Abstract
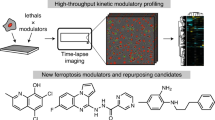
The chemical diversification of natural products provides a robust and general method for the creation of stereochemically rich and structurally diverse small molecules. The resulting compounds have physicochemical traits different from those in most screening collections, and as such are an excellent source for biological discovery. Herein, we subject the diterpene natural product pleuromutilin to reaction sequences focused on creating ring system diversity in few synthetic steps. This effort resulted in a collection of compounds with previously unreported ring systems, providing a novel set of structurally diverse and highly complex compounds suitable for screening in a variety of different settings. Biological evaluation identified the novel compound ferroptocide, a small molecule that rapidly and robustly induces ferroptotic death of cancer cells. Target identification efforts and CRISPR knockout studies reveal that ferroptocide is an inhibitor of thioredoxin, a key component of the antioxidant system in the cell. Ferroptocide positively modulates the immune system in a murine model of breast cancer and will be a useful tool to study the utility of pro-ferroptotic agents for treatment of cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the paper and Supplementary Information and are available from the corresponding author upon request. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the data set identifier PXD012805 (www.ebi.ac.uk/pride/archive/projects/PXD012805). The RNA sequencing data have been deposited to the GEO repository with accession number GSE126868 (www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE126868). The X-ray crystallography data have been deposited in the Cambridge Crystallographic Data Centre (CCDC) using the following identifiers (www. ccdc.cam.ac.uk/structures/): 1851845 (compound P1) and 1849494 (Ferroptocide).
References
Lovering, F., Bikker, J. & Humblet, C. Escape from flatland: increasing saturation as an approach to improving clinical success. J. Med. Chem. 52, 6752–6756 (2009).
Swinney, D. C. & Anthony, J. How were new medicines discovered? Nat. Rev. Drug Discov. 10, 507–519 (2011).
Galloway, W. R. J. D., Isidro-Llobet, A. & Spring, D. R. Diversity-oriented synthesis as a tool for the discovery of novel biologically active small molecules. Nat. Commun. 1, 80–93 (2010).
Gerry, C. J. & Schreiber, S. L. Chemical probes and drug leads from advances in synthetic planning and methodology. Nat. Rev. Drug Discov. 17, 333–352 (2018).
Morrison, K. C. & Hergenrother, P. J. Natural products as starting points for the synthesis of complex and diverse compounds. Nat. Prod. Rep. 31, 6–14 (2014).
Huigens, R. W. III et al. A ring-distortion strategy to construct stereochemically complex and structurally diverse compounds from natural products. Nat. Chem. 5, 195–202 (2013).
Ciardiello, J. J. et al. A novel complexity-to-diversity strategy for the diversity-oriented synthesis of structurally diverse and complex macrocycles from quinine. Bioorg. Med. Chem. 25, 2825–2843 (2017).
Rafferty, R. J., Hicklin, R. W., Maloof, K. A. & Hergenrother, P. J. Synthesis of complex and diverse compounds through ring distortion of abietic acid. Angew. Chem. Int. Ed. 53, 220–224 (2014).
Garcia, A., Drown, B. S. & Hergenrother, P. J. Access to a structurally complex compound collection via ring distortion of the alkaloid sinomenine. Org. Lett. 18, 4852–4855 (2016).
Tasker, S. Z., Cowfer, A. E. & Hergenrother, P. J. Preparation of structurally diverse compounds from the natural product lycorine. Org. Lett. 20, 5894–5898 (2018).
Paciaroni, N. G. et al. A tryptoline ring‐distortion strategy leads to complex and diverse biologically active molecules from the indole alkaloid yohimbine. Chem. Eur. J. 23, 4327–4335 (2017).
Govindaraju, K. et al. Novel topologically complex scaffold derived from alkaloid haemanthamine. Molecules 23, 255–263 (2018).
Charaschanya, M. & Aubé, J. Reagent-controlled regiodivergent ring expansions of steroids. Nat. Commun. 9, 934–942 (2018).
Laurent, E. et al. A ring‐distortion strategy from marine natural product ilimaquinone leads to quorum sensing modulators. Eur. J. Org. Chem. 2018, 2486–2497 (2018).
Xu, H. et al. Identification of a diverse synthetic abietane diterpenoid library for anticancer activity. Bioorg. Med. Chem. Lett. 27, 505–510 (2017).
Luca, L. et al. Discovery of novel cinchona‐alkaloid‐inspired oxazatwistane autophagy inhibitors. Angew. Chem. Int. Ed. 56, 2145–2150 (2017).
Richter, M. F. et al. Predictive compound accumulation rules yield a broad-spectrum antibiotic. Nature 545, 299–304 (2017).
Poulsen, S. M., Karlsson, M., Johansson, L. B. & Vester, B. The pleuromutilin drugs tiamulin and valnemulin bind to the RNA at the peptidyl transferase centre on the ribosome. Mol. Microbiol. 41, 1091–1099 (2001).
Ma, X. et al. Directed C–H bond oxidation of (+)-pleuromutilin. J. Org. Chem. 83, 6843–6892 (2018).
Farney, E. P., Feng, S. S., Schäfers, F. & Reisman, S. E. Total synthesis of (+)-pleuromutilin. J. Am. Chem. Soc. 140, 1267–1270 (2018).
Murphy, S. K., Zeng, M. & Herzon, S. B. A modular and enantioselective synthesis of the pleuromutilin antibiotics. Science 356, 956–959 (2017).
Thirring, K. et al. 12-epi-pleuromutilins. US patent WO2015110481A1 (2015).
Gibbons, E. G. Total synthesis of (±)-pleuromutilin. J. Am. Chem. Soc. 104, 1767–1769 (1982).
Paquette, L. A., Wiedeman, P. E. & Bulman-Page, P. C. (+)-Pleuromutilin synthetic studies. Degradative and de novo acquisition of a levorotatory tricyclic lactone subunit. J. Org. Chem. 53, 1441–1450 (1988).
Liu, J., Lotesta, S. D. & Sorensen, E. J. A concise synthesis of the molecular framework of pleuromutilin. Chem. Commun. 47, 1500–1502 (2011).
Birch, A. J., Holzapfel, C. W. & Rickards, R. W. The structure and some aspects of the biosynthesis of pleuromutilin. Tetrahedron 22, 359–387 (1966).
Arigoni, D. Some studies in the biosynthesis of terpenes and related compounds. Pure Appl. Chem. 17, 331–348 (1968).
Drews, J. et al. Antimicrobial activities of 81.723 hfu, a new pleuromutilin derivative. Antimicrob. Agents Chemother. 7, 507–516 (1975).
Yang, L. P. & Keam, S. J. Retapamulin: a review of its use in the management of impetigo and other uncomplicated superficial skin infections. Drugs 68, 855–873 (2008).
Berner, H., Schulz, G. & Schneider, H. Synthese ab-trans-anellierter derivate des tricyclischen diterpens pleuromutilin durch intramolekulare 1,5-hydrid-verschiebung. Tetrahedron 36, 1807–1811 (1980).
Andemichael, Y. et al. Process development for a novel pleuromutilin-derived antibiotic. Org. Process Res. Dev. 13, 729–738 (2009).
Uccello, D. P. et al. The synthesis of C-13 functionalized pleuromutilins via C–H amidation and subsequent novel rearrangement product. Tetrahedron Lett. 52, 4247–4251 (2011).
Hicklin, R. W., López Silva, T. L. & Hergenrother, P. J. Synthesis of bridged oxafenestranes from pleuromutilin. Angew. Chem. Int. Ed. 53, 9880–9883 (2014).
Botham, R. C. et al. Dual small-molecule targeting of procaspase-3 dramatically enhances zymogen activation and anticancer activity. J. Am. Chem. Soc. 136, 1312–1319 (2014).
Parkinson, E. I., Bair, J. S., Cismesia, M. & Hergenrother, P. J. Efficient NQO1 substrates are potent and selective anticancer agents. ACS Chem. Biol. 8, 2173–2183 (2013).
Palchaudhuri, R. et al. A small molecule that induces intrinsic pathway apoptosis with unparalleled speed. Cell Rep. 13, 2027–2036 (2015).
Linkermann, A., Stockwell, B. R., Krautwald, S. & Anders, H.-J. Regulated cell death and inflammation: an auto-amplification loop causes organ failure. Nat. Rev. Immunol. 14, 759–767 (2014).
Galluzzi, L., Senovilla, L., Zitvogel, L. & Kroemer, G. The secret ally: immunostimulation by anticancer drugs. Nat. Rev. Drug Discov. 11, 215–233 (2012).
Lee, H. Y. et al. Reactive oxygen species synergize to potently and selectively induce cancer cell death. ACS Chem. Biol. 12, 1416–1424 (2017).
Stockwell, B. R. et al. Ferroptosis: a regulated cell death nexus linking metabolism, redox biology and disease. Cell 171, 273–285 (2017).
Dixon, S. J. et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149, 1060–1072 (2012).
Yang, W. S. et al. Regulation of ferroptotic cancer cell death by GPX4. Cell 156, 317–331 (2014).
Shimada, K. et al. Global survey of cell death mechanisms reveals metabolic regulation of ferroptosis. Nat. Chem. Biol. 12, 497–503 (2016).
Gaschler, M. M. et al. FINO2 initiates ferroptosis through GPX4 inactivation and iron oxidation. Nat. Chem. Biol. 14, 507–515 (2018).
Lewerenz, J. et al. Oxytosis/ferroptosis—(re-)emerging roles for oxidative stress-dependent non-apoptotic cell death in diseases of the central nervous system. Front. Neurosci. 12, 214 (2018).
Murphy, T. H. et al. Calcium-dependent glutamate cytotoxicity in a neuronal cell line. Brain Res. 444, 325–332 (1988).
Miyamoto, M., Murphy, T. H., Schnaar, R. L. & Coyle, J. T. Antioxidants protect against glutamate-induced cytotoxicity in a neuronal cell line. J. Pharmacol. Exp. Ther. 250, 1132–1140 (1989).
Seiler, A. et al. Glutathione peroxidase 4 senses and translates oxidative stress into 12/15-lipoxygenase dependent- and AIF-mediated cell death. Cell Metabol. 8, 237–248 (2008).
Yang, W. S. & Stockwell, B. R. Synthetic lethal screening identifies compounds activating iron-dependent, nonapoptotic cell death in oncogenic-RAS-harboring cancer cells. Chem. Biol. 15, 234–245 (2008).
Subramanian, A. et al. A next generation connectivity map: L1000 platform and the first 1,000,000 profiles. Cell 171, 1437–1452 (2017).
Dixon, S. J. et al. Pharmacological inhibition of cystine–glutamate exchange induces endoplasmic reticulum stress and ferroptosis. eLife 3, e02523 (2014).
Banerjee, R., Pace, N. J., Brown, D. R. & Weerapana, E. 1,3,5-Triazine as a modular scaffold for covalent inhibitors with streamlined target identification. J. Am. Chem. Soc. 135, 2497–2500 (2013).
Holmgren, A. Thioredoxin structure and mechanism: conformational changes on oxidation of the active-site sulfhydryls to a disulfide. Structure 3, 239–243 (1995).
Nordberg, J. & Arnér, E. S. J. Reactive oxygen species, antioxidants and the mammalian thioredoxin system. Free Radic. Biol. Med. 31, 1287–1312 (2001).
Mukherjee, A. et al. A cellular and molecular investigation of the action of PMX464, a putative thioredoxin inhibitor, in normal and colorectal cancer cell lines. Br. J. Pharmacol. 151, 1167–1175 (2007).
Baker, A. F. et al. The antitumor thioredoxin-1 inhibitor PX-12 (1-methylpropyl 2-imidazolyl disulfide) decreases thioredoxin-1 and VEGF levels in cancer patient plasma. J. Lab. Clin. Med. 147, 83–90 (2006).
Kirkpatrick, D. L. et al. Mechanisms of inhibition of the thioredoxin growth factor system by antitumor 2-imidazolyl disulfides. Biochem. Pharmacol. 55, 987–994 (1998).
Sexton, D. W. Targeting airway inflammation: PMX464 and the epithelial bulls eye. Br. J. Pharmacol. 155, 620–622 (2008).
Reynoso, E. et al. Thioredoxin-1 actively maintains the pseudokinase MLKL in a reduced state to suppress disulfide bond-dependent MLKL polymer formation and necroptosis. J. Biol. Chem. 292, 17514–17524 (2017).
You, B. R., Shin, H. R. & Park, W. H. PX-12 inhibits the growth of A549 lung cancer cells via G2/M phase arrest and ROS-dependent apoptosis. Int. J. Oncol. 44, 301–308 (2014).
Li, Y., Qian, L. & Yuan, J. Small molecule probes for cellular death machines. Curr. Opin. Chem. Biol. 39, 74–82 (2017).
Galluzzi, L. et al. Molecular mechanisms of cell death: recommendations of the nomenclature committee on cell death 2018. Cell Death Differ. 25, 486–541 (2018).
Mai, T. T. et al. Salinomycin kills cancer stem cells by sequestering iron in lysosomes. Nat. Chem. 9, 1025–1033 (2017).
Bruni, A. et al. Ferroptosis-inducing agents compromise in vitro human islet viability and function. Cell Death Dis. 9, 595–605 (2018).
Acknowledgements
The authors acknowledge support from the University of Illinois and Cancer Scholars for Translational and Applied Research (C*STAR) programme and thank the NIH (R01GM118575) for support of some of the synthetic studies. The authors thank W. Woods and P. Perez Pinera for assistance with CRISPR–Cas9 studies, D. Gray and T. Woods for X-ray analysis of compounds, L. Dirikolu for calculation of pharmacokinetic (PK) parameters, B. Drown for chemoinformatic analysis of compounds, M.E. Vinyard and A. Sowers for synthetic assistance and S. Tasker for many helpful discussions.
Author information
Authors and Affiliations
Contributions
P.J.H., E.L. and R.W.H. conceived this study. R.W.H. designed and synthesized all compounds. E.L. designed and performed all biological experiments and analysed data. H.Y.L. performed the mouse model studies. S.E.M. synthesized the lead compound for the mouse model. L.A.C. and E.W. collected and analysed the LC/LC–MS/MS data. P.J.H. supervised this research. P.J.H. and E.L. wrote this manuscript with the assistance of R.W.H. All authors have given their approval of the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The University of Illinois has filed patents on some compounds described in this manuscript.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Information
The Supplementary Information contains Supplementary Figs. 1–15, methods, chemical schemes, and NMR data
Crystallographic data
Crystallographic data for compound P1. CCDC reference 1851845
Crystallographic data
Structure factors file for compound P1. CCDC reference 1851845
Crystallographic data
Crystallographic data for Ferroptocide. CCDC reference 1849494
Crystallographic data
Structure factors file for Ferroptocide. CCDC reference 1849494
Rights and permissions
About this article
Cite this article
Llabani, E., Hicklin, R.W., Lee, H.Y. et al. Diverse compounds from pleuromutilin lead to a thioredoxin inhibitor and inducer of ferroptosis. Nat. Chem. 11, 521–532 (2019). https://doi.org/10.1038/s41557-019-0261-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-019-0261-6
This article is cited by
-
Modulating ferroptosis sensitivity: environmental and cellular targets within the tumor microenvironment
Journal of Experimental & Clinical Cancer Research (2024)
-
Ferroptosis in cancer: From molecular mechanisms to therapeutic strategies
Signal Transduction and Targeted Therapy (2024)
-
Sulfasalazine promotes ferroptosis through AKT-ERK1/2 and P53-SLC7A11 in rheumatoid arthritis
Inflammopharmacology (2024)
-
The small molecule raptinal can simultaneously induce apoptosis and inhibit PANX1 activity
Cell Death & Disease (2024)
-
Quantitative reactive cysteinome profiling reveals a functional link between ferroptosis and proteasome-mediated degradation
Cell Death & Differentiation (2023)