Abstract
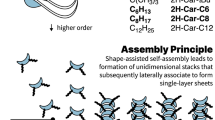
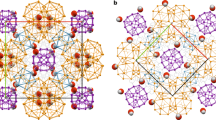
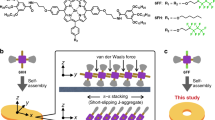
Self-assembly is a powerful method to obtain large discrete functional molecular architectures. When using a single building block, self-assembly generally yields symmetrical objects in which all the subunits relate similarly to their neighbours. Here we report the discovery of a family of self-constructing cyclic macromolecules with stable folded conformations of low symmetry, which include some with a prime number (13, 17 and 23) of units, despite being formed from a single component. The formation of these objects amounts to the production of polymers with a perfectly uniform length. Design rules for the spontaneous emergence of such macromolecules include endowing monomers with a strong potential for non-covalent interactions that remain frustrated in competing entropically favoured yet conformationally restrained smaller cycles. The process can also be templated by a guest molecule that itself has an asymmetrical structure, which paves the way to molecular imprinting techniques at the level of single polymer chains.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The authors declare that all the data supporting the findings of this study are available within the article, in the source data files and in the Supplementary Information. Mass and NMR spectra are stored locally in native format and are available upon request. Crystallographic data for the structures reported in this article have been deposited at the Cambridge Crystallographic Data Centre, under deposition numbers CCDC 1942977 for (4b)16 and 1999456 for (4d)23, respectively. Copies of the data can be obtained free of charge via https://www.ccdc.cam.ac.uk/structures/. Source data are provided with this paper.
References
Fujita, D. et al. Self-assembly of tetravalent Goldberg polyhedra from 144 small components. Nature 540, 563–566 (2016).
Bale, J. B. et al. Accurate design of megadalton-scale two-component icosahedral protein complexes. Science 353, 389–394 (2016).
Pasquale, S., Sattin, S., Escudero-Adan, E. C., Martinez-Belmonte, M. & de Mendoza, J. Giant regular polyhedra from calixarene carboxylates and uranyl. Nat. Commun. 3, 785 (2012).
Sasaki, E. et al. Structure and assembly of scalable porous protein cages. Nat. Commun. 8, 14663 (2017).
Anderson, K. M., Goeta, A. E. & Steed, J. W. Supramolecular synthon frustration leads to crystal structures with Z′ > 1. Cryst. Growth Des. 8, 2517–2524 (2008).
Banerjee, R., Bhatt, P. M., Kirchner, M. T. & Desiraju, G. R. Structural studies of the system Na(saccharinate)·nH2O: a model for crystallization. Angew. Chem. Int. Ed. 44, 2515–2520 (2005).
Rhys, G. G. et al. Maintaining and breaking symmetry in homomeric coiled-coil assemblies. Nat. Commun. 9, 4132 (2018).
Whitesides, G. M. & Ismagilov, R. F. Complexity in chemistry. Science 284, 89–92 (1999).
Lehn, J.-M. From supramolecular chemistry towards constitutional dynamic chemistry and adaptive chemistry. Chem. Soc. Rev. 36, 151–160 (2007).
Lehn, J.-M. Constitutional dynamic chemistry: bridge from supramolecular chemistry to adaptive chemistry. Top. Curr. Chem. 322, 1–32 (2012).
Vantomme, G. & Meijer, E. W. The construction of supramolecular systems. Science 363, 1396–1397 (2019).
Nitschke, J. R. Systems chemistry: molecular networks come of age. Nature 462, 736–738 (2009).
Li, J., Nowak, P. & Otto, S. Dynamic combinatorial libraries: from exploring molecular recognition to systems chemistry. J. Am. Chem. Soc. 135, 9222–9239 (2013).
Corbett, P. T. et al. Dynamic combinatorial chemistry. Chem. Rev. 106, 3652–3711 (2006).
Cougnon, F. B. L. & Sanders, J. K. M. Evolution of dynamic combinatorial chemistry. Acc. Chem. Res. 45, 2211–2221 (2012).
Tsiamantas, C. et al. Selective dynamic assembly of disulfide macrocyclic helical foldamers with remote communication of handedness. Angew. Chem. Int. Ed. 55, 6848–6852 (2016).
Steinkruger, J. D., Woolfson, D. N. & Gellman, S. H. Side-chain pairing preferences in the parallel coiled-coil dimer motif: insight on ion pairing between core and flanking sites. J. Am. Chem. Soc. 132, 7586–7588 (2010).
Hadley, E. B., Testa, O. D., Woolfson, D. N. & Gellman, S. H. Preferred side-chain constellations at antiparallel coiled-coil interfaces. Proc. Natl Acad. Sci. USA 105, 530–535 (2008).
Krishnan-Ghosh, Y. & Balasubramanian, S. Dynamic covalent chemistry on self-templating peptides: formation of a disulfide-linked beta-hairpin mimic. Angew. Chem. Int. Ed. 42, 2171–2173 (2003).
Oh, K., Jeong, K. S. & Moore, J. S. Folding-driven synthesis of oligomers. Nature 414, 889–893 (2001).
Liu, B. et al. Complex molecules that fold like proteins can emerge spontaneously. J. Am. Chem. Soc. 141, 1685–1689 (2019).
Carnall, J. M. A. et al. Mechanosensitive self-replication driven by self-organization. Science 327, 1502–1506 (2010).
Komáromy, D. et al. Self-assembly can direct dynamic covalent bond formation toward diversity or specificity. J. Am. Chem. Soc. 139, 6234–6241 (2017).
Bartolec, B., Altay, M. & Otto, S. Template-promoted self-replication in dynamic combinatorial libraries made from a simple building block. Chem. Commun. 54, 13096–13098 (2018).
Sadownik, J. W., Mattia, E., Nowak, P. & Otto, S. Diversification of self-replicating molecules. Nat. Chem. 8, 264–269 (2016).
Seo, J. et al. An infrared spectroscopy approach to follow β-sheet formation in peptide amyloid assemblies. Nat. Chem. 9, 39–44 (2016).
Käseborn, M., Holstein, J. J., Clever, G. H. & Lützen, A. A rotaxane-like cage-in-ring structural motif for a metallosupramolecular Pd6L12 aggregate. Angew. Chem. Int. Ed. 57, 12171–12175 (2018).
Huang, B., Prantil, M. A., Gustafson, T. L. & Parquette, J. R. The effect of global compaction on the local secondary structure of folded dendrimers. J. Am. Chem. Soc. 125, 14518–14530 (2003).
BelBruno, J. J. Molecularly imprinted polymers. Chem. Rev. 119, 94–119 (2019).
Allen, S. J., Giles, K., Gilbert, T. & Bush, M. F. Ion mobility mass spectrometry of peptide, protein, and protein complex ions using a radio-frequency confining drift cell. Analyst 141, 884–891 (2016).
Warnke, S., von Helden, G. & Pagel, K. Analyzing the higher order structure of proteins with conformer-selective ultraviolet photodissociation. Proteomics 15, 2804–2812 (2015).
Revercomb, H. E. & Mason, E. A. Theory of plasma chromatography/gaseous electrophoresis. Review. Anal. Chem. 47, 970–983 (1975).
Nurizzo, D. et al. The ID23-1 structural biology beamline at the ESRF. J. Synchrotron Rad 13, 227–238 (2006).
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010).
Sheldrick, G. M. SHELXT-integrated space-group and crystal-structure determination. Acta Crystallogr. A 71, 3–8 (2015).
Sheldrick, G. M. Crystal structure refinement with SHELXL. Acta Crystallogr. C 71, 3–8 (2015).
Dolomanov, O. V., Bourhis, L. J., Gildea, R. J., Howard, J. A. K. & Puschmann, H. OLEX2: a complete structure solution, refinement and analysis program. J. Appl. Cryst. 42, 339–341 (2009).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Spek, A. L. Structure validation in chemical crystallography. Acta Crystallogr. D 65, 148–155 (2009).
Cianci, M. et al. P13, the EMBL macromolecular crystallography beamline at the low-emittance PETRA III ring for high- and low-energy phasing with variable beam focussing. J. Synchrotron Rad. 24, 323–332 (2017).
CrysAlisPRO (Agilent Technologies Ltd, 2014).
Acknowledgements
We thank P. van der Meulen and J. Kemmink for their assistance with the NMR experiments and analysing the data. We thank I. Melnikov (ID23-1, ERSF) and G. Pompidor (PETRA III, DESY) for assistance during data collection at the synchrotron beamlines. This research was supported by the ERC (AdG 741774), the EU (MCIF 745805−DSR), NWO (VICI grant), Zernike Dieptestrategie and the Dutch Ministry of Education, Culture and Science (Gravitation program 024.001.035).
Author information
Authors and Affiliations
Contributions
S.O. and I.H. supervised the overall project. C.G.P. conceived and designed the study, synthesized the building blocks, analysed the DCL compositions by UPLC and isolated and characterized the foldamers by CD and NMR spectroscopy. B.L. performed the UPLC-MS experiments and analysed the data. P.K.M. and B.K. carried out the crystallographic studies. X.M. and K.L. performed the ITC experiments. D.K., W.H., C.M., R.C. and K.P. performed the IM-MS experiments and analysed the data. C.G.P., I.H. and S.O. co-wrote the paper. All the authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–126 and Tables 1–4.
Supplementary Video 1
Crystal structure of macrocycle (4b)16.
Supplementary Video 2
Crystal structure of macrocycle (4d)23.
Supplementary Data 1
Raw UPLC chromatograms, raw temperature-dependent CD spectral data and ion-mobility data.
Supplementary Data 2
Crystallographic information file including structure factor for the structure of (4b)16.
Supplementary Data 3
Crystallographic information file including structure factor for the structure of (4d)23.
Source data
Source Data Fig. 2
Raw UPLC chromatograms, raw ion-mobility and CD spectral data.
Rights and permissions
About this article
Cite this article
Pappas, C.G., Mandal, P.K., Liu, B. et al. Emergence of low-symmetry foldamers from single monomers. Nat. Chem. 12, 1180–1186 (2020). https://doi.org/10.1038/s41557-020-00565-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-020-00565-2
This article is cited by
-
Catalytic length-controlled oligomerization with synthetic programmable templates
Nature Synthesis (2023)