Abstract
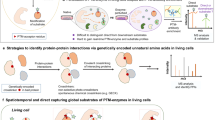
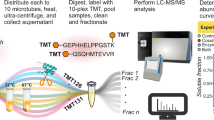
Pyridoxal phosphate (PLP) is an enzyme cofactor required for the chemical transformation of biological amines in many central cellular processes. PLP-dependent enzymes (PLP-DEs) are ubiquitous and evolutionarily diverse, making their classification based on sequence homology challenging. Here we present a chemical proteomic method for reporting on PLP-DEs using functionalized cofactor probes. We synthesized pyridoxal analogues modified at the 2′-position, which are taken up by cells and metabolized in situ. These pyridoxal analogues are phosphorylated to functional cofactor surrogates by cellular pyridoxal kinases and bind to PLP-DEs via an aldimine bond which can be rendered irreversible by NaBH4 reduction. Conjugation to a reporter tag enables the subsequent identification of PLP-DEs using quantitative, label-free mass spectrometry. Using these probes we accessed a significant portion of the Staphylococcus aureus PLP-DE proteome (73%) and annotate uncharacterized proteins as novel PLP-DEs. We also show that this approach can be used to study structural tolerance within PLP-DE active sites and to screen for off-targets of the PLP-DE inhibitor d-cycloserine.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Supplementary information, chemical characterization and proteomic data analysis are available in the online version of the paper. Crystallographic data of alanine racemase structures have been deposited in the Protein Data Bank68 (www.rcsb.org) under PDB codes 6G56, 6G58 and 6G59. The proteomic MS data (raw data and MaxQuant output tables for protein groups and peptides) have been deposited in the ProteomeXchange Consortium69 (http://proteomecentral.proteomexchange.org) via the PRIDE70 partner repository (data set identifier: PXD006483).
References
Amadasi, A. et al. Pyridoxal 5′-phosphate enzymes as targets for therapeutic agents. Curr. Med. Chem. 14, 1291–1324 (2007).
Salvo, Di, M. L, Budisa, N. & Contestabile, R. Beilstein Bozen Symposium on Molecular Engineering and Control. (Beilstein Institut: Frankfurt am Main, 2013) 27–66.
Eliot, A. C. & Kirsch, J. F. Pyridoxal phosphate enzymes: mechanistic, structural, and evolutionary considerations. Annu. Rev. Biochem. 73, 383–415 (2004).
Toney, M. D. Reaction specificity in pyridoxal phosphate enzymes. Arch. Biochem. Biophys. 433, 279–287 (2005).
Percudani, R. & Peracchi, A. The B6 database: a tool for the description and classification of vitamin B6-dependent enzymatic activities and of the corresponding protein families. BMC Bioinformatics 10, 273 (2009).
Percudani, R. & Peracchi, A. A genomic overview of pyridoxal-phosphate-dependent enzymes. EMBO Rep. 4, 850–854 (2003).
Kappes, B., Tews, I., Binter, A. & Macheroux, P. PLP-dependent enzymes as potential drug targets for protozoan diseases. Biochim. Biophys. Acta 1814, 1567–1576 (2011).
Mehta, P. K. & Christen, P. The molecular evolution of pyridoxal-5′-phosphate-dependent enzymes. Adv. Enzymol. Relat. Areas Mol. Biol. 74, 129–184 (2000).
Catazaro, J., Caprez, A., Guru, A., Swanson, D. & Powers, R. Functional evolution of PLP-dependent enzymes based on active-site structural similarities. Proteins 82, 2597–2608 (2014).
Denessiouk, K. A., Denesyuk, A. I., Lehtonen, J. V., Korpela, T. & Johnson, M. S. Common structural elements in the architecture of the cofactor-binding domains in unrelated families of pyridoxal phosphate-dependent enzymes. Proteins 35, 250–261 (1999).
Fleischman, N. M. et al. Molecular characterization of novel pyridoxal-5′-phosphate-dependent enzymes from the human microbiome. Protein Sci. 23, 1060–1076 (2014).
Whittaker, M. M., Penmatsa, A. & Whittaker, J. W. The Mtm1p carrier and pyridoxal 5′-phosphate cofactor trafficking in yeast mitochondria. Arch. Biochem. Biophys. 568, 64–70 (2015).
Simon, E. S. & Allison, J. Determination of pyridoxal-5′-phosphate (PLP)-bonding sites in proteins: a peptide mass fingerprinting approach based on diagnostic tandem mass spectral features of PLP-modified peptides. Rapid Commun. Mass Spectrom. 23, 3401–3408 (2009).
Wu, Y Chen, J., Liu, Z. & Wang, F. Identification of pyridoxal phosphate modified proteins using mass spectrometry. Rapid Commun. Mass Spectrom. 32, 195–200 2018).
Schnackerz, K. D. & Cook, P. F. Resolution of pyridoxal 5′-phosphate from O-acetylserine sulfhydrylase from Salmonella typhimurium and reconstitution of apoenzyme with cofactor and cofactor analogues as a probe of the cofactor binding site. Arch. Biochem. Biophys. 324, 71–77 (1995).
Morino, Y. & Snell, E. E. Coenzyme activity of homologues of pyridoxal phosphate. Biochemistry 57, 1692–1699 (1967).
Mechanik, M. L., Torchinsky, Y. M., Florentiev, V. L. & Karpeisky, M. Y. Interaction of the apoenzyme of l-glutamate decarboxylase with pyridoxal phosphate analogues. FEBS Lett. 13, 177–180 (1971).
Cravatt, B. F., Wright, A. T. & Kozarich, J. W. Activity-based protein profiling: from enzyme chemistry to proteomic chemistry. Annu. Rev. Biochem. 77, 383–414 (2008).
Korytnyk, W., Srivastava, S. C., Angelino, N., Potti, P. G. & Paul, B. A general method for modifying the 2-methyl group of pyridoxol. Synthesis and biological activity of 2-vinyl- and 2-ethynylpyridoxols and related compounds. J. Med. Chem. 16, 1096–1101 1973).
Kaiser, E. M. et al. Regiointegrity of carbanions derived by selective metalations of dimethylpyridines and -quinolines. J. Organomet. Chem. 213, 405–417 (1981).
Kim, Y.-C. & Jacobson, K. A. Versatile synthesis of 6-alkyl and aryl substituted pyridoxal derivatives. Synthesis 1, 119–122 (2000).
Korytnyk, W. & Ahrens, H. 5-homopyridoxals, 5-thiopyridoxal, and related compounds. Synthesis, tautomerism, and biological properties. J. Med. Chem. 14, 947–952 1971).
di Salvo, M. L., Contestabile, R. & Safo, M. K. Vitamin B6 salvage enzymes: mechanism, structure and regulation. Biochim. Biophys. Acta 1814, 1597–1608 (2011).
Denesyuk, A. I., Denessiouk, K. A., Korpela, T. & Johnson, M. S. Functional attributes of the phosphate group binding cup of pyridoxal phosphate-dependent enzymes. J. Mol. Biol. 316, 155–172 (2002).
Nodwell, M. B., Koch, M. F., Alte, F., Schneider, S. & Sieber, S. A. A subfamily of bacterial ribokinases utilizes a hemithioacetal for pyridoxal phosphate salvage. J. Am. Chem. Soc. 136, 4992–4999 (2014).
Strominger, J. L., Ito, I. & Threnn, R. H. Competitive inhibition of enzymatic reactions by oxamycin. J. Am. Chem. Soc. 82, 998–999 (1960).
Griswold, W. R. & Toney, M. D. Role of the pyridine nitrogen in pyridoxal 5′-phosphate catalysis: activity of three classes of PLP enzymes reconstituted with deazapyridoxal 5′-phosphate. J. Am. Chem. Soc. 133, 14823–14830 (2011).
Scaletti, E. R., Luckner, S. R. & Krause, K. L. Structural features and kinetic characterization of alanine racemase from Staphylococcus aureus (Mu50). Acta Crystallogr. D 68, 82–92 (2012).
Rostovtsev, V. V., Green, L. G., Fokin, V. V. & Sharpless, K. B. A stepwise huisgen cycloaddition process: copper(I)-catalyzed regioselective ‘ligation’ of azides and terminal alkynes. Angew. Chem. Int. Ed. 41, 2596–2599 (2002).
Speers, A. E. & Cravatt, B. F. Profiling enzyme activities in vivo using click chemistry methods. Chem. Biol. 11, 535–546 (2004).
Tornoe, C. W., Christensen, C. & Meldal, M. Peptidotriazoles on solid phase: [1,2,3]-triazoles by regiospecific copper(I)-catalyzed 1,3-dipolar cycloadditions of terminal alkynes to azides. J. Org. Chem. 67, 3057–3064 (2002).
Mukherjee, T., Hanes, J., Tews, I., Ealick, S. E. & Begley, T. P. Pyridoxal phosphate: biosynthesis and catabolism. Biochim. Biophys. Acta 1814, 1585–1596 (2011).
Fey, P. D.et al. A genetic resource for rapid and comprehensive phenotype screening of nonessential Staphylococcus aureus genes. mBio 4, e00537-12 (2013).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell. Proteomics. 13, 2513–2526 (2014).
Saxon, E. & Bertozzi, C. R. Cell surface engineering by a modified Staudinger reaction. Science 287, 2007–2010 (2000).
Ito, T. et al. Conserved pyridoxal protein that regulates Ile and Val metabolism. J. Bacteriol. 195, 5439–5449 (2013).
Prunetti, L. et al. Evidence that COG0325 proteins are involved in PLP homeostasis. Microbiology 162, 694–706 (2016).
Mozzarelli, A. & Bettati, S. Exploring the pyridoxal 5′-phosphate-dependent enzymes. Chem. Rec. 6, 275–287 (2006).
Finn, R. D. et al. InterPro in 2017—beyond protein family and domain annotations. Nucleic Acids Res. 45, D190–D199 (2017).
Kuhn, M. L., Majorek, K. A., Minor, W. & Anderson, W. F. Broad-substrate screen as a tool to identify substrates for bacterial Gcn5-related N-acetyltransferases with unknown substrate specificity. Protein Sci. 22, 222–230 (2013).
Mukherjee, J. J. & Dekker, E. E. Purification, properties, and N-terminal amino acid sequence of homogeneous Escherichia coli 2-amino-3-ketobutyrate CoA ligase, a pyridoxal phosphate-dependent enzyme. J. Biol. Chem. 262, 14441–14447 (1987).
Ye, Q. Z., Liu, J. & Walsh, C. T. p-Aminobenzoate synthesis in Escherichia coli: purification and characterization of PabB as aminodeoxychorismate synthase and enzyme X as aminodeoxychorismate lyase. Proc. Natl Acad. Sci. USA 87, 9391–9395 (1990).
Soper, T. S. & Manning, J. M. Different modes of action of inhibitors of bacterial d-amino acid transaminase. A target enzyme for the design of new antibacterial agents. J. Biol. Chem. 256, 4263–4268 (1981).
Malashkevich, V. N., Strop, P., Keller, J. W., Jansonius, J. N. & Toney, M. D. Crystal structures of dialkylglycine decarboxylase inhibitor complexes. J. Mol. Biol. 294, 193–200 (1999).
Yew, W. W., Wong, C. F., Wong, P. C., Lee, J. & Chau, C. H. Adverse neurological reactions in patients with multidrug-resistant pulmonary tuberculosis after coadministration of cycloserine and ofloxacin. Clin. Infect. Dis. 17, 288–289 (1993).
Caminero, J. A., Sotgiu, G., Zumla, A. & Migliori, G. B. Best drug treatment for multidrug-resistant and extensively drug-resistant tuberculosis. Lancet Infect. Dis. 10, 621–629 (2010).
Neuhaus, F. C. Selective inhibition of enzymes utilizing alanine in the biosynthesis of peptidoglycan. Antimicrob. Agents Chemother. 7, 304–313 (1967).
Peisach, D., Chipman, D. M., Van Ophem, P. W., Manning, J. M. & Ringe, D. d-Cycloserine inactivation of d-amino acid aminotransferase leads to a stable noncovalent protein complex with an aromatic cycloserine-PLP derivative. J. Am. Chem. Soc. 120, 2268–2274 (1998).
Fenn, T. D., Stamper, G. F., Morollo, A. A. & Ringe, D. A side reaction of alanine racemase: transamination of cycloserine. Biochemistry 42, 5775–5783 (2003).
Sieradzki, K. & Tomasz, A. Suppression of beta-lactam antibiotic resistance in a methicillin-resistant Staphylococcus aureus through synergic action of early cell wall inhibitors and some other antibiotics. J. Antimicrob. Chemother. 39, 47–51 (1997).
Roze, U. & Strominger, J. L. Alanine racemase from Staphylococcus aureus: conformation of its substrates and its inhibitor, d-cycloserine. Mol. Pharm. 2, 92–94 (1966).
Lambert, M. P. & Neuhaus, F. C. Mechanism of d-cycloserine action: alanine racemase from Escherichia coli W. J. Bacteriol. 110, 978–987 (1972).
Prosser, G. A. & de Carvalho, L. P. Metabolomics reveal d-alanine:d-alanine ligase as the target of d-cycloserine in Mycobacterium tuberculosis. ACS Med. Chem. Lett. 4, 1233–1237 (2013).
Halouska, S. et al. Metabolomics analysis identifies d-alanine-d-lanine ligase as the primary lethal target of d-cycloserine in mycobacteria. J. Proteome Res. 13, 1065–1076 (2014).
Contestabile, R. et al. l-Threonine aldolase, serine hydroxymethyltransferase and fungal alanine racemase. A subgroup of strictly related enzymes specialized for different functions. Eur. J. Biochem. 268, 6508–6525 (2001).
di Salvo, M. L. et al. Alanine racemase from Tolypocladium inflatum: a key PLP-dependent enzyme in cyclosporin biosynthesis and a model of catalytic promiscuity. Arch. Biochem. Biophys. 529, 55–65 (2013).
Takenaka, T., Ito, T., Miyahara, I., Hemmi, H. & Yoshimura, T. A new member of MocR/GabR-type PLP-binding regulator of d-alanyl-d-alanine ligase in Brevibacillus brevis. FEBS J. 282, 4201–4217 (2015).
Kleiner, P., Heydenreuter, W., Stahl, M., Korotkov, V. S. & Sieber, S. A. A whole proteome inventory of background photocrosslinker binding. Angew. Chem. Int. Ed. 56, 1396–1401 (2017).
Anderson, L. N. et al. Live cell discovery of microbial vitamin transport and enzyme–cofactor interactions. ACS Chem. Biol. 11, 345–354 (2016).
Broncel, M., Serwa, R. A., Bunney, T. D., Katan, M. & Tate, E. W. Global profiling of Huntingtin-associated protein E (HYPE)-mediated AMPylation through a chemical proteomic approach. Mol. Cell. Proteomics 15, 715–725 (2016).
Grammel, M. & Hang, H. C. Chemical reporters for biological discovery. Nat. Chem. Biol. 9, 475–484 (2013).
Romine, M. F. et al. Elucidation of roles for vitamin B12 in regulation of folate, ubiquinone, and methionine metabolism. Proc. Natl Acad. Sci. USA 114, E1205–E1214 (2017).
Westcott, N. P., Fernandez, J. P., Molina, H. & Hang, H. C. Chemical proteomics reveals ADP-ribosylation of small GTPases during oxidative stress. Nat. Chem. Biol. 13, 302–308 (2017).
Wright, M. H. et al. Validation of N-myristoyltransferase as an antimalarial drug target using an integrated chemical biology approach. Nat. Chem. 6, 112–121 (2014).
Eirich, J. et al. Pretubulysin derived probes as novel tools for monitoring the microtubule network via activity-based protein profiling and fluorescence microscopy. Mol. Biosyst. 8, 2067–2075 (2012).
Tyanova, S., Temu, T. & Cox, J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat. Protoc. 11, 2301–2319 (2016).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Vizcaino, J. A. et al. ProteomeXchange provides globally coordinated proteomics data submission and dissemination. Nat. Biotechnol. 32, 223–226 (2014).
Vizcaino, J. A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44, 11033 (2016).
Acknowledgements
This project received funding from the European Research Council (ERC) and the European Union’s Horizon 2020 research and innovation programme (grant agreement no. 725085, CHEMMINE, ERC consolidator grant). Further financial support includes doctoral scholarships to A.H. from the Deutscher Akademischer Austausch Dienst (DAAD) and to M.S. from the Studienstiftung des Deutschen Volkes. The authors thank the Network on Antimicrobial Resistance in Staphylococcus aureus (NARSA) for the supply of the Nebraska Transposon Mutant Library (NTML). The authors also thank M. Wolff and K. Bäuml for technical assistance, the Swiss Light Source (SLS) and European Synchrotron Radiation Facility (ESRF) for beam time, and the staff of beamlines PX I (SLS), ID23-2 and ID29 (ESRF) for setting up of the beamlines for data collection. The authors thank M. H. Wright and B. M. Williams for critical proofreading of the manuscript.
Author information
Authors and Affiliations
Contributions
A.H., M.B.N. and S.A.S. conceived and designed the project. A.H., M.B.N. and M.P. synthesized PL probes. A.H., M.B.N. and M.P. conducted biochemical characterization of probes and PLP-DEs, including purification of recombinant proteins, enzyme kinetics assays, UV–vis measurements and in vitro protein labelling experiments (MS and gel-based). S.S. performed crystallization and determined X-ray structures of Alr. A.H. completed cell-based labelling experiments and growth curves. V.C.K. prepared and analysed targeted metabolomic samples and characterized JW8. A.H. designed, executed and analysed proteomic experiments. N.C.B. and M.S. contributed proteomics expertise and analysed PLP binding sites. A.H. and S.A.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Data, Supplementary Schemes, Supplementary Figures and Supplementary Methods and Protocols
Proteomics tables and binding-site identification
Supplementary proteomics tables and identification of binding-sites
Rights and permissions
About this article
Cite this article
Hoegl, A., Nodwell, M.B., Kirsch, V.C. et al. Mining the cellular inventory of pyridoxal phosphate-dependent enzymes with functionalized cofactor mimics. Nature Chem 10, 1234–1245 (2018). https://doi.org/10.1038/s41557-018-0144-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-018-0144-2
This article is cited by
-
Bioorthogonally activatable cyanine dye with torsion-induced disaggregation for in vivo tumor imaging
Nature Communications (2022)
-
Underground metabolism facilitates the evolution of novel pathways for vitamin B6 biosynthesis
Applied Microbiology and Biotechnology (2021)
-
Oxygen reactivity with pyridoxal 5′-phosphate enzymes: biochemical implications and functional relevance
Amino Acids (2020)