Abstract
Recently reported detection of SARS-CoV-2 in wastewater around the world has led to emerging concerns on potential risk in water bodies receiving treated wastewater effluent. This review aims to provide an up-to-date state of key knowledge on the impact of SARS-CoV-2 in natural water bodies receiving treated wastewater. In this review, SARS-CoV-2 concentrations in wastewater, expected removal in WWTPs, and possible dilution and decay in water bodies are reviewed based on past studies on SARS-CoV-2 and related enveloped viruses. We suggest a quantitative microbial risk assessment (QMRA) framework to estimate the potential risk of SARS-CoV-2 in natural water bodies through various water activities. Dose–response model of SARS-CoV and Poisson’s distribution is employed to estimate possible viral ingestion and the annual chance of infection through several water activities in natural water bodies. Finally, future perspectives and research needs have been addressed to overcome the limitations and uncertainty in the risk assessment of SARS-CoV-2 in natural water bodies.
Similar content being viewed by others
Introduction
A number of studies have reported the detection of SARS-CoV-2 in wastewater in the context of wastewater-based epidemiology for early warning of COVID-19 outbreak1,2,3,4,5,6,7. Increased concern has recently emerged, pertaining to the occurrence of SARS-CoV-2 in water environments during the current COVID-19 pandemic8,9. The awareness of the potential risk of SARS-CoV-2 from wastewater has increased since RNA detection of SARS-CoV-2 in wastewater reached public domains3,10,11,12. Recently, possible transmission of COVID-19 from sewage was reported by a cohort study in Guanzhou, China13. Although the occurrence of fecal–oral route transmission and infectivity of SARS-CoV-2 in wastewater is still controversial, there are growing concerns about the exposure risk of SARS-CoV-2 in natural water bodies that receive treated effluent from wastewater treatment plants (WWTPs), among citizens, administrative sectors, and policymakers. Because of the limited prior knowledge, the fate of SARS-CoV-2 from wastewater treatment to the water environment is still being scholarly speculated in a qualitative manner. For quantitative discussion on the potential risk of SARS-CoV-2 in water bodies, the following knowledge base is necessary: i) possible ranges of SARS-CoV-2 concentrations in wastewater, (ii) reduction of SARS-CoV-2 in wastewater treatment, (iii) decay of SARS-CoV-2 in ambient water bodies, and iv) dose–response model of SARS-CoV-2. While these key knowledges are not readily available or limited yet, revisiting the previous studies on viruses in water and wastewater brings valuable knowledge for quantitative discussion on the potential risk of SARS-CoV-2 in wastewater. There have been a number of studies on behaviors of the related coronaviruses and enveloped viruses, e.g., SARS-CoV, murine hepatitis virus (MHV), φ6, and nonenveloped enteric viruses in wastewater treatment and water bodies. Behaviors of these related viruses (e.g., disinfection efficiency, their stability, and decay in aerosol and on environmental surfaces) enable to provide more probable assumptions on the fate of SARS-CoV-2 in wastewater.
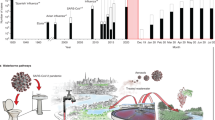
This review aims to provide an up-to-date state of key knowledge to understand the potential risk of SARS-CoV-2 in natural water bodies receiving treated wastewater in water activities. For this purpose, we present recent findings on the prevalence of SARS-CoV-2 in untreated and treated wastewater reported around the globe, revisited past studies on reduction of SARS-CoV-2-related enveloped viruses and nonenveloped enteric viruses in various wastewater treatment and disinfection processes, and dilution and decay of SARS-CoV-2 and related viruses under environmental stresses in natural water bodies. Based on this knowledge, a framework of quantitative microbial risk assessment (QMRA) for SARS-CoV-2 in natural water bodies receiving treated wastewater is suggested, along with the intensive collection of quantitative data for readily calculations of potential risks of SARS-CoV-2 based on water activities (Fig. 1). The potential risk of the various water activities was estimated based on several example scenarios of exposure by using dose–response and probability models. Finally, perspectives are addressed on limitations and applicability of the suggested QMRA framework, and possible impacts of untreated wastewater discharge like combined sewer overflows on the occurrence of SARS-CoV-2 in water bodies. This study would be beneficial to understand the potential impacts of SARS-CoV-2 in wastewater on natural water bodies and to implement QMRA-based risk management during the COVID-19 pandemic.
Concentrations of SARS-CoV-2 in wastewater
Detection of SARS-CoV-2 RNA in feces of symptomatic and asymptomatic patients of COVID-19 has directed toward the presence of SARS-CoV-2 RNA in the wastewaters10,14,15,16. Moreover, SARS-CoV-2 RNA in feces can be detected for a longer duration (22 days) followed by respiratory samples (18 days) and serum samples (16 days)17. Due to such clinical observations, it became the need of time to conduct wastewater surveillance for assessing the presence of the actual number of COVID-19 patients. To date, 11 countries have reported the presence of molecular SARS-CoV-2 RNA in wastewater samples, including untreated wastewater, treated wastewater, and sludge (Table 1). According to most of these studies, SARS-CoV-2 RNA in wastewater was detected when the number of confirmed cases reached 1–100 per million population1,2,7,11,18,19,20,21.
Table 1 summarizes the reported detection of SARS-CoV-2 RNA in treated and untreated wastewaters and sludge samples. The detection frequency was the highest in the untreated wastewater with maximum concentration ranging from 102 to 106.5 copies/L1,2,3,4,5,6,7,11,18,19,20,21,22,23. Though many studies reported no detection of SARS-CoV-2 RNA in treated wastewater5,6,7,23, some cases reported detection with maximum concentration ranging from 103 to 105 copies/L3,11,24. Interestingly, during the study conducted on Wuchang Fangcang Hospital wastewater samples, SARS-CoV-2 RNA was not detected in the untreated wastewater but reported in treated wastewater. This indicates accumulation during the treatment as well as low efficacy of current disinfection practices on hospital effluents of China24. There is a report of the detection of SARS-CoV-2 RNA in treated wastewater before chlorination, which was attributed to the low limit of detection and high filtration volume of the wastewater sample3. Previously, the sludge settled in the wastewater treatment plants has shown to contain a broad variety of human viruses including the coronaviruses25. With this knowledge, two cases, in the United States26 and Spain6 reported successful detection of SARS-CoV-2 RNA in sludge with maximum concentration ranging from 105 to 107 copies/L.
The detected concentrations of SARS-CoV-2 RNA copies in wastewater samples were consistent with possible viral shedding through excretion27. Tscharke et al.28 estimated the average quantity of wastewater produced per person, as 250 L/person/day, although it is bound to vary with geographical locations, water availability, and lifestyle. The feces quantity was modeled as a normal distribution with a mean of 2.11 log g/person and a standard deviation of 0.25 log obtained from high-income countries1,29. During the period of heaviest shedding among the mild cases of COVID-19, the shedding rate of SARS-CoV-2 RNA in feces was reported in the range 2–8 log copies/g15,17.
Reduction of SARS-CoV-2 in wastewater treatment
According to the environmental surveillance reports so far, SARS-CoV-2 RNA in treated wastewater is mostly under detection limit (~104 copies/L) (Table 1), while the typical range of the detected SARS-CoV-2 RNA in untreated wastewater has been 104–106 copies/L. Fate and removal of SARS-CoV-2 in wastewater treatment possibly differ from those of well-studied enteric viruses. SARS-CoV-2 is an enveloped virus, which is encased within an outer lipid bilayer membrane called as “envelope”, while most human enteric viruses detected in wastewater, such as norovirus, rotavirus, enterovirus, and others are nonenveloped viruses, which do not possess envelope30. This difference in surface chemical property is likely to affect virus removal efficiency in wastewater treatment, which typically ranges between 2 and 3 log10 from untreated wastewater to the final effluent of WWTP for most of the known enteric viruses (Fig. 2).
In general, LRV for conventional activated sludge (CAS) processes exhibits 1–3 log; membrane bioreactor (MBR) processes often have 1 or 2 log higher LRVs than CAS processes. Anaerobic–anoxic–oxic (A2O) processes reportedly have lower LRVs (ranging 1–2 log) than CAS (Table 2). Major removal mechanisms of viruses in wastewater are partitioning to wastewater solids in primary treatment and adsorption on sludge flocs in secondary treatment31,32. However, there is still limited knowledge of the removal of enveloped viruses in wastewater. Ye et al. investigated partitioning behaviors of two enveloped viruses, i.e., murine hepatitis virus (MHV) and φ6 in wastewater and found that enveloped viruses exhibit four times larger adsorption than the nonenveloped virus of MS2 and T333. It is known that the adsorption of virus particles is governed by extended Derjaguin–Landau–Verwey–Overbeek (XDLVO) forces, i.e., attractive Lewis van der Waals (LW), acid–base (AB) forces, and repulsive electrostatic (EL) force34,35. Since apolar or hydrophobic substances generally have larger AB force and exhibit more adhesiveness36, an enveloped virus with a relatively hydrophobic lipid bilayer membrane is also likely to be more adhesive to wastewater solids and sludge flocs than a nonenveloped enteric virus.
Disinfection is often applied to the secondary treatment effluent for inactivation of bacteria and viruses before discharge. Although a number of studies reported on disinfection of nonenveloped human enteric viruses37,38,39, only a limited number of studies reported disinfection efficiency of enveloped viruses in wastewater. Wang et al. investigated inactivation of SARS-CoV in untreated wastewater by chlorination and reported 100% reduction by a dose of free chlorine at 1 mg Cl L−1 (free residue chlorine 0.40 mg Cl/L) for 10 min40. Inactivation efficiencies of F2 phage and E. coli under the same condition were 28 and 0%. Under all tested conditions of dose and contact time, SARS-CoV was inactivated more easily than F2 phage and E. coli. Also, in treated wastewater, disinfection efficiency by chlorination could be better because it contains less amount of interfering substances that can consume free chlorine. To our knowledge, no prior studies on the efficiency of ozonation for inactivation of enveloped viruses in water are available in the public domain. Since the viral envelope is vulnerable to chemical oxidation, inactivation of SARS-CoV-2 by ozonation could be also more efficient than enteric viruses as expected in chlorination. Ultraviolet irradiation is also probably effective for the inactivation of SARS-CoV-2 in wastewater. Inagaki et al. reported SARS-CoV-2 in water could be inactivated to 3 log by UV at 250–300 nm with exposure of 3.7 mW/cm2 for 10 s (=37 mJ/cm2)41. Inactivation efficiency by UV depends on viral genome types. Generally, the single-stranded RNA (ssRNA) genome is reported to be more vulnerable than the double-stranded RNA (dsRNA) genome, and the double-stranded DNA (dsDNA) genome exhibits the highest tolerance42. Typical LRV of enteric viruses by UV disinfection process is 0.5–2 (Table 2). Walker and Ko reported that MHV coronavirus, a ssRNA virus-like SARS-CoV-2, had much lower survival than MS2, another ssRNA virus, and dsDNA adenovirus had even lower survival, under a dose of UV 254 nm43. Briefly, SARS-CoV-2 is expected to have higher (or at least equivalent) LRV than other nonenveloped enteric viruses for any disinfection processes applied in wastewater treatment.
Dilution and decay of SARS-CoV-2 in receiving water bodies
Viruses present in treated wastewater are typically released into environmental waters (i.e., rivers, lakes, and oceans) where virus concentrations are affected by a number of environmental processes categorized into two major processes of dilution and decay. While dilution is driven by seasonal dynamics of water flows, it is the primary process affecting concentration levels of any contaminant that vary significantly in time and space depending on various climatic, geological, hydrological, and anthropogenic conditions44,45,46. The decay of viruses in receiving waters involves processes like adsorption, aggregation, sedimentation, resuspension, inactivation by temperature and solar radiation, and predation. It is governed by environmental factors such as heat, light, dryness, pH, salinity, ammonia, organic matter, microbial activity, and biofilms47,48,49.
Keller et al. evaluated dilution factors of effluents from WWTPs in rivers around the world based on spatial modeling in 0.5° resolution (with an average cell size of 2250 km2)50. The authors found that annual median dilution factors differ very largely among countries, for example, Canada (33,500), Indonesia (4240), New Zealand (1070), Vietnam (508), Switzerland (260), China (145), United States (120), Bangladesh (87), Japan (58), Korea (40), United Kingdom (37), India (36), Spain (26), Oman (17), Belgium (8.7), and Morocco (5). Large variations in dilution factors also exist within a country, e.g., annual dilution factors in different locations in the United States illustrated a variation of two orders of magnitude (from 33 to 3600) difference between 25th and 75th percentiles, and four orders of magnitude (from 4.5 to 89,000) difference between 5th and 95th percentiles. Spatial variation usually becomes larger for a higher spatial resolution (i.e., more local scale)51,52,53. Temporal/seasonal variations also cause up to seven orders of magnitude variations in some tropical countries50. Similarly, lakes, ponds, and coastal waters have a diverse range of dilution factors for wastewater-derived contaminants, depending on the size of water bodies, local hydrology (e.g., inflow/outflow, vertical/horizontal transport, and exchange with the outer sea), and wastewater discharge volumes. In engineering practice, a ratio of river to wastewater flow (a dilution factor) of 40 is recommended to prevent risk54. Overall, it is pertinent to note that in stagnant and (semi) closed water systems, contaminants that are not readily attenuated (e.g., by sedimentation and degradation) can persist in the waters and pose infection risks for a long time.
The dilution of wastewater in the receiving waters can be determined by using specific chemicals or microbes as indicators/tracers. Various pharmaceuticals, personal care products, and artificial sweeteners have been suggested as chemical tracers of domestic wastewater in surface and marine waters55,56,57,58. Microbial indicators or tracers of wastewater include a diverse range of specific bacteria and viruses having various source specificity and environmental stability, which have been summarized as microbial source tracking tools59,60,61. Recent studies have employed both chemical and microbial indicators of wastewater for evaluating fecal pollution62,63. Migration of wastewater (fecal sources) to environmental waters can be sensitively detected by using these indicators/tracers, for example, raw wastewater with up to a dilution factor of 19,000 (4.3 log10) can be detected by using PMMoV and human adenovirus (AdV)63.
Among the factors affecting the decay of viruses, the temperature has been most studied and often regarded as the most influential factor on virus survival in environmental waters49,64. Other important factors include virus association with solids, exposure to UV, and the presence of microbial flora47. In general, enveloped viruses have less stability against environmental stresses than nonenveloped viruses owing to the easy disruption of proteins and lipids of the envelope65. The reported stability of various coronavirus strains varies significantly with temperature40,66,67. In water at 25 °C, the time for virus titer to decrease by 2 log10 (T99), was 22 days for transmissible gastroenteritis virus (TGEV) and 17 days for mouse hepatitis virus (MHV). On the other hand, in the water at 4 °C, both viruses did not exhibit decay over 6 weeks67. Gundy et al. showed that two coronavirus strains, human coronavirus 229E (HCoV) and feline infectious peritonitis virus (FIPV), were less stable than nonenveloped poliovirus 1 LSc-2ab (PV-1) in water and wastewater in a dark condition66. It was found that T99 in secondary effluent at 23 °C was 1.85 days for HCoV, 1.62 days for FIPV, and 3.83 days for PV-1. The difference in the stability between the two enveloped viruses and the nonenveloped virus was even larger at 23 °C (T99: 8.09 days for HCoV, 8.32 days for FIPV, and 47.5 days for PV-1). The study also reported shorter viral survival in the case of secondary effluent, encountering aggregation and adsorption to solid phases, removal by sedimentation, and envelop disruptions by solvents and detergents66.
The occurrence of enteric viruses in environmental waters has been extensively studied47,68. Interestingly, surface water and seawater can contain virus concentrations more than treated wastewater by a few orders of magnitude, particularly in the case of combined sewer overflow (CSO) events, where untreated wastewater is discharged into water bodies69,70. A recent survey in Tokyo Bay showed that both E. coli and viruses had an increase in abundance for few days after a heavy rain involving CSOs (highest concentration: 9.8 × 104 CFU/L of E. coli, 6.8 × 106 copies/L of PMMoV, 2.9 × 104 copies/L of Aichi virus, 5.6 × 103 copies/L of norovirus genogroup I, and 1.2 × 104 copies/L of norovirus genogroup II)71. However, while the increase and subsequent decrease in E. coli level were large (e.g., three orders of magnitude), virus concentrations had small fluctuations. It can probably be due to the survival of viruses derived from the previous CSOs, low removal of the viruses at WWTPs, and high inhibition in molecular detection of the viruses by environmental matrices. This highlights the stability of viruses in coastal waters, where various recreational activities can take place, as well as the importance of CSOs as a source of viruses in such waters. It is also important to use appropriate process controls and/or additional treatment of virus concentrates21,68 in analyzing viruses in environmental waters, which have various degree of inhibitory substances that reduce the efficacy of RNA extraction and RT-PCR72.
Estimated risk of exposure to SARS-CoV-2 in receiving water bodies
COVID-19 is likely to have similar transmission potential with SARS, MERS, and 1918 pandemic flu A (H1N1). The transmission potential of disease is represented by the basic reproduction number (R0), which is the number of secondary infections resulting from a single primary infection into an otherwise susceptible population. The basic reproduction number (R0) of COVID-19 reportedly ranged 2.2–3.673,74,75, which is in the range of the reported R0 values for SARS, MERS, and 1918 pandemic flu A (H1N1)74,76. Since the basic reproduction number depends on not only the infectivity of the virus but also its infectious period, the similarity of the transmission potential does not directly mean the similarity of viral infectivity indicated as a dose–response relationship. Although sufficient data are not available for modeling the dose–response relations of SARS-CoV-2 yet, reported viral doses in some animal experiments to indicate that SARS-CoV and SARS-CoV-2 are more infectious than influenza virus77.
Infection of SARS-CoV-2 by ingestion is still controversial but potentially occurs9. SARS-CoV-2 is the phylogenetically related species of SARS-CoV, which caused the SARS outbreaks in 2003. SARS-CoV and SARS-CoV-2 have the same infection mechanisms, in which the virus enters human cells by binding their spike (S) proteins to ACE2 receptors of human cells8. Since the ACE2 receptor is abundant in small intestine cells78, SARS-CoV-2 infection possibly occurs in the gastrointestinal tract when the virus survives in gastric acid and reaches small intestines79. Survival of SARS-CoV-2 was observed at a high titer under acidic conditions (pH 2.2), which mimics the gastric environment80. Moreover, gastric pH could be around 4.0–5.0 by medication of acid-reducing drugs and gastrointestinal feeding81. In such cases, SARS-CoV-2 is more likely to survive in gastric acid and cause infection, as reported with SARS-CoV by Darnell et al.82.
In QMRA, the chances of infection by a virus are calculated by a dose–response model, which describes relations of viral exposure dose and the probability of infection. Currently, the dose–response model of SARS-CoV-2 is not available yet; however, a dose–response model of SARS-CoV through intranasal infection has been developed by Watanabe et al.83. This model could be applied to complement the lack of a dose–response model of SARS-CoV-2 with careful consideration of its uncertainty84. Watanabe’s model suggested an exponential model with the following equation:
where p(r|d): the chance of infection at the viral dose of d, d: dose of virus (PFU, plaque-forming unit), k: 4.2 × 102 (PFU). According to this model, 1 PFU of viral dose causes 0.24% of infection and 2 PFU causes 0.49%.
The expected dose of the virus is estimated from the volume of water ingestion and viral concentration in the water. SARS-CoV-2 concentration in treated wastewater expectedly ranges between 1 and 100 copies/L, assuming the presence of 104–105 copies/L of SARS-CoV-2 in untreated wastewater and 3 log of removal (equivalent to that of nonenveloped enteric viruses) in wastewater treatment. In water bodies receiving the treated wastewater, SARS-CoV-2 concentrations are expected to be even lower by dilution and decay. As discussed above, factors of dilution and decay are variable to be five times to 4–5 log, depending on the locations of discharge. Hence, SARS-CoV-2 RNA in receiving water bodies does not probably exceed <100 copies/L, except in the worst case like in an urban river that consists of a large proportion of treated wastewater from highly infected areas. Moreover, the viable count of the virus as PFU is probably smaller than viral RNA concentration since viral RNA concentration possibly includes viral RNA from a virus that has lost its infectivity.
The expected volume of water ingestion depends on the activity to expose the water body. For example, the median volume of water ingestion per event is reported as 2.0 mL in fishing, 6.0 mL in swimming85. When the water body contained 1 PFU/L of viral concentration, the expected viral dose will be 0.002 PFU/event in fishing and 0.006 PFU/event in swimming. When the expected viral dose is smaller than 1 PFU, the probability where virus particles are ingested should also be considered. Assuming that viruses are randomly distributed in the water, the viral dose from a certain volume of water follows the Poisson distribution:
where p(r|d): the probability of the event where the viral dose of d occurs under the expected dose of λ, λ: the expected dose per event.
Therefore, the chance of infection under the expected dose of λ, P(r|λ), can be calculated as
The potential chance of infection of SARS-CoV from a given expected viral dose in water activities is derived as in Fig. 3 by assuming all the ingestion dose reaches the nasal cavity. For example, when the water body contained 1 PFU/L of virus concentration, the expected viral dose will be 0.002 PFU/event in fishing and 0.006 PFU/event in swimming. The estimated chance of infection per event is derived from equation (3) as 4.9 × 10−6 in fishing and 1.5 × 10−5 in swimming (Table 3).
SARS-CoV-2 is phylogenetically related to SARS-CoV with 79% (in ~30,000 bp) of genomic similarity to SARS-CoV86. Both species have the spike (S) protein to enter the host cells by interacting with angiotensin-converting enzyme 2 (ACE2). The structure of the receptor-binding domain of SARS-CoV-2 has been evolved from SARS-CoV, and possibly causes higher infectivity86,87. Viral doses in some animal model experiments suggest that SARS-CoV-2 is potentially more infectious than SARS-CoV77. Hence, the estimated chance of infection for COVID-19 in this study could be underestimated and has large uncertainty.
Perspectives
We presented a QMRA framework to estimate the risk of SARS-CoV-2 in treated wastewater and receiving water bodies by employing the best of the currently available related knowledge. However, the estimated risk values obtained potentially have large uncertainty due to various assumptions and limitations. The major limitations that possibly cause uncertainties in the estimation are (i) employment of the dose–response model for SARS-CoV, and (ii) unknown ratio of SARS-CoV-2 in wastewater as RNA copies to infectious viable count as PFU. The dose–response model for SARS-CoV used in this study needs to be replaced with yet-to-be-developed actual dose–response model of SARS-CoV-2. The dose–response model of SARS-CoV was developed from data sets of challenges in mice with intranasal inoculation of murine hepatitis virus 1 (MHV-1) as a surrogate for SARS-CoV. There must be large uncertainties regarding differences of SARS-CoV-2 from MHV-1, gastrointestinal from intranasal infection including a possible reduction in gastric acid, and human from mice. Although SARS-CoV-2 and SARS-CoV have a similar infectious mechanism, there could be a significant difference in gastrointestinal dose–response relations of SARS-CoV-2 from the dose–response model for SARS-CoV employed in this study. Moreover, viral RNA copies are assumed to be the maximum viable count as PFU because virus quantity as RNA copies possibly includes the remaining RNA from viruses, which has decayed or lost their infectivity. There are limitations pertaining to the use of virus quantity by molecular methods, e.g., RNA copies, for QMRA as addressed by Haas et al.88. The actual infectivity of SARS-CoV-2 in wastewater quantified as RNA copies is still uncertain. While the persistence of SARS-CoV-2 is being studied under different environmental circumstances, Remoldi et al. reported insignificant infectivity of SARS-CoV-2 in untreated and treated wastewater from the observation of no cytopathic effects on Vero E6 cells following the inoculation of 48 h and 72 h89. It seems that the most SARS-CoV-2 in wastewater possibly already lost its infectivity or inherently inactive owing to its vulnerability to detergents and alcohols, as well as exposure to detergents in sewer lines for hours90,91,92. Hence, the estimated risk in this study could be overestimated from this assumption, because viable count as PFU can be smaller than virus quantity as RNA copies. However, even with such large uncertainty, decision-makers are required to put the best efforts to manage the possible risk by SARS-CoV-2 in treated wastewater, because WWTPs need to continue their operation even under COVID-19 outbreaks. The use of QMRA could be helpful for the management of the potential risk of SARS-CoV-2 in water bodies rather than without any quantitative idea on possible risk.
Although this study focused on the impacts of SARS-CoV-2 in treated wastewater on water bodies, many natural water bodies are often affected by untreated wastewater, as the decay of SARS-CoV-2 is still a debated issue93. Discharge of untreated wastewater from combined sewer overflows (CSOs) is highly significant and very common in many countries, including Central Europe (around 70% systems are combined sewer systems) and in the United States (serving more than 40 million individuals)94,95. The quantity of untreated CSOs reached 20% of the total annual discharge of combined sewage in the United States95. CSOs possess a higher concentration of bacteria, viruses, and other pathogens than the WWTP effluents, as it contains untreated sewage and hence causes pathogenic contamination in the ambient waters62,96,97,98. Loads of fecal indicator bacteria from CSOs into natural water bodies can be 4-log larger than those from treated effluent when 23% of combined sewage was discharged as untreated CSOs96. The detection of SARS-CoV-2 RNA is already reported in rivers in Italy and Ecuador, indicating contaminated streams of untreated wastewater12,89. It should be noted that the occurrence of CSO events during COVID-19 outbreaks possibly causes a substantial increase in the infection risk of SARS-CoV-2 by exposure to receiving water bodies. Thus, more careful risk assessment and management are necessary for water bodies that possibly receive untreated wastewater including CSOs.
Overall, the motivation of this paper was to provide information on potential risk in treated wastewater from the best of the current knowledge for such decision-making. It should be carefully noted that the estimated risk of SARS-CoV-2 presented in our study should be considered as the bottom line of calculating possible risk under the QMRA framework for SARS-CoV-2 in treated wastewater and water environment. It must further be updated with the development of dose–response model of SARS-CoV-2, the proportion of viable SARS-CoV-2 to RNA copies in wastewater and water environment, for better confidence and reliability.
References
Medema, G., Heijnen, L., Elsinga, G., Italiaander, R. & Brouwer, A. Presence of SARS-Coronavirus-2 RNA in sewage and correlation with reported COVID-19 prevalence in the early stage of the epidemic in the Netherlands. Environ. Sci. Technol. 7, 511–516 (2020).
Ahmed, W. et al. First confirmed detection of SARS-CoV-2 in untreated wastewater in Australia: A proof of concept for the wastewater surveillance of COVID-19 in the community. Sci. Total Environ. 728, 138764 (2020).
Haramoto, E., Malla, B., Thakali, O. & Kitajima, M. First environmental surveillance for the presence of SARS-CoV-2 RNA in wastewater and river water in Japan. Sci. Total Environ. 737, 140405 (2020).
La Rosa, G. et al. First detection of SARS-CoV-2 in untreated wastewaters in Italy. Sci. Total Environ. 736, 139652 (2020).
Sherchan, S. P. et al. First detection of SARS-CoV-2 RNA in wastewater in North America: A study in Louisiana, USA. Sci. Total Environ. 743, 140621 (2020).
Randazzo, W. et al. SARS-CoV-2 RNA in wastewater anticipated COVID-19 occurrence in a low prevalence area. Water Res. 181, 115942 (2020).
Kumar, M. et al. First proof of the capability of wastewater surveillance for COVID-19 in India through detection of genetic material of SARS-CoV-2. Sci. Total Environ. 746, 141326 (2020).
Ding, S. & Liang, T. J. Is SARS-CoV-2 also an enteric pathogen with potential fecal oral transmission: A COVID-19 virological and clinical review. Gastroenterology 159, 53–61 (2020).
Amirian, E. S. Potential fecal transmission of SARS-CoV-2: Current evidence and implications for public health. Int. J. Infect. Dis. 95, 363–370 (2020).
Zhang, D. et al. Potential spreading risks and disinfection challenges of medical wastewater by the presence of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) viral RNA in septic tanks of Fangcang Hospital. Sci. Total Environ. 741, 140445 (2020).
Wurtzer, S., Marechal, V., Mouchel, J. M. & Moulin, L. Time course quantitative detection of SARS-CoV-2 in Parisian wastewaters correlates with COVID-19 confirmed cases. Preprint at https://www.medrxiv.org/content/10.1101/2020.04.12.20062679v1.full.pdf (2020).
Guerrero-latorre, L. et al. SARS-CoV-2 in river water: Implications in low sanitation countries. Sci. Total Environ. 743, 140832 (2020).
Yuan, J. et al. Sewage as a possible transmission vehicle during a coronavirus disease 2019 outbreak in a densely populated community: Guangzhou, China. Clin. Infect. Dis. 1494, 1494 (2020).
Cheung, K. S. et al. Gastrointestinal manifestations of SARS-CoV-2 infection and virus load in fecal samples from the Hong Kong Cohort and systematic review and meta-analysis. Gastroenterology 159, 81–95 (2020).
Wölfel, R. et al. Virological assessment of hospitalized cases of coronavirus disease 2019. Nature 581, 465–469 (2020).
Lavezzo, E. et al. Suppression of a SARS-CoV-2 outbreak in the Italian municipality of Vo’. Nature 584, 425–429 (2020).
Zheng, S. et al. Viral load dynamics and disease severity in patients infected with SARS-CoV-2 in Zhejiang province, China, January–March 2020: retrospective cohort study. BMJ. 369, m1443 (2020).
Bar-Or, I. et al. Regressing SARS-CoV-2 sewage measurements onto COVID-19 burden in the population: a proof-of-concept for quantitative environmental surveillance. Preprint at https://www.medrxiv.org/content/10.1101/2020.04.26.20073569v1 (2020).
Nemudryi, A. et al. Temporal detection and phylogenetic assessment of SARS-CoV-2 in municipal wastewater. Cell Rep. Med. 1, 100098 (2020).
Wu, F. et al. SARSCoV-2 titers in wastewater are higher than expected from clinically confirmed cases. Preprint at https://www.medrxiv.org/content/10.1101/2020.04.05.20051540v1.full.pdf (2020).
Hata, A., Hara-Yamamura, H., Meuchi, Y., Imai, S. & Honda, R. Detection of SARS-CoV-2 in wastewater in Japan during a COVID-19 outbreak. Sci. Total Environ. 758, 143578 (2020).
Kocamemi, B. A. et al. First data-set on SARS-CoV-2 detection for Istanbul wastewaters in Turkey. Preprint at https://www.medrxiv.org/content/10.1101/2020.05.03.20089417v1 (2020).
Balboa, S. et al. The fate of SARS-CoV-2 in wastewater treatment plants points out the sludge line as a suitable spot for incidence monitoring. Preprint at https://www.medrxiv.org/content/10.1101/2020.05.25.20112706v1 (2020).
Zhang, W. et al. Molecular and serological investigation of 2019-nCoV infected patients: implication of multiple shedding routes. Emerg. Microbes Infect. 9, 386–389 (2020).
Bibby, K. & Peccia, J. Identification of viral pathogen diversity in sewage sludge by metagenome analysis. Environ. Sci. Technol. 47, 1945–1951 (2013).
Peccia, J. et al. Measurement of SARS-CoV-2 RNA in wastewater tracks community infection dynamics. Nat. Biotechnol. 38, 1164–1167 (2020).
Hata, A. & Honda, R. Potential sensitivity of wastewater monitoring for SARS-CoV-2: comparison with norovirus cases. Environ. Sci. Technol. 54, 6451–6452 (2020).
Tscharke, B. J. et al. Harnessing the power of the census: characterizing wastewatertreatment plant catchment populations for wastewater-based epidemiology. Environ. Sci. Technol. 53, 10303–10311 (2019).
Rose, C., Parker, A., Jefferson, B. & Cartmell, E. The characterization of feces and urine:a review of the literature to inform advanced treatment technology. Crit. Rev. Environ. Sci. Technol. 45, 1827–1879 (2015).
Qiu, Y. et al. Assessment of human virus removal during municipal wastewater treatment in Edmonton, Canada. J. Appl. Microbiol. 119, 1729–1739 (2015).
Gerba, C. P. Virus survival in wastewater treatment. In: Viruses and Wastewater Treatment (Goddard M, Butler M. eds) Pergamon Press, New York, pp. 39–48 (1981).
Hurst, C. J. & Gerba, C. P. Fate of viruses during wastewater sludge treatment processes. Crit. Rev. Env. Sci. Technol 18, 317–343 (2009).
Ye, Y., Ellenberg, R. M., Graham, K. E. & Wigginton, K. R. Survivability, partitioning, and recovery of enveloped viruses in untreated municipal wastewater. Environ. Sci. Technol. 50, 5077–5085 (2016).
Hermansson, M. The DLVO theory in microbial adhesion. Colloids and Surf. B: Biointerfaces 14, 105–119 (1999).
Chrysikopoulos, C. V. & Syngouna, V. I. Attachment of bacteriophages MS2 and ΦX174 onto kaolinite and montmorillonite: extended-DLVO interactions. Colloids and Surf. B: Biointerfaces 92, 74–83 (2012).
van Oss, C. J. Interfacial Forces in Aqueous Media, second edition (CRC Press, 2006).
Aieta, E. M. & Berg, J. D. A review of chlorine dioxide in drinking water treatment. J. AWWA 78, 62–72 (1986). 1986.
Kim, J. G., Yousef, A. E. & Dave, S. Application of ozone for enhancing the microbiological safety and quality of foods: a review. J. Food Prot. 62, 1071–1087 (1999).
Hijnen, W. A. M., Beerendonk, E. F. & Medema, G. J. Inactivation credit of UV radiation for viruses, bacteria and protozoan (oo)cysts in water: a review. Water Res. 40, 3–22 (2006). 2006.
Wang, X. et al. Excretion and detection of SARS coronavirus and its nucleic acid from digestive system. World J. Gastroenterol. 11, 4390–4395 (2005).
Inagaki, H., Saito, A., Sugiyama, H., Okabayashi, T. & Fujimoto, S. Rapid inactivation of SARS-CoV-2 with deep-UV LED irradiation. Emerg. Microbes Infect. 9, 1–8 (2020).
Pepper, I. L., Gerba, C. P., Gentry, T. J. & Maier, R. M. Environmental Microbiology, 2nd edn. Elsevier, Academic Press, London (2011).
Walker, C. M. & Ko, G. Effect of ultraviolet germicidal irradiation on viral aerosols. Environ. Sci. Technol. 41, 5460–5465 (2007).
Keller, V. Risk assessment of “down-the-drain” chemicals: search for a suitable model. Sci. Total Environ. 360, 305–318 (2006).
Johnson, A. C. Natural variations in flow are critical in determining concentrations of point source contaminants in rivers: an estrogen example. Environ Sci Technol. 44, 7865–7870 (2010).
Osorio, V. et al. Occurrence and modeling of pharmaceuticals on a sewage-impacted Mediterranean river and their dynamics under different hydrological conditions. Sci. Total Environ. 440, 3–13 (2012).
Bosch, A., Pintó, R. M. & Abad, F. X. Survival and transport of enteric viruses in the environment. In: Goyal S. M. (ed.) Viruses in foods Springer, Boston, MA, pp 151–187 (2006).
Brookes, J. D. et al. Fate and transport of pathogens in lakes and reservoirs. Environ. Int. 30, 741–759 (2004).
Pinon, A. & Vialette, M. Survival of viruses in water. Intervirology 61, 214–222 (2018).
Keller, V. D., Williams, R. J., Lofthouse, C. & Johnson, A. C. Worldwide estimation of river concentrations of any chemical originating from sewage‐treatment plants using dilution factors. Environ. Toxicol. Chem. 33, 447–452 (2014).
Kehrein, N., Berlekamp, J. & Klasmeier, J. Modeling the fate of down-the-drain chemicals in whole watersheds: new version of the GREAT-ER software. Environ Model Softw. 64, 1–8 (2015).
Kuroda, K. et al. Hospital-use pharmaceuticals in Swiss waters modeled at high spatial resolution. Environ. Sci. Technol. 50, 4742–4751 (2016).
Siddiqui, S., Conkle, J. L., Scarpa, J. & Sadovski, A. An analysis of US wastewater treatment plant effluent dilution ratio: implications for water quality and aquaculture. Sci. Total Environ. 721, 137819 (2020).
Gupta, R. S. Solutions Manual to Accompany Hydrology and Hydraulic Systems (Waveland Press, 2008).
Nakada, N. et al. Evaluation of pharmaceuticals and personal care products as water-soluble molecular markers of sewage. Environ. Sci. Technol. 42, 6347–6353 (2008).
Benotti, M. J. & Brownawell, B. J. Distributions of pharmaceuticals in an urban estuary during both dry-and wet-weather conditions. Environ. Sci. Technol. 41, 5795–5802 (2007).
Buerge, I. J., Buser, H. R., Kahle, M., Muller, M. D. & Poiger, T. Ubiquitous occurrence of the artificial sweetener acesulfame in the aquatic environment: an ideal chemical marker of domestic wastewater in groundwater. Environ. Sci. Technol. 43, 4381–4385 (2009).
Cantwell, M. G. et al. Spatial patterns of pharmaceuticals and wastewater tracers in the Hudson River Estuary. Water Res. 137, 335–343 (2018).
Harwood, V. J., Staley, C., Badgley, B. D., Borges, K. & Korajkic, A. Microbial source tracking markers for detection of fecal contamination in environmental waters: relationships between pathogens and human health outcomes. FEMS. Microbiol. Rev. 38, 1–40 (2014).
Stoeckel, D. M. & Harwood, V. J. Performance, design, and analysis in microbial source tracking studies. Appl. Environ. Microbiol. 73, 2405–2415 (2007).
Tran, N. H., Gin, K. Y. H. & Ngo, H. H. Fecal pollution source tracking toolbox for identification, evaluation and characterization of fecal contamination in receiving urban surface waters and groundwater. Sci. Total Environ. 538, 38–57 (2015).
Kumar, M. et al. Concurrence of antibiotic resistant bacteria (ARB), viruses, pharmaceuticals and personal care products (PPCPs) in ambient waters of Guwahati, India: urban vulnerability and resilience perspective. Sci. Total Environ. 693, 133640 (2019).
Kuroda, K. et al. Pepper mild mottle virus as an indicator and a tracer of fecal pollution in water environments: comparative evaluation with wastewater-tracer pharmaceuticals in Hanoi, Vietnam. Sci. Total Environ. 506, 287–298 (2015).
Bertrand, I. et al. The impact of temperature on the inactivation of enteric viruses in food and water: a review. J. Appl. Microbiol. 112, 1059–1074 (2012).
Fitzgibbon, J. E. & Sagripanti, J. Analysis of the survival of Venezuelan equine encephalomyelitis virus and possible viral simulants in liquid suspensions. J. Appl. Microbiol. 105, 1477–1483 (2008).
Gundy, P. M., Gerba, C. P. & Pepper, I. L. Survival of coronaviruses in water and wastewater. Food Environ. Virol. 1, 10 (2009).
Casanova, L., Rutala, W. A., Weber, D. J. & Sobsey, M. D. Survival of surrogate coronaviruses in water. Water Res. 43, 1893–1898 (2009).
Haramoto, E. et al. A review on recent progress in the detection methods and prevalence of human enteric viruses in water. Water Res. 135, 168–186 (2018).
Phillips, P. J. et al. Combined sewer overflows: an environmental source of hormones and wastewater micropollutants. Environ. Sci. Technol. 46, 5336–5343 (2012).
Passerat, J., Ouattara, N. K., Mouchel, J. M., Rocher, V. & Servais, P. Impact of an intense combined sewer overflow event on the microbiological water quality of the Seine River. Water Res. 45, 893–903 (2011).
Inoue, K., Asami, T., Shibata, T., Furumai, H. & Katayama, H. Spatial and temporal profiles of enteric viruses in the coastal waters of Tokyo Bay during and after a series of rainfall events. Sci. Total Environ. 727, 138502 (2020).
Hata, A., Kitajima, M. & Katayama, H. Occurrence and reduction of human viruses, F-specific RNA coliphage genogroups and microbial indicators at a full-scale wastewater treatment plant in Japan. J. Appl. Microbiol. 114, 545–554 (2013).
D'Arienzo, M. & Coniglio, A. Assessment of the SARS-CoV-2 basic reproduction number, R0, based on the early phase of COVID-19 outbreak in Italy. Biosaf. Heal. 2, 57–59 (2020).
Zhao, S. et al. Preliminary estimation of the basic reproduction number of novel coronavirus (2019-nCoV) in China, from 2019 to 2020: a data-driven analysis in the early phase of the outbreak. Int. J. Infect. Dis. 92, 214–217 (2020).
Lai, A., Bergna, A., Acciarri, C., Galli, M. & Zehender, G. Early phylogenetic estimate of the effective reproduction number of SARS-CoV-2. J. Med. Virol. 92, 675–679 (2018).
Callaway, E. Coronavirus vaccines: five key questions as trials begin. Nature 579, 481 (2020).
Kitajima, M. et al. SARS-CoV-2 in wastewater: state of the knowledge and research needs. Sci. Total Environ. 739, 139076 (2020).
Hamming, I. et al. Tissue distribution of ACE2 protein, the functional receptor for SARS coronavirus. A first step in understanding SARS pathogenesis. J. Pathol. 203, 631–637 (2004).
Xiao, F. et al. Evidence for gastrointestinal infection of SARS-CoV-2. Gastroenterology 158, 1831–1833.e3 (2020).
Sun, Z. et al. Survival of SARS-COV-2 under liquid medium, dry filter paper and acidic conditions. Cell Discov. 6, 57 (2020).
Price, E. Could the severity of COVID-19 be increased by low gastric acidity? Crit. Care 24, 456 (2020).
Darnell, M. E. R., Subbarao, K., Feinstone, S. M. & Taylor, D. R. Inactivation of the coronavirus that induces severe acute respiratory syndrome, SARS-CoV. J. Virol. Methods 121, 85–91 (2004).
Watanabe, T., Bartrand, T. A., Weir, M. H., Omura, T. & Haas, C. N. Development of a dose-response model for SARS coronavirus. Risk Anal. 30, 1129–1138 (2010).
Haas, C. Coronavirus and risk analysis. Risk Anal. 40, 660–661 (2020).
Dorevitch, S. et al. Water ingestion during water recreation. Water Res. 45, 2020–2028 (2011).
Lu, R. et al. Genomic characterization and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet 395, 565–574 (2020).
Shang, J. et al. Structural basis of receptor recognition by SARS-CoV-2. Nature 581, 221–224 (2020).
Haas, C. N. Quantitative microbial risk assessment and molecular biology: paths to integration. Environ. Sci. Technol. 54, 8539–8546 (2020).
Rimoldi, S. G. et al. Presence and infectivity of SARS-CoV-2 virus in wastewaters and rivers. Sci. Total Environ. 744, 140911 (2020).
Camacho-Muñoz, D., Martín, J., Santos, J. L., Aparicio, I. & Alonso, E. Occurrence of surfactants in wastewater: Hourly and seasonal variations in urban and industrial wastewaters from Seville (Southern Spain). Sci. Total Environ. 468–469, 977–984 (2014).
Mungray, A. K. & Kumar, P. Fate of linear alkylbenzene sulfonates in the environment: a review. Int. Biodeterior. Biodegrad. 63, 981–987 (2009).
Teerlink, J., Hering, A. S., Higgins, C. P. & Drewes, J. E. Variability of trace organic chemical concentrations in raw wastewater at three distinct sewershed scales. Water Res. 46, 3261–3271 (2012).
Kumar, M. et al. Decay of SARS-CoV-2 RNA along the wastewater treatment outfitted with upflow anaerobic sludge blanket (UASB) system evaluated through two sample concentration techniques. Sci. Total Environ. 754, 142329 (2021).
Butler, D. & Davies, J. W. Combined sewers and combined sewer overflows. In: Butler, D. & Davies, J. W. (eds) Urban Drainage. Spon Press, Taylor and Francis Group, New York, USA, pp. 254−289 (2004).
U.S. Environmental Protection Agency (EPA). Chapter 5: Environmental Impacts of CSOs and SSOs. Report to Congress on Impacts and Control of Combined Sewer Overflows and Sanitary Sewer Overflows, Water. Office of Water, Washington, DC, USA, pp. 5-1~5-27 (2004).
Honda, R. et al. Estimated discharge of antibiotic-resistant bacteria from combined sewer overflows of urban sewage system. npj Clean Water 3, 15 (2020).
Honda, R. et al. Impacts of urbanization on the prevalence of antibiotic-resistant Escherichia coli in the Chaophraya river and its tributaries. Water Sci. Technol. 73, 362–374 (2016).
Kumar, M., Chaminda, G. T. & Honda, R. Seasonality impels the antibiotic resistance in Kelani river of the emerging economy of Sri Lanka. npj Clean Water 3, 1–8 (2020).
Sano, D., Amarasiri, M., Hata, A., Watanabe, T. & Katayama, H. Risk management of viral infectious diseases in wastewater reclamation and reuse: review. Environ. Int. 91, 220–229 (2016).
Kitajima, M., Iker, B. C., Pepper, I. L. & Gerba, C. P. Relative abundance and treatment reduction of viruses during wastewater treatment processes—identification of potential viral indicators. Sci. Total Environ. 488–489, 290–296 (2014).
Nordgren, J., Bucardo, F., Dienus, O., Svensson, L. & Lindgren, P. E. Novel light-upon-extension real-time PCR assays for detection and quantification of genogroup I and II noroviruses in clinical specimens. J. Clin. Microbiol. 46, 164–170 (2008).
Miura, T., Schaeffer, J., Le Saux, J.-C., Le Mehaute, P. & Le Guyader, F. S. Virus type-specific removal in a full-scale membrane bioreactor treatment process. Food Environ. Virol. 10, 176–186 (2018).
Da Silva, A. K. et al. Evaluation of removal of noroviruses during wastewater treatment, using real-time reverse transcription-PCR: different behaviors of genogroups I and II. Appl. Environ. Microbiol. 73, 7891–7897 (2007).
Chaudhry, R. M., Nelson, K. L. & Drewes, J. E. Mechanisms of pathogenic virus removal in a full-scale membrane bioreactor. Environ. Sci. Technol. 49, 2815–2822 (2015).
Simmons, F. J., Kuo, D. H. W. & Xagoraraki, I. Removal of human enteric viruses by a full-scale membrane bioreactor during municipal wastewater processing. Water Res. 45, 2739–2750 (2011).
Miura, T., Okabe, S., Nakahara, Y. & Sano, D. Removal properties of human enteric viruses in a pilot-scale membrane bioreactor (MBR) process. Water Res. 75, 282–291 (2015).
Katayama, H. et al. One-year monthly quantitative survey of noroviruses, enteroviruses, and adenoviruses in wastewater collected from six plants in Japan. Water Res. 42, 1441–1448 (2008).
Hewitt, J., Leonard, M., Greening, G. E. & Lewis, G. D. Influence of wastewater treatment process and the population size on human virus profiles in wastewater. Water Res. 45, 6267–6276 (2011).
Rosario, K., Symonds, E. M., Sinigalliano, C., Stewart, J. & Breitbart, M. Pepper mild mottle virus as an indicator of fecal pollution. Appl. Environ. Microbiol. 75, 7261–7267 (2009).
Papp, K., Moser, D. & Gerrity, D. Viral surrogates in potable reuse applications: evaluation of a membrane bioreactor and full advanced treatment. J. Environ. Eng. 146, 04019103 (2020).
Kuo, D. et al. Assessment of human adenovirus removal in a full-scale membrane bioreactor treating municipal wastewater. Water Res. 44, 1520–1530 (2010).
Jumat, M. R. et al. “Membrane bioreactor-based wastewater treatment plant in Saudi Arabia: reduction of viral diversity, load, and infectious capacity.”. Water 9, 534 (2017).
Ottoson, J., Hansen, A., Bjorlenius, B., Norder, H. & Stenstrom, T. A. Removal of viruses, parasitic protozoa and microbial indicators in conventional and membrane processes in a wastewater pilot plant. Water Res. 40, 1449–1457 (2006).
Li, D. et al. Monitoring and evaluation of infectious rotaviruses in various wastewater effluents and receiving waters revealed correlation and seasonal pattern of occurrences. J. Appl. Microbiol. 110, 1129–1137 (2011).
Tandukar, S., Sherchan, S. P. & Haramoto, E. Reduction of human enteric and indicator viruses at a wastewater treatment plant in Southern Louisiana, USA. Food Environ. Virol. 0123456789, 10–13 (2020).
Gomes, J., Frasson, D., Quinta-Ferreira, R. M., Matos, A. & Martins, R. C. Removal of enteric pathogens from real wastewater using single and catalytic ozonation. Water 11, 1–12 (2019).
Wang, H. et al. Differential removal of human pathogenic viruses from sewage by conventional and ozone treatments. Int. J. Hyg. Environ. Health 221, 479–488 (2018).
Acknowledgements
This study was supported by JST MIRAI Program (Grant No. JPMJMI18DC) and Hiramoto-Gumi Inc., a civil engineering company in Japan.
Author information
Authors and Affiliations
Contributions
M.K.: writing, reviewing, and editing all parts of the paper; M.A.: literature review and writing in the section “Reduction of SARS-CoV-2 in wastewater treatment”; K.K.: literature review and writing in the section “Dilution and decay of SARS-CoV-2 in receiving water bodies”; K.D.: literature review and writing in the section “Concentrations of SARS-CoV-2 in wastewater”; A.H.: review and editing all parts of the paper; H.Y.: formal analysis, reviewing, and editing in the section “Estimated risk of exposure to SARS-CoV-2 in receiving water bodies”; R.H.: conceptualization, formal analysis, writing the sections “Estimated risk of exposure to SARS-CoV-2 in receiving water bodies” and “Perspectives”, reviewing, and editing all parts of the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kumar, M., Alamin, M., Kuroda, K. et al. Potential discharge, attenuation and exposure risk of SARS-CoV-2 in natural water bodies receiving treated wastewater. npj Clean Water 4, 8 (2021). https://doi.org/10.1038/s41545-021-00098-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41545-021-00098-2
This article is cited by
-
Enhanced detection of viruses for improved water safety
Scientific Reports (2023)
-
Environmental monitoring of SARS-CoV-2 in the metropolitan area of Porto Alegre, Rio Grande do Sul (RS), Brazil
Environmental Science and Pollution Research (2023)
-
Detection and identification of potentially infectious gastrointestinal and respiratory viruses at workplaces of wastewater treatment plants with viability qPCR/RT-qPCR
Scientific Reports (2022)
-
Reply: Potential discharge, attenuation and exposure risk of SARS-CoV-2 in natural water bodies receiving treated wastewater
npj Clean Water (2021)
-
The impact of coronavirus SARS-CoV-2 (COVID-19) in water: potential risks
Environmental Science and Pollution Research (2021)