Abstract
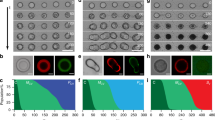
Multicellularity enables the growth of complex life forms as it allows for the specialization of cell types, differentiation and large-scale spatial organization. In a similar way, modular construction of synthetic multicellular systems will lead to dynamic biomimetic materials that can respond to their environment in complex ways. To achieve this goal, artificial cellular communication and developmental programs still have to be established. Here, we create geometrically controlled spatial arrangements of emulsion-based artificial cellular compartments containing synthetic in vitro gene circuitry, separated by lipid bilayer membranes. We quantitatively determine the membrane pore-dependent response of the circuits to artificial morphogen gradients, which are established via diffusion from dedicated organizer cells. Utilizing different types of feedforward and feedback in vitro gene circuits, we then implement artificial signalling and differentiation processes, demonstrating the potential for the realization of complex spatiotemporal dynamics in artificial multicellular systems.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Raw data used for the generation of the figures are available from the authors upon request. Plasmids pSB1A3-AD009, pSB1A3-AD010 and pSB1A3-AD011 are available on Addgene.
References
Szathmáry, E. & Smith, J. M. The major evolutionary transitions. Nature 374, 227–232 (1995).
Grosberg, R. K. & Strathmann, R. R. The evolution of multicellularity: a minor major transition? Annu. Rev. Ecol. Evol. System. 38, 621–654 (2007).
Bell, G. & Mooers, A. O. Size and complexity among multicellular organisms. Biol. J. Linn. Soc. 60, 345–363 (1997).
Crick, F. Diffusion in embryogenesis. Nature 225, 420 (1970).
Christian, J. L. Morphogen gradients in development: from form to function. Wiley Interdiscip. Rev. Dev. Biol. 1, 3–15 (2012).
Green, J. B. A. & Sharpe, J. Positional information and reaction-diffusion: two big ideas in developmental biology combine. Development 142, 1203–1211 (2015).
Elowitz, M. B. & Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000).
Danino, T., Mondragón-Palomino, O., Tsimring, L. & Hasty, J. A synchronized quorum of genetic clocks. Nature 463, 326–330 (2010).
Basu, S., Gerchman, Y., Collins, C. H., Arnold, F. H. & Weiss, R. A synthetic multicellular system for programmed pattern formation. Nature 434, 1130–1134 (2005).
Baccouche, A., Montagne, K., Padirac, A., Fujii, T. & Rondelez, Y. Dynamic DNA-toolbox reaction circuits: a walkthrough. Methods 67, 234–249 (2014).
Kim, J., White, K. S. & Winfree, E. Construction of an in vitro bistable circuit from synthetic transcriptional switches. Mol. Syst. Biol. 2, 68 (2006).
Ayukawa, S., Takinoue, M. & Kiga, D. RTRACS: a modularized RNA-dependent RNA transcription system with high programmability. Acc. Chem. Res. 44, 1369–1379 (2011).
Shin, J. & Noireaux, V. An E. coli cell-free expression toolbox: application to synthetic gene circuits and artificial cells. ACS Synth. Biol. 1, 29–41 (2012).
Karzbrun, E., Tayar, A. M., Noireaux, V. & Bar-Ziv, R. H. Programmable on-chip DNA compartments as artificial cells. Science 345, 829–832 (2014).
Isalan, M., Lemerle, C. & Serrano, L. Engineering gene networks to emulate Drosophila embryonic pattern formation. PLoS Biol. 3, e64 (2005).
Zadorin, A. S., Rondelez, Y., Galas, J.-C. & Estevez-Torres, A. Synthesis of programmable reaction–diffusion fronts using DNA catalyzers. Phys. Rev. Lett. 114, 068301 (2015).
Tayar, A. M., Karzbrun, E., Noireaux, V. & Bar-Ziv, R. H. Propagating gene expression fronts in a one-dimensional coupled system of artificial cells. Nat. Phys. 11, 1037–1041 (2015).
Zadorin, A. S. et al. Synthesis and materialization of a reaction–diffusion French flag pattern. Nat. Chem. 9, 990–996 (2017).
Chirieleison, S. M., Allen, P. B., Simpson, Z. B., Ellington, A. D. & Chen, X. Pattern transformation with DNA circuits. Nat. Chem. 5, 1000–1005 (2013).
Adamala, K. P., Martin-Alarcon, D. A., Guthrie-Honea, K. R. & Boyden, E. S. Engineering genetic circuit interactions within and between synthetic minimal cells. Nat. Chem. 9, 431–439 (2017).
Hasatani, K. et al. High-throughput and long-term observation of compartmentalized biochemical oscillators. Chem. Commun. 49, 8090–8092 (2013).
Weitz, M. et al. Diversity in the dynamical behaviour of a compartmentalized programmable biochemical oscillator. Nat. Chem. 6, 295–302 (2014).
Genot, A. J. et al. High-resolution mapping of bifurcations in nonlinear biochemical circuits. Nat. Chem. 8, 760 (2016).
Hansen, M. M. K. et al. Macromolecular crowding creates heterogeneous environments of gene expression in picolitre droplets. Nat. Nanotech. 11, 191–197 (2016).
Niederholtmeyer, H. et al. Rapid cell-free forward engineering of novel genetic ring oscillators. eLife 4, e09771 (2015).
Semenov, S. N. et al. Rational design of functional and tunable oscillating enzymatic networks. Nat. Chem. 7, 160–165 (2015).
Lentini, R. et al. Integrating artificial with natural cells to translate chemical messages that direct E. coli behaviour. Nat. Commun. 5, 4012 (2014).
Weitz, M. et al. Communication and computation by bacteria compartmentalized within microemulsion droplets. J. Am. Chem. Soc. 136, 72–75 (2014).
Schwarz-Schilling, M., Aufinger, L., Muckl, A. & Simmel, F. C. Chemical communication between bacteria and cell-free gene expression systems within linear chains of emulsion droplets. Integrat. Biol. 8, 564–570 (2016).
Qiao, Y., Li, M., Booth, R. & Mann, S. Predatory behaviour in synthetic protocell communities. Nat. Chem. 9, 110–119 (2016).
Villar, G., Graham, A. D. & Bayley, H. A tissue-like printed material. Science 340, 48–52 (2013).
Thiam, A. R., Bremond, N. & Bibette, J. From stability to permeability of adhesive emulsion bilayers. Langmuir 28, 6291–6298 (2012).
Yasuga, H. et al. Logic gate operation by DNA translocation through biological nanopores. PLoS ONE 11, e0149667 (2016).
Booth, M. J., Restrepo Schild, V., Box, S. J. & Bayley, H. Light-patterning of synthetic tissues with single droplet resolution. Sci. Rep. 7, 9315 (2017).
Elani, Y., Law, R. V. & Ces, O. Vesicle-based artificial cells as chemical microreactors with spatially segregated reaction pathways. Nat. Commun. 5, 5305 (2014).
Elani, Y., Law, R. V. & Ces, O. Protein synthesis in artificial cells: using compartmentalisation for spatial organisation in vesicle bioreactors. Phys. Chem. Chem. Phys. 17, 15534–15537 (2015).
Bayley, H. et al. Droplet interface bilayers. Mol. Biosyst. 4, 1191–1208 (2008).
Song, L. et al. Structure of Staphylococcal α-hemolysin, a heptameric transmembrane pore. Science 274, 1859–1865 (1996).
Tilley, S. J. & Saibil, H. R. The mechanism of pore formation by bacterial toxins. Curr. Opin. Struct. Biol. 16, 230–236 (2006).
Sun, Z. Z. et al. Protocols for implementing and Escherichia coli based TX-TL cell-free expression system for synthetic biology. J. Vis. Exp. 79, e50762 (2013).
Ghose, A. K., Viswanadhan, V. N. & Wendoloski, J. J. A knowledge-based approach in designing combinatorial or medicinal chemistry libraries for drug discovery. 1. A qualitative and quantitative characterization of known drug databases. J. Comb. Chem. 1, 55–68 (1999).
Petit, J., Meurice, N., Kaiser, C. & Maggiora, G. Softening the Rule of Five—where to draw the line? Bioorg. Med. Chem. 20, 5343–5351 (2012).
Lipinski, C. A. Rule of Five in 2015 and beyond: target and ligand structural limitations, ligand chemistry structure and drug discovery project decisions. Adv. Drug Deliv. Rev. 101, 34–41 (2016).
Yang, N. J. & Hinner, M. J. Getting across the cell membrane: an overview for small molecules, peptides, and proteins. Methods Mol. Biol. 1266, 29–53 (2015).
Bindels, D. S. et al. mScarlet: a bright monomeric red fluorescent protein for cellular imaging. Nat. Methods 14, 53 (2016).
Iizuka, R., Yamagishi, M. & Funatsu, T. Kinetic study of de novo chromophore maturation of fluorescent proteins. Anal. Biochem. 414, 173–178 (2011).
Mangan, S., Itzkovitz, S., Zaslaver, A. & Alon, U. The incoherent feed-forward loop accelerates the response-time of the gal system of Escherichia coli. J. Mol. Biol. 356, 1073–1081 (2006).
Paige, J. S., Wu, K. Y. & Jaffrey, S. R. RNA mimics of green fluorescent protein. Science 333, 642–646 (2011).
Eldar, A. & Elowitz, M. B. Functional roles for noise in genetic circuits. Nature 467, 167–173 (2010).
Chambers, I. et al. Nanog safeguards pluripotency and mediates germline development. Nature 450, 1230–1234 (2007).
Hoffmann, M. et al. Noise-driven stem cell and progenitor population dynamics. PLoS ONE 3, e2922 (2008).
Ozbudak, E. M., Thattai, M., Lim, H. N., Shraiman, B. I. & van Oudenaarden, A. Multistability in the lactose utilization network of Escherichia coli. Nature 427, 737–740 (2004).
Hildebrand, A., Pohl, M. & Bhakdi, S. Staphylococcus aureus alpha-toxin. Dual mechanism of binding to target cells. J. Biol. Chem. 266, 17195–17200 (1991).
Müller, P. & Schier, A. F. Extracellular movement of signaling molecules. Dev. Cell 21, 145–158 (2011).
Acknowledgements
This work was supported by the European Research Council (grant agreement no. 694410, AEDNA) and the DFG Cluster of Excellence Nanosystems Initiative Munich (DFG EXC 4/3). A.D. acknowledges additional support by the DFG Research Training Group ‘Chemical Foundations of Synthetic Biology’ GRK 2062/1. The authors thank B. Tinao for her preliminary work on the artificial multicellular assemblies. They also thank D. Ziegler and J. List for their help with set-up construction, S. Sagredo and E. Falgenhauer for providing purified enzymes, and M. Schwarz-Schilling for helpful discussions. Correspondence and request for materials should be addressed to F.C.S.
Author information
Authors and Affiliations
Contributions
A.D. and F.C.S. designed the experiments and wrote the manuscript. A.D. performed the experiments and analysed the data. A.D. and F.C.S. performed the modelling.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary methods, text, figures, tables, videos and references
Supplementary Video 1
This video shows the time lapse of Fig. 1b–c
Supplementary Video 2
Time lapse of the diffusion of DFHBI into a network of receivers containing a Spinach RNA transcription
Supplementary Video 3
This video shows 6 arrays containing the pulse propagation circuit, with different numbers of receivers
Supplementary Video 4
These videos show 2D networks containing the pulse generator
Supplementary Video 5
This video shows specifically a time-lapse of Fig. 3g
Supplementary Video 6
This video shows a differentiating network of one sender (indicated by S) and 4 receivers containing the circuit described in Fig. 4b, left panel
Rights and permissions
About this article
Cite this article
Dupin, A., Simmel, F.C. Signalling and differentiation in emulsion-based multi-compartmentalized in vitro gene circuits. Nature Chem 11, 32–39 (2019). https://doi.org/10.1038/s41557-018-0174-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-018-0174-9
This article is cited by
-
DNA as a universal chemical substrate for computing and data storage
Nature Reviews Chemistry (2024)
-
Superstructural ordering in self-sorting coacervate-based protocell networks
Nature Chemistry (2024)
-
Engineering cellular communication between light-activated synthetic cells and bacteria
Nature Chemical Biology (2023)
-
A simple method to make, trap and deform a vesicle in a gel
Scientific Reports (2023)
-
Dissipative DNA nanotechnology
Nature Chemistry (2022)