Abstract
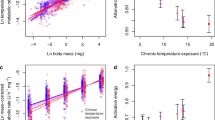
Anthropogenic climate warming affects many biological systems, ranging in scale from microbiomes to biomes. In many animals, warming-related fitness depression appears more closely linked to changes in ecological community interactions than to direct thermal stress. This biotic community framework is commonly applied to warming studies at the scale of ecosystems but is rarely applied at the scale of microbiomes. Here, we used replicated bromeliad microecosystems to show warming effects on tadpole gut microbiome dysbiosis mediated through biotic community interactions. Warming shifted environmental bacteria and arthropod community composition, with linkages to changes in microbial recruitment that promoted dysbiosis and stunted tadpole growth. Tadpole growth was more strongly associated with cascading effects of warming on gut dysbiosis than with direct warming effects or indirect effects on food resources. These results suggest that assessing warming effects on animal health requires an ecological community perspective on microbiome structure and function.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
Sequence data that support the findings of this study have been deposited in the NCBI Sequence Read Archive with the accession code PRJNA613682. Other data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Ockendon, N. et al. Mechanisms underpinning climatic impacts on natural populations: altered species interactions are more important than direct effects. Glob. Change Biol. 20, 2221–2229 (2014).
Cahill, A. E. et al. How does climate change cause extinction? Proc. R. Soc. B 280, 20121890 (2013).
Penuelas, J., Filella, I. & Comas, P. Changed plant and animal life cycles from 1952 to 2000 in the Mediterranean region. Glob. Change Biol. 8, 531–544 (2002).
Parmesan, C. et al. Poleward shifts in geographical ranges of butterfly species associated with regional warming. Nature 399, 579–583 (1999).
Freeman, B. G., Yaw, J. A. L., Sunday, J. M. & Hargreaves, A. L. Expanding, shifting and shrinking: the impact of global warming on species’ elevational distributions. Glob. Ecol. Biogeogr. 27, 1268–1276 (2018).
Draper, A. M. & Weissburg, M. J. Impacts of global warming and elevated CO2 on sensory behavior in predator–prey interactions: a review and synthesis. Front. Ecol. Evol. 7, 72 (2019).
Romero, G. Q. et al. Global predation pressure redistribution under future climate change. Nat. Clim. Change 8, 1087–1091 (2018).
Hudson, P. J., Cattadori, I. M., Boag, B. & Dobson, A. P. Climate disruption and parasite–host dynamics: patterns and processes associated with warming and the frequency of extreme climatic events. J. Helminthol. 80, 175–182 (2006).
Robinson, C. J., Bohannan, B. J. M. & Young, V. B. From structure to function: the ecology of host-associated microbial communities. Microbiol. Mol. Biol. Rev. 74, 453–476 (2010).
Longo, A. V. & Zamudio, K. R. Temperature variation, bacterial diversity and fungal infection dynamics in the amphibian skin. Mol. Ecol. 26, 4787–4797 (2017).
Longo, A. V., Savage, A. E., Hewson, I. & Zamudio, K. R. Seasonal and ontogenetic variation of skin microbial communities and relationships to natural disease dynamics in declining amphibians. R. Soc. Open Sci. 2, 140377 (2015).
Kohl, K. D. & Yahn, J. Effects of environmental temperature on the gut microbial communities of tadpoles. Environ. Microbiol. 18, 1561–1565 (2016).
Fontaine, S. S., Novarro, A. J. & Kohl, K. D. Environmental temperature alters the digestive performance and gut microbiota of a terrestrial amphibian. J. Exp. Biol. 221, 187559 (2018).
Woodhams, D. C. et al. Interacting symbionts and immunity in the amphibian skin mucosome predict disease risk and probiotic effectiveness. PLoS ONE 9, e96375 (2014).
Muletz-Wolz, C. R. et al. Inhibition of fungal pathogens across genotypes and temperatures by amphibian skin bacteria. Front. Microbiol. 8, 1551 (2017).
Bestion, E. et al. Climate warming reduces gut microbiota diversity in a vertebrate ectotherm. Nat. Ecol. Evol. 1, 0161 (2017).
Meyer, E. A., Cramp, R. L., Bernal, M. H. & Franklin, C. E. Changes in cutaneous microbial abundance with sloughing: possible implications for infection and disease in amphibians. Dis. Aquat. Organ. 101, 235–242 (2012).
Flury, S. & Gessner, M. O. Experimentally simulated global warming and nitrogen enrichment effects on microbial litter decomposers in a marsh. Appl. Environ. Microbiol. 77, 803–809 (2011).
Belden, L. K. & Harris, R. N. Infectious diseases in wildlife: the community ecology context. Front. Ecol. Environ. 5, 533–539 (2007).
Bernabé, T. N. et al. Warming weakens facilitative interactions between decomposers and detritivores, and modifies freshwater ecosystem functioning. Glob. Change Biol. 24, 3170–3186 (2018).
Hoekman, D. Turning up the heat: temperature influences the relative importance of top-down and bottom-up effects. Ecology 91, 2819–2825 (2010).
Becker, C. G. et al. Low-load pathogen spillover predicts shifts in skin microbiome and survival of a terrestrial-breeding amphibian. Proc. R. Soc. B 286, 20191114 (2019).
Jiménez, R. R. & Sommer, S. The amphibian microbiome: natural range of variation, pathogenic dysbiosis, and role in conservation. Biodivers. Conserv. 26, 763–786 (2017).
Greenspan, S. E. et al. Arthropod–bacteria interactions influence assembly of aquatic host microbiome and pathogen defense. Proc. R. Soc. B 286, 20190924 (2019).
Walker, W. A. in The Microbiota in Gastrointestinal Pathophysiology: Implications for Human Health, Prebiotics, Probiotics, and Dysbiosis (eds Floch, M. H. et al.) 227–232 (Academic Press, 2016).
Zaneveld, J. R., McMinds, R. & Vega Thurber, R. Stress and stability: applying the Anna Karenina principle to animal microbiomes. Nat. Microbiol. 2, 17121 (2017).
Jin Song, S. et al. Engineering the microbiome for animal health and conservation. Exp. Biol. Med. 244, 494–504 (2019).
Kriss, M., Hazleton, K. Z., Nusbacher, N. M., Martin, C. G. & Lozupone, C. A. Low diversity gut microbiota dysbiosis: drivers, functional implications and recovery. Curr. Opin. Microbiol. 44, 34–40 (2018).
Collins, M. et al. in Climate Change 2013: The Physical Science Basis (eds. Stocker, T. F. et al.) 1029–1136 (IPCC, Cambridge Univ. Press, 2013).
Sabagh, L. T., Ferreira, G. L., Branco, C. W. C., Rocha, C. F. D. & Dias, N. Y. N. Larval diet in bromeliad pools: a case study of tadpoles of two species in the genus Scinax (Hylidae). Copeia 2012, 683–689 (2012).
Angilletta, M. J., Niewiarowski, P. H. & Navas, C. A. The evolution of thermal physiology in ectotherms. J. Therm. Biol. 27, 249–268 (2002).
Becker, C. G., Longo, A. V., Haddad, C. F. B. & Zamudio, K. R. Land cover and forest connectivity alter the interactions among host, pathogen and skin microbiome. Proc. R. Soc. B 284, 20170582 (2017).
Fukami, T. & Nakajima, M. Community assembly: alternative stable states or alternative transient states? Ecol. Lett. 14, 973–984 (2011).
Wang, J. et al. Phylogenetic beta diversity in bacterial assemblages across ecosystems: deterministic versus stochastic processes. ISME J. 7, 1310–1321 (2013).
Longo, A. V. & Zamudio, K. R. Environmental fluctuations and host skin bacteria shift survival advantage between frogs and their fungal pathogen. ISME J. 11, 349–361 (2017).
Friedman, B. A. & Dugan, P. R. Identification of Zoogloea species and the relationship to zoogloeal matrix and floc formation. J. Bacteriol. 95, 1903–1909 (1968).
Gao, N. et al. Both widespread PEP-CTERM proteins and exopolysaccharides are required for floc formation of Zoogloea resiniphila and other activated sludge bacteria. Environ. Microbiol. 20, 1677–1692 (2018).
Merritt, R. W., Dadd, R. H. & Walker, E. D. Feeding behavior, natural food, and nutritional relationships of larval mosquitos. Annu. Rev. Entomol. 37, 349–376 (1992).
Moghadam, F. S. & Zimmer, M. Effects of warming and nutrient enrichment on how grazing pressure affects leaf litter–colonizing bacteria. J. Environ. Qual. 43, 851–858 (2014).
Zander, A., Bersier, L. & Gray, S. M. Effects of temperature variability on community structure in a natural microbial food web. Glob. Change Biol. 23, 56–67 (2017).
Aguirre, A. A. & Tabor, G. M. Global factors driving emerging infectious diseases: Impact on wildlife populations. Ann. NY Acad. Sci. 1149, 1–3 (2008).
Greenspan, S. E. et al. Infection increases vulnerability to climate change via effects on host thermal tolerance. Sci. Rep. 7, 9349 (2017).
Neely, W. J. et al. Synergistic effects of warming and disease linked to high mortality in cool-adapted terrestrial frogs. Biol. Conserv. 245, 108521 (2020).
Raffel, T. R. et al. Disease and thermal acclimation in a more variable and unpredictable climate. Nat. Clim. Change 3, 146–151 (2013).
Raffel, T. R., Halstead, N. T., Mcmahon, T. A., Davis, A. K. & Rohr, J. R. Temperature variability and moisture synergistically interact to exacerbate an epizootic disease. Proc. R. Soc. B 282, 20142039 (2015).
Greenspan, S. E. et al. White blood cell profiles in amphibians help to explain disease susceptibility following temperature shifts. Dev. Comp. Immunol. 77, 280–286 (2017).
Dézerald, O. et al. Food-web structure in relation to environmental gradients and predator–prey ratios in tank-bromeliad ecosystems. PLoS ONE 8, e71735 (2013).
Kitching, R. L. Food Webs and Container Habitats: The Natural History and Ecology of Phytotelmata (Cambridge Univ. Press, 2000).
Richardson, B. A. The bromeliad microcosm and the assessment of faunal diversity in a Neotropical forest. Biotropica 31, 321–336 (1999).
Leroy, C. et al. What drives detrital decomposition in Neotropical tank bromeliads? Hydrobiologia 802, 85–95 (2017).
Giongo, A. et al. Seasonal physiological parameters and phytotelmata bacterial diversity of two bromeliad species (Aechmea gamosepala and Vriesea platynema) from the Atlantic Forest of Southern Brazil. Diversity 11, 111 (2019).
Frank, J. H. & Lounibos, L. P. Insects and allies associated with bromeliads: a review. Terr. Arthropod Rev. 1, 125–153 (2009).
Ruano-Fajardo, G., Toledo, L. F. & Mott, T. Jumping into a trap: high prevalence of chytrid fungus in the preferred microhabitats of a bromeliad-specialist frog. Dis. Aquat. Organ. 121, 223–232 (2016).
Gomez-Hoyos, D. A. et al. Phytotelmata selection by anurans and implications for their conservation at Las Tablas Protected Zone, Costa Rica. Alytes 35, 1–11 (2018).
Haddad, C. F. B. et al. Guia dos Anfíbios da Mata Atlântica: Diversidade e Biologia (Anolis Books, 2013).
Gosner, K. L. A simplified table for staging anuran embryos and larvae with notes on identification. Herpetologica 16, 183–190 (1960).
Oksanen, J. et al. vegan: community ecology package. R package version 2.4-1 https://CRAN.R-project.org/package=vegan (2016).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2018).
Caporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 6, 1621–1624 (2012).
Caporaso, J. G. et al. EMP 16S Illumina Amplicon Protocol (protocols.io, 2018).
Kozich, J. J., Westcott, S. L., Baxter, N. T., Highlander, S. K. & Schloss, P. D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl. Environ. Microbiol. 79, 5112–5120 (2013).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 848–857 (2019).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Amir, A. et al. Deblur rapidly resolves single-nucleotide community sequence patterns. mSystems 2, e00191–16 (2017).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Bokulich, N. A. et al. Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2’s q2-feature-classifier plugin. Microbiome 6, 90 (2018).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072 (2006).
McDonald, D. et al. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 6, 610–618 (2012).
Bokulich, N. A. et al. Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat. Methods 10, 57–59 (2013).
Faith, D. P. Conservation evaluation and phylogenetic diversity. Biol. Conserv. 61, 1–10 (1992).
Lozupone, C. & Knight, R. UniFrac: a new phylogenetic method for comparing microbial communities. Appl. Environ. Microbiol. 71, 8228–8235 (2005).
Lozupone, C. A., Hamady, M., Kelley, S. T. & Knight, R. Quantitative and qualitative β diversity measures lead to different insights into factors that structure microbial communities. Appl. Environ. Microbiol. 73, 1576–1585 (2007).
Lozupone, C., Lladser, M. E., Knights, D., Stombaugh, J. & Knight, R. UniFrac: an effective distance metric for microbial community comparison. ISME J. 5, 169–172 (2011).
Lefcheck, J. S. piecewiseSEM: Piecewise structural equation modelling in R for ecology. Methods Ecol. Evol. 7, 573–579 (2016).
Deegan, J. On the occurrence of standardized regression coefficients greater than one. Educ. Psychol. Meas. 38, 873–888 (1978).
JMP v.14.0.0 (SAS Institute, 2019).
Warnes, G. et al. gplots: various R programming tools for plotting data. R package version 3.0.1.1 https://CRAN.R-project.org/package=gplots (2019).
Acknowledgements
We thank R. Bell, T. Bernabe, M. Bletz, T. Jenkinson, R. Martins, D. Medina, W. Neely and R. Salla Jacob. São Paulo Research Foundation (FAPESP) provided grants to M.L.L. (grant no. 2017/26162-8), L.P.R. (grant nos. 2018/23622-0 and 2016/25358-3), L.F.T. (grant nos. 2016/25358-3 and 2019/18335-5), C.F.B.H. (grant no. 2013/50741-7) and G.Q.R. (grant nos. 2017/09052-4 and 2018/12225-0). National Council for Scientific and Technological Development (CNPq) provided research fellowships to L.F.T. (grant no. 300896/2016-6), C.F.B.H. (grant no. 306623/2018-8) and G.Q.R. The Royal Society provided a Newton Advanced Fellowship to G.Q.R. (grant no. NAF\R2\180791).
Author information
Authors and Affiliations
Contributions
C.G.B. and S.E.G. designed the study. All authors carried out the study. C.G.B., S.E.G. and G.Q.R. analysed the data. S.E.G. drafted the manuscript. All authors critically revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Climate Change thanks Obed Hernandez-Gomez and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2, and Tables 1–8.
Rights and permissions
About this article
Cite this article
Greenspan, S.E., Migliorini, G.H., Lyra, M.L. et al. Warming drives ecological community changes linked to host-associated microbiome dysbiosis. Nat. Clim. Chang. 10, 1057–1061 (2020). https://doi.org/10.1038/s41558-020-0899-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41558-020-0899-5
This article is cited by
-
Effects of grassland controlled burning on symbiotic skin microbes in Neotropical amphibians
Scientific Reports (2024)
-
Disentangling direct vs indirect effects of microbiome manipulations in a habitat-forming marine holobiont
npj Biofilms and Microbiomes (2024)
-
Networks of climate change: connecting causes and consequences
Applied Network Science (2023)
-
Comparing western (Megascops kennicottii) and whiskered (M. trichopsis) screech-owl microbiomes in southern Arizona using a novel 16S rRNA sequencing method
Animal Microbiome (2022)
-
Signatures of functional bacteriome structure in a tropical direct-developing amphibian species
Animal Microbiome (2022)