Abstract
Activated sludge (AS) microbial communities were analyzed for seasonal variation, a disturbance-recovery event, and separated small aggregates (SAG) to study the influent immigration effect using both neutral immigration model and mass-balance model with operational parameters. SAG differed with AS, and higher immigration impact on SAG was confirmed by both models. Adding the SAG community segregation in the latter model to evaluate the contribution of influent immigration to community disturbance-recovery showed increased impact of immigration.
Similar content being viewed by others
Introduction
The activated sludge (AS) wastewater treatment plant provides a well-defined model system to study microbial ecology. The AS community is shaped by deterministic (Niche) factors and stochastic (Neutral) factors (e.g., neutral immigration from wastewater influent)1. Two types of numerical models have been used to quantify the influent immigration effect. One is the neutral community (immigration) model (NIM)2, employing the abundance and frequency of operational taxonomic units (OTUs) in metacommunities to calculate the community-level migration probability1,3,4,5,6. The other is mass-balance model (MBM) that defines the immigration as the proportion of reactor biomass derived from influent biomass \(\left( {\frac{{{\mathrm{influent}}\;{\mathrm{biomass}}}}{{{\mathrm{influent}}\;{\mathrm{biomass}} + {\mathrm{local}}\;{\mathrm{growth}}}}} \right)\), using abundances of specific OTUs to calculate the immigration rates and net growth rates7. The NIM immigration describes the probability that a dead individual in the reactor is replaced by an immigrant from the influent, differentiated from the reproduction of local community. Intrinsically, the MBM bases on the fractionation of influent biomass and local growth, which also aims to differentiate the influent immigration and local community. The MBM has been increasingly used because of the quantitative measurements of the OTU growth activity8. Although steady-state metacommunities demonstrated that operational parameters (e.g., solid retention time, SRT) are important for the evaluation of immigration4,9, the operation parameters were not directly employed in the original MBM. Recently, Frigon and Wells10 and Guo11 developed the mass-flow immigration model, which is a MBM incorporating operational parameters (MBM-OP), to calculate the immigration rate of specific OTUs. By using operational parameters, this model is comprehensible for both wastewater engineers and microbiologists.
In wastewater treatment bioreactors, community variation has been shown in relation to the size and status (settled and unsettled) of the aggregates12,13. In conventional AS, the relative importance of influent immigration on different-sized aggregate communities (i.e., easily settleable large-size flocs and low-settleability small-size particles) has not been explored. Besides steady-state modeling, the bioreactors often experience disturbances9,14, such as sudden high flow rates which lead to a biomass washout. The non-steady-state community serves as a good example to study the community recovery and to compare the contributions of local community growth and influent immigration.
Our study aimed at monitoring the long-term variation of AS communities in different seasons and comparing the influent immigration effect on different-sized AS communities using NIM and MBM-OP models. In addition, the short-term non-steady-state (under operational high-flow disturbance) AS community’s recovery attributed to local community growth and influent immigration was evaluated.
AS community variation
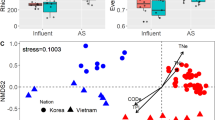
The AS communities (between 2013 and 2019) showed that abundant microorganisms were relatively consistent during the long-term observation, and consisted of a core group at the order level (Fig. 1b) and at genus level (Supplementary Fig. S1). Seasonal changes and disturbance did not alter the core group14. The variation between small aggregates (SAG) and AS was clearly shown (Fig. 1b), being consistent in different seasons. The most abundant members in SAG were different from AS communities, but similar with the influent community’s most abundant members, indicating that influent microbiome is an important origin for the SAG. The community richness (number of observed OTUs) was slightly higher in SAG than in AS (Supplementary Fig. S2), inferring that the SAG community was a union of AS and influent communities. The shared numbers and abundances of OTUs are indicators of immigration effect7,9,15. A large number of OTUs were shared among the influent, AS, and SAG (Fig. 1c). The shared OTUs between influent and AS took up 89.1 ± 0.1% of the influent community total number of reads and 54.0 ± 4.3% of the AS community total number of reads, which increased between influent and SAG (97.0 ± 0.1% and 73.8 ± 3.0%, respectively), indicating higher immigration effect for influent–SAG than for influent–AS. The Bray-Curtis distances also showed higher similarity between influent and SAG than between influent and AS (Fig. 1d). Permutational multivariate analysis of variance (Bray-Curtis distance, permutation = 999) was performed to test the two-way factor effect of sampling time and aggregate size. Our results showed significant effect of aggregated size (p = 0.001), but not sampling time (p = 0.911) or their interaction (p = 0.866).
a Schematic chart of sample collection. b Assembly of the top 10 abundant bacterial orders from influent and AS. c Venn diagram of shared OTUs among influent, activated sludge (AS) and small aggregates (SAG), and the relative abundances of shared OTUs. d PCoA of Bray-Curtis distances among the set of seasonal AS communities and high-flow disturbed and recovered communities (pyrosequencing), and the size-separated bioreactor samples (Illumina Miseq sequencing); variation between the long-term data and current data is likely due to different sequencing technologies. e Quantified influent immigration to AS and SAG using neutral immigration model (mNIM) and mass-balance model with operational parameters (mi,MBM-OP).
This phenomenon of community segregation was observed in different forms of biomass in biofilm and granular sludge reactors13,16,17. Less work has been reported for conventional AS. SAG have higher accessibility to substrates and lower retention time and are more subjected to influent immigration effect as revealed in our results. An ecological growth trait of community, weighted rrn gene copy number per genome, showed that SAG has higher average rrn than AS, implying possibly higher growth activity (Supplementary Fig. S3)14,18,19.
The influent immigration impact was quantified using the NIM2,3 and MBM-OP (Eq. 14) models. The migration probability (mNIM) was higher for influent–SAG (0.40) than for influent–AS (0.33) (Fig. 1e). The distribution of immigration rates for each OTU based on MBM-OP (mi,MBM-OP) showed that influent–SAG shifted towards higher immigration rate compared to influent–AS (Fig. 1e), in accordance with the observation of mNIM. These observations suggest a neglected fact that the influent immigration to bioreactors is associated with the aggregate size-differentiated community. The SAG community is less settleable compared to flocs and more subjected to washing out from the reactor, leading to higher dependency on the influent immigration. Moreover, the influent community that enters the reactor and the SAG community share a more similar suspension status compared to flocs, which may attribute to the higher community similarity (Fig. 1d) and immigration (Fig. 1e).
Immigration contribution to community recovery
The AS community was disturbed by a high-flow event which led to a drop in biomass density, shifting away from the steady-state community and recovered after 3 days (Fig. 1d). The recovery could be attributed to different mechanisms, including growth of the indigenous microorganisms9,15 and influent immigration1,2,9.
Using the MBM-OP immigration model, the influent community can be partitioned to growth and neutral immigration (non-growth) (Fig. 2a). A conceptual model MBM-OP-SAG was constructed (Fig. 2c). In both models, OTUs were classified into three groups based on their relative abundances in the influent and in the AS: (i) not in influent but in reactor community, (ii) growth group (calculated net growth rate ui,MBM-OP > 0, Eq. 16), (iii) complete immigration group (calculated net growth rate ui,MBM-OP ≤ 0, Eq. 16). The relative abundances (Fig. 2b, d) showed that the immigration group was using the MOM-OP-SAG model more (Fig. 2d) than using the MBM-OP-AS model (Fig. 2b). For the disturbed and recovered AS communities, the MBM-OP-AS model showed similar abundances of immigration group (Fig. 2b). The MBM-OP-SAG model estimated higher contribution of immigration in recovered than in disturbed community (Fig. 2d). The two models differed in certain taxa’s growth rates and immigration rates, resulting in a list of genera classified into growth group by MBM-OP-AS model while into immigration group by MBM-OP-SAG model (Supplementary Table S1). Microorganisms such as Methanobacterium, Methanobrevibacter, Clostridiales, Synergistales, and Bifidobacterium are known anaerobic microorganisms and likely to be carried by wastewater from sewer20 and not to grow in AS. The MBM-OP-SAG model was successful to identify these genera as influent immigration microorganisms.
a Influent to AS immigration model. b Proportions of the immigration and growth groups calculated by MBM-OP-AS model using the relative abundances of OTUs in influent and AS. c Proposed influent to small aggregates immigration model. d Proportions of the immigration and growth groups calculated by MBM-OP-SAG model using the relative abundances of OTUs in influent and small aggregates. Error bar represents standard deviation of n samples (n = 2 for influent; n = 6 for SAG and AS; n = 2 for disturbed and recovered).
Our results demonstrated AS community variation in different-sized aggregates. NIM and MBM-OP models were compared, both resulted in higher immigration impact for the small aggregate (SAG) community than for the AS community. The different-sized community variation led to an approach (MBM-OP-SAG model) of modeling influent and segregated communities for immigration effects, which was applied to investigate the community recovery. The MBM-OP-SAG model showed higher immigration contribution to recovery than the MBM-OP-AS model. This approach could be applied to other bioreactor systems to increase modeling accuracy of microbial growth and immigration, compared to the conventional NIM and MBM-OP models.
Methods
AS and influent biomass sampling from full-scale wastewater treatment plant
Influent (treated by primary clarifier) and AS samples under various flow and seasonal conditions were taken from a full-scale biological nutrient removal wastewater treatment plant with a capacity of treating 310 MLD in Edmonton, Canada. For long-term seasonal comparison, AS samples were collected spanning 6 years (2013–2019), which covers seasonal variations and various flow conditions (240–310 MLD). A high-flow event resulted in a decrease of mixed liquor volatile suspended solids from 2840 to 2200 mg/L. The most recent two AS samples (Fall 2018 and Spring 2019) were used to analyze different-sized AS communities. AS samples were taken from the aeration tank to the on-site lab and immediately settled for 30 min using a stir tank at 5 rpm. The supernatant (containing SAG), the interface, and the settled AS were transferred to 50 mL sterile centrifuge tubes and stored on ice before centrifugation (within 1 h). The particle size in the supernatant was determined using Zetasizer (model ZEN 1600, Malvern Instruments Ltd.), resulting in a median size of 25 ± 4.5 µm and range of 1.9–29.0 µm.
Influent samples and small aggregate samples were centrifuged in 50 mL tubes at 5000g for 10 min to collect the pellets. The AS samples were centrifuged in 2 mL tubes at 13,000g for 2 min. The samples were stored at −20 °C until DNA extraction.
DNA extraction, 16S rRNA gene amplicon sequencing
DNA was extracted from influent and bioreactor samples using DNeasy PowerSoil Kit (Qiagen Inc., Toronto, Canada) according to the manufacturer’s protocol. The long-term monitoring of the community was conducted using pyrosequencing of the 16S rRNA genes amplified by 28F (5′-GAGTTTGATCNTGGCTCAG- 3′) and 519R (5′-GTNTTACNGCGGCKGCTG-3′). The different-sized AS and influent communities were analyzed using Illumina sequencing. The extracted DNA was used for PCR with modified primers with adaptors: 515 forward primer (5′- ACA CTG ACG ACA TGG TTC TAC AGT GYC AGC MGC CGC GGT AA −3′) and 806 reverse primer (5′- TAC GGT AGC AGA GAC TTG GTC TGG ACT ACN VGG GTW TCT AAT −3′)21,22, targeting the V4 region of 16S rRNA genes. The PCR was performed under the following conditions: primer concentration at 0.3 µm, Mg2+ at 1.5 mM, dNTP at 200 µm, Taq DNA polymerase at 1.25 U/50 µL reaction (New England Biolabs, Whitby, ON, Canada), initial denaturation at 94 °C for 3 min, then 25 cycles of denaturation at 94 °C for 30 s, annealing at 62 °C for 45 s and elongation at 68 °C for 1 min, then final elongation at 68 °C for 10 min. The PCR products were purified using QIAquick PCR Purification Kit (Qiagen, Toronto, ON, Canada), then sent for barcoding and sequencing at McGill University and Génome Québec Innovation Centre (Montréal, QC, Canada) using Illumina MiSeq PE250 platform. Sequence data were deposited to the National Center for Biotechnology Information (NCBI) GenBank (Bio Project Accession number PRJNA580466).
The raw sequences (2 × 250 nt) were paired, quality-filtered using DADA223 in Qiime2 pipelines24. The OTUs were defined using the default 100% similarity. The taxonomic identification was assigned using GreenGenes (version gg_13_8) reference database at 99% similarity25,26. Microbial community diversities (alpha-diversity, beta-diversity) were analyzed using R “vegan” package27. The weighted rrn copy number was estimated by the normalized OTU table using PICRUSt28 with metagenome references in Integrated Microbial Genomes (IMG) system29.
where f16S_norm,i is the relative abundance of the ith OTU in total 16S rRNA gene amplicon sequence reads normalized by PICRUSt; Nrrn,i is the rrn copy number of the ith OTU in the IMG database.
Neutral immigration model
The probability of influent immigration was defined by Sloan et al2. and calculated using the neutral immigration model. The model described that in a microbial community with NT individuals, an individual die or leave the system and is immediately replaced by an immigrant with probability of m, or by reproduction of a local member of the community with probability of 1–m.
For the ith species (initial number Ni), the probability (Pr) of increase by one, stay the same, or decrease by one is expressed, respectively, by the following three equations:
where pi is the relative abundance of the ith species, αi is the advantage parameter of the ith species over the other species.
The relative abundance can be expressed as \(x_i = \frac{{N_i}}{{N_T}}\). Assume that NT is large enough and xi becomes continuous, and αi = 0 so the species has no advantages, then the probability density function of xi,ϕi=(xi,t) can be expressed as (for more details please see Sloan et al2.):
where c is a constant so that \({\int\limits_0^1} \phi _i(x_i)dx_i = 1\).
The probability of the ith species observed in any local community is:
where d is the detection threshold.
This equation can be used to calibrate community data (frequency and mean relative abundance plot) and solve for m.
AS and SAG communities included six replicates each. The immigration rate m was estimated for influent and AS communities and influent and SAG communities, using the R codes from Burns et al3. with 100 permutations (R code in Supplementary Information).
Mass-balance model with operational parameters
The MBM with operational parameters (mass-flow immigration model) calculation for net growth rates was described by Guo11.
The mixed liquor heterotrophic biomass is contributed from the influent biomass and the microbial growth from the consumption of substrate resources. The immigration is defined as the fraction of influent biomass contribution:
where m is the mass-flow immigration rate; XOHO,Inf and XOHO,ML are the active heterotrophic biomass in influent and mixed liquor; fOHO,Capt is the fraction of influent biomass captured by the mixed liquor which is default 1;YOHO is the heterotrophic growth yield; Sconsumed is the consumed substrates.
For continuous stirred tank reactors, the heterotrophic biomass based on mass-balance calculation30 is:
Rearrangement gives:
Under the assumption that \(f_{{\mathrm{OHO,Capt}}} \ast X_{{\mathrm{OHO,Inf}}} \ll Y_{{\mathrm{OHO}}} \ast S_{{\mathrm{consumed}}}\), substitution of Eq. 10 to Eq. 8 gives:
The immigration rate of a specific ith OTU is:
The proportion of the ith OTU in the biomass sample (f16S,i) can be quantified from the 16S rRNA amplicon sequencing data. The biomass concentration of the ith OTU can be defined by:
where XOHO,i is the biomass concentration of the ith OTU; XTot is the total concentration of solids; γDNA is the mass of DNA per solids; f16S,i is the proportion of the ith genus in total 16S rRNA gene amplicon sequence reads.
Therefore, the immigration rate of a specific ith OTU is expressed using 16S rRNA amplicon sequencing data:
Note that the maximum value of mi is 1, so the calculated mi values when greater than 1 were set to 1. When mi = 1 (i.e., the growth rate [μOHO,i] is 0):
The net growth rate equals the growth rate minus decay rate, therefore the net growth rate of the ith OTU is:
The OTUs then can be classified into three groups:
-
(i)
not in influent but in reactor community
-
(ii)
growth group (calculated net growth rate uOHO,Net,i > 0)
-
(iii)
complete immigration group (calculated net growth rate uOHO,Net,i ≤ 0).
Data availability
The raw sequence data of 16S rRNA gene amplicons can be accessed at GenBank (Bio Project Accession number PRJNA580466).
References
Ofiteru, I. D. et al. Combined niche and neutral effects in a microbial wastewater treatment community. Proc. Natl Acad. Sci. USA 107, 15345–15350 (2010).
Sloan, W. T. et al. Quantifying the roles of immigration and chance in shaping prokaryote community structure. Environ. Microbiol. 8, 732–740 (2006).
Burns, A. R. et al. Contribution of neutral processes to the assembly of gut microbial communities in the zebrafish over host development. ISME J. 10, 655–664 (2016).
Yuan, H., Mei, R., Liao, J. & Liu, W. T. Nexus of stochastic and deterministic processes on microbial community assembly in biological systems. Front. Microbiol. 10, 1536 (2019).
Mei, R., Kim, J., Wilson, F. P., Bocher, B. T. W. & Liu, W. T. Coupling growth kinetics modeling with machine learning reveals microbial immigration impacts and identifies key environmental parameters in a biological wastewater treatment process. Microbiome 7, 65 (2019).
Ning, D., Deng, Y., Tiedje, J. M. & Zhou, J. A general framework for quantitatively assessing ecological stochasticity. Proc. Natl Acad. Sci. USA 116, 16892–16898 (2019).
Saunders, A. M., Albertsen, M., Vollertsen, J. & Nielsen, P. H. The activated sludge ecosystem contains a core community of abundant organisms. ISME J. 10, 11–20 (2016).
Mei, R. & Liu, W. T. Quantifying the contribution of microbial immigration in engineered water systems. Microbiome 7, 144 (2019).
Vuono, D. C., Munakata-Marr, J., Spear, J. R. & Drewes, J. E. Disturbance opens recruitment sites for bacterial colonization in activated sludge. Environ. Microbiol. 18, 87–99 (2016).
Frigon, D. & Wells, G. Microbial immigration in wastewater treatment systems: analytical considerations and process implications. Curr. Opin. Biotechnol. 57, 151–159 (2019).
Guo, B. Cellular Metabolic Markers and Growth Dynamics Definition of Functional Groups in Activated Sludge Wastewater Treatment Heterotrophic Population. Doctor of Philosophy thesis, McGill Univ. (2019).
Sun, X., Sheng, Z. & Liu, Y. Effects of silver nanoparticles on microbial community structure in activated sludge. Sci. Total Environ. 443, 828–835 (2013).
Ali, M. et al. Importance of species sorting and immigration on the bacterial assembly of different-sized aggregates in a full-scale aerobic granular sludge plant. Environ. Sci. Technol. 53, 8291–8301 (2019).
Perez, M.V., Guerrero, L.D., Orellana, E., Figuerola, E.L. & Erijman, L. Time series genome-centric analysis unveils bacterial response to operational disturbance in activated sludge. mSystems 4(4), e00169–19, https://doi.org/10.1128/mSystems.00169-19 (2019).
Lee, S. H., Kang, H. J. & Park, H. D. Influence of influent wastewater communities on temporal variation of activated sludge communities. Water Res. 73, 132–144 (2015).
Laureni, M. et al. Biomass segregation between biofilm and flocs improves the control of nitrite-oxidizing bacteria in mainstream partial nitritation and anammox processes. Water Res. 154, 104–116 (2019).
Wells, G. F. et al. Microbial biogeography across a full-scale wastewater treatment plant transect: evidence for immigration between coupled processes. Appl. Microbiol. Biotechnol. 98, 4723–4736 (2014).
Klappenbach, J. A., Dunbar, J. M. & Schmidt, T. M. rRNA operon copy number reflects ecological strategies of bacteria. Appl. Environ. Microbiol. 66, 1328–1333 (2000).
Nemergut, D. R. et al. Decreases in average bacterial community rRNA operon copy number during succession. ISME J. 10, 1147–1156 (2016).
Guo, B., Liu, C., Gibson, C. & Frigon, D. Wastewater microbial community structure and functional traits change over short timescales. Sci. Total Environ. 662, 779–785 (2019).
Parada, A. E., Needham, D. M. & Fuhrman, J. A. Every base matters: assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples. Environ. Microbiol 18, 1403–1414 (2016).
Apprill, A., McNally, S., Parsons, R. & Weber, L. Minor revision to V4 region SSU rRNA 806R gene primer greatly increases detection of SAR11 bacterioplankton. Aquat. Microb. Ecol. 75, 129–137 (2015).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
McDonald, D. et al. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 6, 610–618 (2012).
Werner, J. J. et al. Impact of training sets on classification of high-throughput bacterial 16s rRNA gene surveys. ISME J. 6, 94–103 (2012).
Oksanen, F.J. et al. vegan: Community Ecology Package. R package Version 2.4-3. https://CRAN.R-project.org/package=vegan (2017).
Langille, M. G. et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 31, 814–821 (2013).
Markowitz, V. M. et al. IMG: the Integrated Microbial Genomes database and comparative analysis system. Nucleic Acids Res. 40, D115–D122 (2012).
Grady, C. P. L. & Grady, C. P. L. Biological Wastewater Treatment. 3rd edn. (Taylor & Francis, 2011).
Acknowledgements
This work was funded by the Canada Research Chair (CRC) in Future Water Services (Y.L.) and Natural Sciences and Engineering Research Council of Canada (NSERC) Discovery Grant (Y.L.).
Author information
Authors and Affiliations
Contributions
B.G. collected and analyzed the samples, carried out data analysis, modeling and wrote the manuscript. Z.S. analyzed the samples and carried out analysis. Y.L. conceived and guided the study and revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Guo, B., Sheng, Z. & Liu, Y. Evaluation of influent microbial immigration to activated sludge is affected by different-sized community segregation. npj Clean Water 4, 20 (2021). https://doi.org/10.1038/s41545-021-00112-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41545-021-00112-7