GO terms with significant enrichment of DMGs in Paracyclopina nana exposed to elevated CO2 (P adjusted < 0.05). b. Significantly enriched pathways by gene set enrichment analysis (GSEA) comparing DMGs of F2M versus control groups. Heatmaps for top and bottom 50 F2M DMGs. The red color indicates hypermethylated genes, and the blue color indicates hypomethylated genes. c. The GSEA of the metabolic process and calcium ion transport related pathways in the F2M groups.
Extended Data Fig. 7 Relationship between gene-body methylation (GBM) and gene expression (GE) with multigenerational plasticity in copepods exposed to elevated CO2 levels.
a. GBM. b. GE. Discriminant function density plots with four measurements. The arrow indicates the shift in discriminant function scores for the F2 ocean acidification-treated groups compared to the control.
Extended Data Fig. 8 Positive correlation between gene body methylation (GBM) and expression levels (GE).
a. Genes enriched in GBM show higher GE than unmethylated genes (P < 0.01). b. The linear graph represents the relationship between the inverse of the coefficient of variation (CV, signal to noise ratio) in gene expression and the log10-transformed mean expression values of all samples. The CV of methylated genes is higher than that of unmethylated genes, suggesting that GBM is involved in inhibition of transcriptional noise.
Supplementary information
Supplementary Information
Extended Data Figure legend, Supplementary Tables 1–6, Figs. 1–10, Methods and Discussion.
Supplementary Data 1
Supplementary Datasets 1–131, Functional annotation result of annotated genes in Paracyclopina nana. 2a, Trimming, mapping and coverage calculations for WGBS and RNA-Seq reads. 2b, Differentially methylated repeat elements. 2c, The CpG bias, methylation density and median methylation on a per-gene basis. 3a, Differentially methylated genes at pH 7.3 generation F0 (F0M), relative to control (pH 8.0 generation F0, F0N). 3b, Differentially methylated genes at pH 7.3 generation F1 (F1M), relative to control (pH 8.0 generation F0, F0N). 3c, Differentially methylated genes at pH 7.3 generation F2 (F2M), relative to control (pH 8.0 generation F0, F0N). 4, Transgenerational effect of ocean acidification on the epigenome. 5, The genome-wide DNA methylation profiles in F3M and F4M. 6, Hyper- and hypo-differentially methylated genes list. 7a, Enriched canonical pathways from Ingenuity Pathway Analysis with DEMGs data. 7b, GO enrichment of biological process in F2M hyper- and hypo-methylated genes from DAVID results. 7c, Up- and down-regulated GO terms in F2M versus F2N methylated genes from GSEA (|NES| > 1.5, nominal P < 0.05). 7d, Detailed gene lists of GO term enriched by GSEA in F2M. 8a, Differentially expressed genes at pH 7.3 generation F0 (F0M), relative to control (pH 8.0 generation F0, F0N). 8b, Differentially expressed genes at pH 7.3 generation F1 (F1M), relative to control (pH 8.0 generation F0, F0N). 8c, Differentially expressed genes at pH 7.3 generation F2 (F2M), relative to control (pH 8.0 generation F0, F0N). 9, Differentially expressed and methylated genes (DEMGs). 10, KEGG pathway enrichment analysis of DEGs, DMGs and DEMGs (combined gene set of DEGs and DMGs) and statistics of reproduction-related pathways’ ranks. 11, Ingenuity Pathway Analysis (IPA) network analysis of the combined set of DMGs and DEGs. 12, Effects of exposure to lower pH levels (pH 7.0) on Paracyclopina nana reproduction. 13, Trimming, mapping and coverage calculations for new WGBS.
Source data
Source Data Fig. 2
Statistical Source Data.
Source Data Fig. 3
Statistical Source Data.
Source Data Fig. 4
Statistical Source Data.
Source Data Extended Data Fig. 1
Statistical Source Data.
Source Data Extended Data Fig. 8
Statistical Source Data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, Y.H., Kim, MS., Wang, M. et al. Epigenetic plasticity enables copepods to cope with ocean acidification. Nat. Clim. Chang. 12, 918–927 (2022). https://doi.org/10.1038/s41558-022-01477-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41558-022-01477-4