Abstract
Antibiotics are widely used in animal husbandry, and various types of antibiotic resistance genes (ARGs) are frequently detected in livestock waste around the world. Conventional livestock waste treatment processes do not completely remove ARGs, resulting in their release to soil and water environments. Various exposure routes of these ARGs to humans, including inhalation and ingestion of antibiotic-resistant bacteria (ARB) that harbor them, may be contributing to the rise in resistant clinical infections that are increasingly difficult to treat with antibiotics. In this review, we assess the occurrence and variability of ARGs in livestock wastes and their potential propagation pathways to human pathogens. We also review the mechanisms and environmental factors that influence the dissemination of ARGs through these pathways, and evaluate the ARG removal efficiency of common livestock waste management approaches. Challenges and research needs for assessing and mitigating the risk of antibiotic resistance dissemination from livestock waste are also presented.
Similar content being viewed by others
Introduction
Annual global deaths from antibiotic-resistant infections are projected to increase from 700,000 in 2014 to 10 million by 2050, with cumulative costs in healthcare and reduced productivity reaching $100 trillion USD.1 Even countries that have made substantial efforts to reduce antibiotic use2,3 still observe increasing rates of clinical antibiotic resistance,4 highlighting the complex global nature of this problem.
Globally, animal husbandry accounts for over one-half of all antibiotic use (Fig. 1), which was estimated at 131,109 tons in 2013 and is projected to reach over 200,000 tons by 2030.5 However, only 10% of publications on antibiotic resistance consider the potential contribution from animal husbandry (Fig. 1).
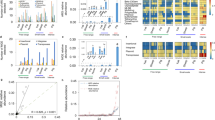
Web of Science results (from 2000 to 2018) show an exponential increase in the number of annual publications related to resistance propagation in the environment. However, the number of publications related to antibiotic resistance in animal husbandry is not commensurate with the dominant use of antibiotics in this sector. Inset shows antibiotic consumption by humans and animals in three major countries/regions.76,154.
Most of the antibiotics used in animal husbandry are for non-therapeutic purposes, such as growth promotion and disease prevention, and are consistently detected in livestock gastrointestinal environments at low and sub-lethal concentrations that slow down the growth of susceptible bacterial populations.6,7 This exerts selective pressure for bacteria in livestock digestive systems to acquire and maintain antibiotic resistance genes (ARGs) and fosters an increase in the relative abundance of resistant populations.8 When these ARGs propagate to surrounding environments, antibiotic resistance becomes an environmental pollution problem, with ARGs as contaminants of emerging concern.9 For example, when antibiotic-resistant bacteria (ARB) in livestock gastrointestinal environments are excreted,10 ARGs are disseminated into receiving environments (e.g., soil, water). Subsequent ARG replication and propagation would increase the likelihood of human exposure, particularly for agricultural workers and those living in neighboring areas.
A clear etiology between ARG propagation from animal husbandry and ARG acquisition by human gut microbiomes has yet to be established,11 which underscores the need for an updated and more holistic perspective of how ARGs released from animal husbandry affect environmental and human resistomes. Furthermore, selective pressure for ARGs in animal husbandry settings is a major operational concern as it may limit the number of antibiotics that are effective for therapeutic treatments, which is conducive to higher incidence of disease and rising production costs.5
In this review, we examine the research progress on the occurrence and release of ARGs from animal husbandry, their propagation into receiving environments, and their potential exposure pathways to human pathogens. Furthermore, we evaluate the removal efficacy of ARGs by different livestock waste treatment processes. Finally, we highlight critical knowledge gaps and research needs, and provide suggestions to mitigate the risk of ARGs from livestock waste.
Occurrence and variability of ARGs in livestock waste
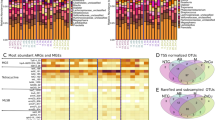
Through three fundamental resistance mechanisms (antibiotic deactivation, extrusion through efflux pumps, and protection of targets—such as ribosomes—by specific proteins12), ARGs can confer resistance to nine major classes of antibiotics: tetracyclines (tet), sulfonamides (sul), β-lactams (bla), macrolide-lincosamid-streptogramin B (MLSB) (erm), aminoglycosides (aac), FCA (fluoroquinolone, quinolone, florfenicol, chloramphenicol, and amphenicol) (fca), colistin (mcr), vancomycin (van) and multidrug (mdr) (Table S1). The mostly frequently detected ARG classes in livestock waste include tet, sul, erm, fca, and bla (Fig. 2 and Table S2),13,14,15 which match the major classes of antibiotics used in animal husbandry (Table S3). Of these five ARG classes, tet and sul are generally the most abundant ARGs appearing in nearly all surveyed livestock waste (Fig. 2 and Table S2). Note that antibiotic usage varies considerably across livestock farm types and locations,16 and residual antibiotics are frequently detected in livestock wastes at widely varying concentrations (μg/kg to mg/kg in manure solids and ng/L to μg/L in wastewater) (Table S4), which leads to considerable differences in the selective pressure for ARGs.17 Here, we discuss the variability in the occurrence and abundance of ARGs in livestock waste, and the underlying factors that influence this variability.
ARGs abundance in livestock waste from swine and chicken farms are higher (by three to five orders of magnitude) than in hospital and municipal waste, whereas the abundance of ARGs on cattle waste and fish ponds is similar to hospital and municipal waste. Some ARGs abundance data from solid waste (for livestock waste) or biosolids (for hospital and municipal wastewater) are still used in this figure because the density of solid wastes and wastewater are on the same magnitude (e.g., 1.02–1.06 g/cm3 for biosolids155 and 0.9–1.1 g/cm3 for livestock waste156), which can facilitate the comparison of ARG concentrations.
The abundance of ARGs in livestock waste is higher than in other reservoirs
ARGs are frequently detected in livestock wastes (including solids used for manure, wastewater, and lagoon slurry and sediments18,19,20) at much higher levels (up to 28,000 times21) than in background soil or upstream water. The abundance of total ARGs in untreated livestock waste (combined solid waste and wastewater) varies from 106 to 1011 copies/g dry weight or 106 to 1012 copies/mL (absolute abundance), and 10–3 to 10−1 copies/16S ribosomal RNA (rRNA; relative abundance) (Table S2). Wastewater from swine and chicken farms harbors three to five orders of magnitude more ARGs than that of hospital and municipal wastewaters, while the abundance of ARGs in cattle and fish wastewaters is comparable to those in hospital and municipal wastewaters (Fig. 2 and Tables S2 and S5). The higher ARG concentrations in livestock wastes than human waste may be due to higher levels of residual antibiotics22,23 (and thus higher selective pressure for ARB) from the consistent use of antibiotics for animal growth promotion and disease prevention. In contrast, residual antibiotic concentrations (and their corresponding ARGs) in wastes from hospitals and municipal settings would likely fluctuate around lower values in accordance to the lower human therapeutic usage of antibiotics.22
The abundance of ARGs in livestock waste varies amongst livestock farm types
Considerable differences in the abundances of ARGs in livestock waste amongst livestock types have been observed, which may be due to varying antibiotic usage and dosing patterns. Generally, swine and chicken waste show higher ARG abundances than cow and fish waste (Fig. 2), with absolute abundance of ARGs in swine farm wastewater being around three orders of magnitude higher than that in fish ponds.13 Chicken and swine manure also exhibit greater ARG diversity than cow manure.14 These differences may be due to the more intensive use of antibiotics for therapeutic, prophylactic, and metaphylactic purposes on swine and chicken farms (at 172 mg/population correction unit (PCU) for swine and 148 mg/PCU for chickens16) in comparison to fish and cattle farms (at 45 mg/PCU for cattle).16
Antibiotic dosing pattern as a function of animal life stage may be another driving factor. Swine receive higher antibiotic dosages early in their life stage and are gradually given lower concentrations over their lifetime.24,25 This is reflected in the livestock waste from finisher swine farms, which shows lower ARG abundances and diversity relative to that of sow and nursery farms.26,27 Although >50% of broiler chicken production in the USA does not use antibiotics,28 those that do administer antibiotics from about two months after hatching until slaughter, which may contribute to higher levels of ARG in chicken waste than other livestock types (Fig. 2). The reason for the relatively low levels of ARGs found in cattle waste is unclear, although we cannot rule out that this observation might reflect later-in-life exposure to antibiotic feed additives than poultry and swine. Similar to most livestock, cattle for meat and dairy production are provided antibiotics for therapeutic purposes as needed, and generally also receive antibiotics in their feed for disease prevention (mostly to prevent liver abscesses). However, this non-therapeutic use occurs later in life for cattle, when they reach the feedlot.29,30
The abundance of ARGs in livestock waste varies among and within countries
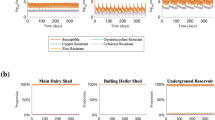
Of the 96 countries for which ARG abundance data have been reported, livestock waste in China, which is the largest producer and consumer of antibiotics,16 generally harbored the highest absolute abundance of ARGs (Fig. 3 and Table S6). Although there is not yet a clear linkage between global antibiotic usage and ARG abundance on a per country basis, it is informative to compare countries in this way. For instance, when comparing tet in swine wastewater in Shandong (China) and in Colorado (USA), ARG abundances in Shandong are 104-fold higher than in Colorado, even though antibiotic usage is relatively similar on a country wide basis (at 23% and 19% of the global consumption (2010), respectively).19,31 ARG abundances also depend on site-specific conditions within the countries. For example, the reported absolute abundance of ARGs in the cattle manure from Shandong, China are about 100-fold higher than that in Shaanxi, China (at 1011 copies/g).19,32 Variations in ARG abundance on this scale is also observed amongst various dairy farms in the Midwestern and Northeastern United States.15,17 These examples illustrate that for some countries ARG abundances may correlate with antibiotic use intensity and resulting residual antibiotic concentrations (Table S7), but wide variability may occur as a result of site-specific physical/chemical conditions that influence ARB growth and ARG propagation and attenuation dynamics.
ARGs have been detected in livestock farms in 95 countries/regions (coral-colored; data presented in Table S6). China, the United States, and the European Union are three most frequently studied countries/regions. The size (diameter) of filled pies represent the total ARGs absolute abundance (with different solid colors representing different ARGs classes) while hollow pies represent the ARGs relative abundance. China, which has a relatively high use of antibiotics for animal farming, exhibits a relatively high abundance of ARGs in livestock waste.
Pathways for ARG transfer from livestock waste to human pathogens
Considerable research has been conducted on the behavior and fate of ARB and ARGs discharged from animal husbandry to soil (e.g., through land application of manure and wastewater irrigation) and aquatic environments (e.g., through wastewater discharge and runoff), although the impact of discharging ARGs in treated livestock wastewater to aquatic systems (and associated ARG amplification and attenuation dynamics) has received less attention in the literature. Intracellular and free ARGs in surface and ground water, soil, and air19,33,34,35 can propagate through horizontal gene transfer (HGT) to indigenous bacteria.36,37 These ARB may eventually reach and colonize humans38,39 through multiple pathways (Fig. 4),40,41 causing acute infections or long-term silent colonization that can eventually evolve into an infection.11 Thus, it is important to consider ARG dissemination and attenuation mechanisms (and associated dynamics) in soil, water, air, and human gut environments.
Bacteria containing ARGs are discharged through drainage, treated wastewater, and solid waste from livestock farms into various receiving environments. HGT may occur between antibiotic-resistant bacteria (ARB) and indigenous bacteria by three main mechanisms: conjugation, transformation, and transduction, with the indicated frequencies based on literature.51,52,53,54,55,56,57,58 Some opportunistic pathogens harboring ARGs may be directly discharged into receiving environments. ARB may enter humans through ingestion (food/water) although such risks remain to be accurately quantified due to insufficient transport and public exposure data (indicated by dashed arrow). ARB can also directly enter humans through occupational exposure (e.g., water ingestion, food ingestion, and inhalation). Then ARB may reproduce in the human body (especially gut) and cause endogenous or exogenous infections.
Application of manure containing ARGs is the predominant initial propagation pathway in the environment,42 as it increases the diversity and abundance of ARGs in soil by up to 105-fold.19,43 ARG abundances in manured soils have been observed at up to 28,000 times more than those in un-manured soil, at 106 to 1010 copies/g soil (absolute abundance) and 10−4 to 10−1 copies/16S rRNA (relative abundance) (Table S8). Post one-time manure application, ARGs can persist in soil for >120 days19,44 and can take from three to six months to attenuate to levels less than or equal to background.45,46 However, variations in manure types and land application methods can significantly influence these outcomes and will be discussed as control strategies in section “ARGs removal by conventional livestock waste treatment”.
ARGs can also propagate through receiving water and air environments, although on a smaller scale. Generally, ARG abundances in receiving surface water environments range from not detected (ND) to 108 copies/L, which is up to 100-fold higher than upstream waters (Table S8). Once in receiving waterbodies, ARGs can accumulate in sediments through sedimentation and adsorption,34 and can be acquired by bacteria colonizing the intestinal mucus of fishes (at concentrations up to 10−1 copies /16S rRNA).47 ARGs have been frequently detected (frequency: 67%–100%, n = 124) in ground water near swine farms33 and as far as 250 m downstream from treatment lagoons.48 ARGs are also found in aerosols downwind of animal husbandry operations up to four orders of magnitude greater than at the source.49,50
Once ARGs reach new environments, propagation relies on the survival and proliferation of the original host (which differ in their ability to maintain, replicate, and transfer ARGs) and the likelihood of ARG acquisition by new hosts through HGT. Several studies have investigated the relative importance of the three HGT mechanisms for ARGs from livestock waste,51,52,53,54,55,56,57 which include conjugation, transformation and transduction. Conjugation is generally thought to be the most prevalent HGT mechanism (with a frequency ranging from 10−5 to 10−2)52,55 while transduction shows the lowest potential (at a frequency of 10−9 to 10−5)51,58 (Fig. 4). Note that these frequency estimations are based on laboratory results (e.g., culture-based experiments), and the transfer of ARGs in natural environments can be greatly influenced by many other factors, including selective pressure by residual antibiotics for ARG maintenance and replication,59 the relative abundance of mobile genetic elements (MGEs) and recipients, nutrient availability,52 and DNA form and availability (e.g., intracellular, free or adsorbed),60 which are further discussed below. For example, antibiotics have been found to increase bacterial competence, promoting HGT via transformation.61
MGEs, such as plasmids, integrons, and transposases, which are indispensable for conjugative HGT,62 have been frequently detected in livestock waste and show strong positive correlations with ARGs.19,20,63 A large body of evidence supports the transmission of ARGs via HGT in animal production environments.37,64,65,66,67 Note that many of these studies lacked the resolution of genomic data to appropriately type strains, and were unable to determine transmission directionality due to methodological limitations.68 Nevertheless, detection of ARGs in extracellular DNA (eDNA) and bacteriophages from livestock waste further supports a potential route for ARGs transfer via transformation and transduction,36,69 and several antibiotics used in the livestock industry are known to enhance HGT via the promotion of DNA damage and induction of the SOS response.70
In addition to available ARG hosts in receiving environments, the attenuation and propagation of ARGs depends on environmental conditions such as levels of residual antibiotics and heavy metals, substrate availability, oxygen, light/ultraiolet (UV) intensity and temperature.52,71,72 Accordingly, spatial and temporal variability in such conditions likely contributes to considerable variation in ARG abundances from one system to another (Table S8).
Positive correlations between ARGs and heavy metal concentrations have been reported.73,74 Heavy metals, which are usually present at concentrations two to three orders of magnitude higher than residual antibiotics in receiving environments,75,76 could exert significant selective pressure for ARGs that code for efflux pumps that excrete both heavy metals and antibiotics (Table S9).73 The co-transfer of ARGs and metal resistance genes via MGEs, has been observed in various environments, such as soil, sediment, and the human gut,77,78 and may also be a driving factor for the maintenance of ARGs.
ARB growth rates, which are a function of variables such as substrate concentrations and the presence of toxic compounds, and physical/chemical factors such as temperature, can also impact the abundance of ARGs in a given environment. Both positive and negative correlations between ARG concentrations and substrate availability31,43,79 have been observed, which may be due to differences in the ability of ARBs to utilize available substrates in the receiving environment. Anaerobic (and fermentative) conditions are thermodynamically less favorable to harvest metabolic energy for ARG maintenance and replication, which may result in greater metabolic burden and faster resistance plasmid curing in the absence of antibiotics.72 The presence of UV (about 5%)80 in visible light can remarkably remove ARGs81 and the production of highly cytotoxic reactive oxygen species can promote the attenuation of ARGs.82 High sub-lethal temperatures may enhance ARB metabolism, which would help alleviate the metabolic burden of ARG maintenance,83 and could induce prophages containing ARGs. Conversely, higher temperatures (e.g., 55 °C in an aerobic digester) would significantly eliminate mesophilic ARB.84
The human gut deserves special attention as it serves as the main reservoir for ARGs in the human body and harbors the highest relative abundance and richness of ARGs that are common to both human and non-human environments.85 The exact pathways by which ARB and ARGs reach human gut environments have not been comprehensively determined, but include oral ingestion of contaminated materials (food, waste, residual waste from occupational exposure or contaminated environments) and inhalation of airborne ARB.35,50,86,87 The fate of ingested bacteria, including ARB, is highly species- and strain-dependent and influenced by numerous factors, including pre-existing microbiome structure, medication status, host age and dietary context.88,89,90,91 Under normal dietary and physiological conditions, gut microbiome diversity and abundance typically represents a substantial barrier towards integration of ingested bacteria,92 and most ingested bacteria only transiently colonize the human body.11 However, antibiotic treatment or ingestion of antibiotic residues may substantially disrupt gut microbiome structure, facilitate the integration of ARBs by removing competition, and promote ARG proliferation. For example, selective pressure from residual antibiotics (e.g., >60 ng/kg in meat products93), has been shown to alter the type and increase the abundance of ARGs in the human gut.94 Furthermore, antibiotic-mediated perturbations to the gut microbiome have also been found to persist for years95,96 in some cases. Note that long-term colonization by ARB may also accelerate HGT in the human gut due to a favorable environment (e.g., high concentrations and proximity of ARG donors and recipients, stable temperature, physiochemical conditions, and nutrient availability)97 and contribute to the emergence of multidrug resistance genes by pathogens such as methicillin-resistant Staphylococcus aureus (MRSA).98
Most ARGs in the human gut are harbored by strictly anaerobic commensal bacteria and can be transferred to gut-dwelling opportunistic pathogens, though with a relatively low frequency.99 The impact of ARB colonizing human gastrointestinal environments is dependent on their virulence properties. Differences in virulence properties have been found between livestock-associated MRSA (LA-MRSA) isolates, and hospital-acquired (HA) or community-associated (CA) MRSA isolates. For example, LA-MRSA CC398 adheres to human cells less efficiently than CA- or HA-MRSA isolates, and has a lower disease burden. Nevertheless, LA-MRSA retains strong cytotoxic potential and certain subtypes have increased invasive potential. Moreover, LA-MRSA colonization of farm workers is believed to have led to introduction to hospitals and communities where HA- and CA-MRSA developed.100 On the other hand, several ARB harbored by livestock may not infect humans due to maladaptation,11 infection barriers101 and immune response.102 For example, a study of LA-MRSA showed that while its presence in farm workers was highly correlated with duration of animal contact, it did not persist long once animal contact had ceased.103 This suggests that LA-MRSA is a poorly persistent human colonizer. However, such risks remain to be accurately quantified due to insufficient transport and public exposure data, and the lack of a clear etiology between resistance observed in livestock waste versus clinical resistance.
ARGs removal by conventional livestock waste treatment
Although current livestock waste treatment technologies are not designed to remove ARGs and ARB specifically, the possible removal of ARGs during waste treatment from concentrated animal feeding operations (CAFO) is an area of interest (Table 1). Typically, in regions where there is a demand for manure, livestock waste treatment processes are more advanced (e.g., anaerobic digestion). On the other hand, some regions may lack sufficient farmland to assimilate or reuse the treated livestock waste (such as Southern China20,104), and the treated wastewater effluent is discharged directly to waterbodies. The general treatment process starts by storing livestock waste in lagoons. The waste is then treated through anaerobic digestion (AD) or composting, where it is separated into solid and liquid waste streams.42 The solids are then usually used as manure to fertilize agriculture soils, while the wastewater is next treated in a bioreactor (usually sequential treatment through an anaerobic reactor and aerobic activated sludge)20,105, through a constructed wetland (CW),106,107 or is discharged directly to waterbodies without treatment. Note that on free-range farms, livestock waste drains directly without treatment.108
AD has been shown to remove ARGs up to 4.23 log,109 but is not a panacea as it can increase the abundance of certain types of ARGs (e.g., sul, fca) up to 52 times110 (Table 1). This may be due to differences in operating temperature,110 abilities of potential hosts to maintain and transfer ARGs,111 and properties of the manure itself.110 Mesophilic or ambient-temperature AD are less efficient than thermophilic AD in removing antibiotics (e.g., tetracyclines and sulfonamides) and some mesophilic ARB (e.g., Bacteroidetes and Proteobacteria111). In addition, higher total volatile solids concentrations (>15%) in AD are conducive to more efficient removal of ARB and associated ARGs.112 Practices that generally mitigate ARG propagation in soil include manure stabilization with lime before application,113 vegetating soil with appropriate plants such as pasture that harbor (in their rhizosphere) indigenous microorganisms that compete with ARB,114 and using land application methods such as incorporation or injection (rather than broadcast, which involves manure distribution on the soil surface) to minimize ARG runoff.115 Furthermore, the addition of an adsorbent (e.g., biochar and wheat straw) can decrease the dissolved concentration of inhibitory residual antibiotics and heavy metals,32,116 and thus improve the digestion process and the associated removal of ARGs.
Composting can significantly decrease the diversity and abundance of ARGs in livestock manure (Table 1). Similar with AD, thermophilic composting has a higher ARG removal efficiency than mesophilic composting.63,111 Additives can further improve ARG removal during composting by reducing the bioavailability of heavy metals (thereby removing not only their inhibitory effect but also their co-selective pressure)117,118 and by decreasing the amount of available carbon (thereby reducing residual antibiotics and placing a metabolic burden on ARBs), resulting in a decreased abundance of ARGs.119,120
Biological processes have demonstrated 3.1 log ARG removal, but removal efficiencies vary among different ARG types (Table 1). Differences in operating conditions and removal rates of ARBs during treatment are probably a factor in this variabilty.105,121 CWs generally have higher ARGs removal efficiencies than bioreactors (Table 1), which may be due to the multiple removal mechanisms (e.g., filtration, physical and chemical adsorption, biodegradation) that take place during treatment.122 Optimizing operating conditions, such as using subsurface flow (instead of surface flow),106 applying proper fillings (e.g., bricks) and vegetation,106,123 and decreasing the hydraulic loading rate (10 cm/day),123 are essential for effective ARG removal by CWs.
Critical research needs
The global prevalence of ARGs in livestock waste and their apparent potential to disseminate to receiving environments and eventually transfer to humans underscore the need for a better understanding of how to reduce in-farm ARG proliferation (source), mitigate ARG discharge (elimination) and attenuate dissemination (fate and transport) (Fig. 5). The development of validated models to quantitatively assess the impact of ARG dissemination from livestock agriculture on human health, and a bridge in the communication gap between environmental and clinical microbiologists, are needed to mitigate the potential health risks of ARGs. Selected research opportunities to address these knowledge gaps are discussed below.
Reduce in-farm ARG proliferation
One of the most pressing needs is to control ARG levels at the source by improving livestock management strategies to reduce the abundance and diversity of ARGs in livestock waste. Modifying farm management approaches can suppress ARG proliferation and reduce the need for prophylactic antibiotic use. Management changes could include adjusting animal diets to minimize disease occurrence (e.g., dysentery in swine,124 rumen acidosis in cattle), decreasing human-to-animal contact, optimizing waste collection methods (e.g., scrape versus flush), increasing the frequency of waste collection, and creating containment areas for sick livestock to reduce the spread of disease.
A better understanding of the mechanisms underlying the maintenance and spread of antibiotic resistance is necessary to understand how antibiotic usage will impact ARG abundances in livestock waste and the environment and lay the foundations for appropriate policy. Although curtailing antibiotic usage in animal husbandry may decrease the abundance of some ARB in animals and multidrug-resistant bacteria by about 15% and 24%–32%, respectively,125 reducing antibiotic use may not necessarily decrease ARG abundances. For example, vancomycin-resistant Enterococci persisted for a long time after avoparcin was banned.126 Frequent conjugation of plasmids carrying ARGs can result in plasmid maintenance in a microbial community in the absence of antibiotics, even if the plasmid incurs a fitness cost.127 Thus, strategies to prevent conjugation and promote resistance plasmid loss may be needed in addition to halting antibiotic use to reduce ARG abundances.
Antibiotic alternatives, such as antimicrobial peptides (effector molecules that can kill pathogens), probiotics (live bacteria and yeasts) and prebiotics (a type of dietary fiber for probiotics),128 represent a promising strategy for selective microbial control. Some of these alternatives have been reported to embody most or all of the essential antibiotics functions (e.g., direct antibacterial activity, immunomodulation, nutrient adsorption), while also being less likely to induce bacterial resistance.129 For example, prebiotics have been reported to modify livestock gut microbial communities and promote growth by regulating metabolism, modulating immune systems, inhibiting pathogens, and establishing a favorable gut microbiome composition.130,131 Design of better drug delivery systems, such as nanomaterials129 or gel vaccine delivery systems132 and in ovo injection,129 have also been proposed to increase therapeutic efficacy. Improved understanding of the dynamics and interactions of microbes in response to the above alternatives should accelerate appropriate innovations.
Mitigate ARG discharge
Enhancing the ARG removal efficiency of current livestock waste treatments can minimize the abundance of ARGs discharged into receiving environments. For solid waste treatment, reasonable and feasible regulations and policy changes to guide manure storage and disposal practices and training of skilled workers to correctly operate treatment facilities (e.g., anaerobic digestion and composting) should be established. Additionally, further studies on the effects of additives (e.g., biochar, clay, surfactant) and optimization of operating conditions (e.g., temperature, solids, and hydraulic retention time) should be carried out to improve ARG removal efficiency through conventional waste treatment. More specifically, CWs design and operation should be optimized for contact time, dissolved oxygen levels, and vegetation selection to improve ARG removal efficiency. In addition, disinfection processes (e.g., chlorination and UV disinfection), which remove ARGs from drinking water and municipal wastewater to varying degrees of success,133,134 should be considered when treating wastewater from animal husbandry.
To do so, it is pertinent to first understand the role of HGT in ARG dissemination in receiving environments and quantify the specific transfer frequencies of conjugation, transformation, and transduction under different conditions (informed by lab experiments and bioinformatic tools). This would help elucidate and control the dissemination of ARGs into receiving environments. Additionally, by identifying the dominant hosts of ARGs and the taxa that are involved in the HGT of ARGs, it would be possible to discern ARG vectors that could be targeted (e.g., using bacteriophages135) to efficiently hinder ARG replication and propagation in livestock waste and receiving environments, including the human gut. Methods that link ARGs directly with their hosts, including long-read shotgun metagenomic sequencing,136 single cell fusion-PCR-based methods,137 and genomic crosslinking methods,138 may be used to advance the fundamental understanding of host–ARG–MGE relationships. Furthermore, the fate and transport of ARGs should be quantitatively assessed through direct measurements or mathematical modeling that considers ARG replication and HGT propagation dynamics, as well as natural attenuation mechanisms (e.g., ARG sedimentation, hydrolysis, and photolysis).
Attenuate ARG exposure to humans
A quantitative understanding of the likelihood of ARG transfer from ARBs in livestock waste to human pathogens is a critical knowledge gap for assessing the risks of ARGs from animal agriculture. Preventive measures, such as not applying livestock waste amended manure to soils used for human food crops, should be taken to reduce direct exposure routes of ARGs to human pathogens. Without a proper understanding of the mechanisms behind exposure and attenuation of ARGs in the human gut environment, only individual level preventative measures, such as proper hygiene and early treatment of bacterial infections, can be justified to reduce the spread of ARGs. Once quantitative models of exposure and risk are developed, monitoring efforts can be established in parallel to those, which already exist via the national antimicrobial resistance monitoring system.139
In order to develop quantitative models, which can inform risk, it is pertinent to address the communication gap between clinical and environmental microbiologists. Clinical investigations are largely focused on characterizing the risk of a specific ARB strain within a geopolitical region,140,141 and primarily rely on culture-based methods.142,143 In contrast, environmental risk assessments generally characterize ARG, ARB, and residual antibiotic concentrations on an ecosystem-wide basis, and thus mainly rely on culture-independent tools such as metagenomic sequencing and PCR20,37 to assess unculturable and low-prevalence species. This difference in approaches to risk assessment hinders data comparison and integration of environmental and clinical risk frameworks. Although environmental microbiologists use correlation studies to synthesize and interpret research efforts from the two groups,144,145 a unifying approach for comparison would be an ARB-based risk framework that facilitates collaboration and enhances integration of environmental and clinical risk assessment. Collaboration on this front would facilitate a more holistic etiological perspective to disambiguate the link between clinical and environmental antibiotic resistance.145,146
Compounding this communications gap, there exists contrasting findings in the comparison of clinically relevant isolates in clinical samples to those in environmental samples, which suggests that there is no consistent and direct association between human exposure to ARGs from the environment and clinical antibiotic-resistant infections.147,148,149 Therefore, environmental and clinical microbiology communities should strengthen communications (e.g., through specialized conferences) about future research directions and tools to tackle problems of shared concern (e.g., ARGs vectors, etiology of infections) and collaboratively develop a risk framework to improve data exchange. In doing so, a consensus can be reached to more effectively understand and combat the potential human health risks from environmental ARGs.
Conclusions
Animal husbandry is a major source of environmental ARGs as reflected by the relatively high concentrations found in various compartments impacted by livestock waste (i.e., air, water, soil). ARGs are prevalent in livestock farms worldwide with varying diversity and abundance; however, data on their absolute and relative abundance are scarce. Thus, augmenting the quantitative ARG database is critical to enhance risk assessment and to develop and validate models that help discern the relationship between antibiotic usage (or lack thereof) in animal husbandry and ARG abundance in receiving environments and clinical settings. Research is also needed to elucidate mechanisms driving in-farm ARG maintenance and transfer. Doing so would aid in understanding how ARGs propagate to new environments and hosts, which is critical to develop a clear etiology between resistance observed in animal husbandry and antibiotic-resistant human infections.
There is an imminent need for collaborative and cross-disciplinary research on ARG pathways from animal wastes to human gastrointestinal environments. Environmental and clinical microbiologists should work in tandem to understand how antibiotic usage drives (or slows) the abundance of ARGs in environmental and human gastrointestinal environments and define favorable environmental conditions and mechanisms for ARG propagation. Research is also needed to enhance source control, including higher ARG removal efficiency during livestock waste treatment (e.g., use of additives and optimization of operating conditions) and improved livestock and waste management strategies. By doing so, policy and operational adjustments can be implemented to mitigate the spread of ARGs from livestock waste while improving animal welfare, ensuring livestock profitability and protecting human health.
Data availability
The authors declare that all data supporting the findings of this study are available within the paper, the cited references, and its supplementary information files.
References
O’neill, J. Review on Antimicrobial Resistance: Tackling A Crisis for the Health and Wealth of Nations. 2014. (HM Government, London, 2016).
Xiao, Y. & Li, L. China’s national plan to combat antimicrobial resistance. Lancet Infect. Dis. 16, 1216–1218 (2016).
Government of India. National Action Plan on Antimicrobial Resistance. (Government of India, 2017).
Van Boeckel, T. P. et al. Global trends in antimicrobial resistance in animals in low- and middle-income countries. Science 365, eaaw1944 (2019).
Van Boeckel, T. P. et al. Reducing antimicrobial use in food animals. Science 357, 1350–1352 (2017).
Zhu, Y. G. et al. Continental-scale pollution of estuaries with antibiotic resistance genes. Nat. Microbiol. 2, 16270 (2017).
Woolhouse, M. E. J. & Ward, M. J. Sources of antimicrobial resistance. Science 341, 1460–1461 (2013).
Gullberg, E. et al. Selection of resistant bacteria at very low antibiotic concentrations. PLoS Pathog. 7, e1002158 (2011).
Rysz, M. & Alvarez, P. J. Amplification and attenuation of tetracycline resistance in soil bacteria: aquifer column experiments. Water Res. 38, 3705–3712 (2004).
Li, B. et al. Metagenomic and network analysis reveal wide distribution and co-occurrence of environmental antibiotic resistance genes. ISME J. 9, 2490–2502 (2015).
Manaia, C. M. Assessing the risk of antibiotic resistance transmission from the environment to humans: Non-direct proportionality between abundance and risk. Trends Microbiol. 25, 173–181 (2017).
Blair, J. M., Webber, M. A., Baylay, A. J., Ogbolu, D. O. & Piddock, L. J. Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 13, 42–51 (2015).
Huang, L. et al. Dissemination of antibiotic resistance genes (ARGs) by rainfall on a cyclic economic breeding livestock farm. Int. Biodeter. Biodeger. 138, 114–121 (2019).
Qian, X. et al. Diversity, abundance, and persistence of antibiotic resistance genes in various types of animal manure following industrial composting. J. Hazard. Mater. 344, 716–722 (2018).
Hurst, J. J. et al. Trends in antimicrobial resistance genes in manure blend pits and long-term storage across dairy farms with comparisons to antimicrobial usage and residual concentrations. Environ. Sci. Technol. 53, 2405–2415 (2019).
Van Boeckel, T. P. et al. Global trends in antimicrobial use in food animals. Proc. Natl Acad. Sci. USA. 112, 5649–5654 (2015).
Peak, N. et al. Abundance of six tetracycline resistance genes in wastewater lagoons at cattle feedlots with different antibiotic use strategies. Environ. Microbiol. 9, 143–151 (2007).
McKinney, C. W., Dungan, R. S., Moore, A. & Leytem, A. B. Occurrence and abundance of antibiotic resistance genes in agricultural soil receiving dairy manure. FEMS Microbiol. Ecol. 94, https://doi.org/10.1093/femsec/fiy010 (2018).
Han, X.-M. et al. Antibiotic resistance genes and associated bacterial communities in agricultural soils amended with different sources of animal manures. Soil Biol. Biochem. 126, 91–102 (2018).
Yuan, Q. B., Zhai, Y. F., Mao, B. Y., Schwarz, C. & Hu, N. Fates of antibiotic resistance genes in a distributed swine wastewater treatment plant. Water Environ. Res. https://doi.org/10.1002/wer.1125 (2019).
Zhu, Y. G. et al. Diverse and abundant antibiotic resistance genes in Chinese swine farms. Proc. Natl Acad. Sci. USA. 110, 3435–3440 (2013).
Sim, W. J. et al. Occurrence and distribution of pharmaceuticals in wastewater from households, livestock farms, hospitals and pharmaceutical manufactures. Chemosphere 82, 179–186 (2011).
Ekpeghere, K. I., Lee, J.-W., Kim, H.-Y., Shin, S.-K. & Oh, J.-E. Determination and characterization of pharmaceuticals in sludge from municipal and livestock wastewater treatment plants. Chemosphere 168, 1211–1221 (2017).
Dunlop, R. H. et al. Antimicrobial drug use and related management practices among Ontario swine producers. Can. Vet. J. 39, 87–96 (1998).
Dewey, C. E. Use of antimicrobials in swine feeds in the United States. Swine Health Prod. v. 7, 19-25-1999 v.1997 no.1991 (1999).
Brooks, J. P., Adeli, A. & McLaughlin, M. R. Microbial ecology, bacterial pathogens, and antibiotic resistant genes in swine manure wastewater as influenced by three swine management systems. Water Res. 57, 96–103 (2014).
Lu, X. M., Li, W. F. & Li, C. B. Characterization and quantification of antibiotic resistance genes in manure of piglets and adult pigs fed on different diets. Environ. Pollut. 229, 102–110 (2017).
National Chicken Council. Questions and Answers about Antibiotics in Chicken Production. https://www.nationalchickencouncil.org/questions-answers-antibiotics-chicken-production/ (2019).
Landers, T. F., Cohen, B., Wittum, T. E. & Larson, E. L. A review of antibiotic use in food animals: perspective, policy, and potential. Public Health Rep. 127, 4–22 (2012).
US Food & Drug Administration. Veterinary Feed Directive (VFD). https://www.fda.gov/animal-veterinary/development-approval-process/veterinary-feed-directive-vfd (2019).
McKinney, C. W., Loftin, K. A., Meyer, M. T., DAVIS, J. G. & Pruden, A. tet and sul antibiotic resistance genes in livestock lagoons of various operation type, configuration, and antibiotic occurrence. Environ. Sci. Technol. 44, 6102–6109 (2010).
Sun, W., Gu, J., Wang, X., Qian, X. & Tuo, X. Impacts of biochar on the environmental risk of antibiotic resistance genes and mobile genetic elements during anaerobic digestion of cattle farm wastewater. Bioresour. Technol. 256, 342–349 (2018).
Koike, S. et al. Monitoring and source tracking of tetracycline resistance genes in lagoons and groundwater adjacent to swine production facilities over a 3-Year Period. Appl. Environ. Microbiol. 73, 4813–4823 (2007).
Fang, H. et al. Dissemination of antibiotic resistance genes and human pathogenic bacteria from a pig feedlot to the surrounding stream and agricultural soils. J. Hazard. Mater. 357, 53–62 (2018).
Sancheza, H. et al. Antibiotic resistance in airborne bacteria near conventional and organic beef cattle farms in California, USA. Water Air Soil Poll. 227. https://doi.org/10.1007/s11270-016-2979-8 (2016).
Zhang, Y., Snow, D. D., Parker, D., Zhou, Z. & Li, X. Intracellular and extracellular antimicrobial resistance genes in the sludge of livestock waste management structures. Environ. Sci. Technol. 47, 10206–10213 (2013).
Fang, H., Wang, H., Cai, L. & Yu, Y. Prevalence of antibiotic resistance genes and bacterial pathogens in long-term manured greenhouse soils as revealed by metagenomic survey. Environ. Sci. Technol. 49, 1095–1104 (2015).
Forsberg, K. J. et al. The shared antibiotic resistome of soil bacteria and human pathogens. Science 337, 1107–1111 (2012).
Zeng, J. et al. Metagenomic insights into the distribution of antibiotic resistome between the gut-associated environments and the pristine environments. Environ. Int. 126, 346–354 (2019).
Reynaga, E. et al. Prevalence of colonization by methicillin-resistant Staphylococcus aureus ST398 in pigs and pig farm workers in an area of Catalonia, Spain. BMC Infect. Dis. 16, 716 (2016).
Heaney, C. D. et al. The prevalence of antibiotic-resistant Staphylococcus aureus nasal carriage among industrial hog operation workers, community residents, and children living in their households: North Carolina, USA. Environ. Health Persp. 125. https://doi.org/10.1289/ehp35 (2016).
Dungan, R. S., McKinney, C. W. & Leytem, A. B. Tracking antibiotic resistance genes in soil irrigated with dairy wastewater. Sci. Total Environ. 635, 1477–1483 (2018).
Chen, Q. et al. Long-term field application of sewage sludge increases the abundance of antibiotic resistance genes in soil. Environ. Int. 92–93, 1–10 (2016).
Gou, M. et al. Aerobic composting reduces antibiotic resistance genes in cattle manure and the resistome dissemination in agricultural soils. Sci. Total Environ. 612, 1300–1310 (2018).
Chen, Z. et al. Antibiotic resistance genes and bacterial communities in cornfield and pasture soils receiving swine and dairy manures. Environ. Pollut. 248, 947–957 (2019).
Lopatto, E. et al. Characterizing the soil microbiome and quantifying antibiotic resistance gene dynamics in agricultural soil following swine CAFO manure application. PLoS ONE 14, e0220770 (2019).
Marti, E. et al. Abundance of antibiotic resistance genes and bacterial community composition in wild freshwater fish species. Chemosphere 196, 115–119 (2018).
Chee-Sanford, J. C., Aminov, R. I., Krapac, I. J., Garrigues-Jeanjean, N. & Mackie, R. I. Occurrence and diversity of tetracycline resistance genes in lagoons and groundwater underlying two swine production facilities. Appl. Environ. Microbiol. 67, 1494–1502 (2001).
McEachran, A. D. et al. Antibiotics, bacteria, and antibiotic resistance genes: aerial transport from cattle feed yards via particulate matter. Environ. Health Persp. 123, 337–343 (2015).
Ling, A. L., Pace, N. R., Hernandez, M. T. & LaPara, T. M. Tetracycline resistance and Class 1 integron genes associated with indoor and outdoor aerosols. Environ. Sci. Technol. 47, 4046–4052 (2013).
Mazaheri Nezhad Fard, R., Barton, M. D. & Heuzenroeder, M. W. Bacteriophage-mediated transduction of antibiotic resistance in enterococci. Lett. Appl. Microbiol. 52, 559–564 (2011).
Guo, M. T., Yuan, Q. B. & Yang, J. Distinguishing effects of ultraviolet exposure and chlorination on the horizontal transfer of antibiotic resistance genes in municipal wastewater. Environ. Sci. Technol. 49, 5771–5778 (2015).
Torres-Barceló, C. The disparate effects of bacteriophages on antibiotic-resistant bacteria. Emerg. Microbes Infec. 7, 168 (2018).
Lu, N., Zilles, J. L. & Nguyen, T. H. Adsorption of extracellular chromosomal DNA and its effects on natural transformation of Azotobacter vinelandii. Appl. Environ. Microbiol. 76, 4179–4184 (2010).
Wang, X. et al. Bacterial exposure to ZnO nanoparticles facilitates horizontal transfer of antibiotic resistance genes. NanoImpact 10, 61–67 (2018).
Nielsen, K. M. et al. Natural transformation and availability of transforming DNA to Acinetobacter calcoaceticus in soil microcosms. Appl. Environ. Micro. 63, 1945–1952 (1997).
Ellison, C. K. et al. Retraction of DNA-bound type IV competence pili initiates DNA uptake during natural transformation in Vibrio cholerae. Nat. Microbiol. 3, 773–780 (2018).
Varga, M. et al. Efficient transfer of antibiotic resistance plasmids by transduction within methicillin-resistant Staphylococcus aureus USA300 clone. FEMS Microbiol. Lett. 332, 146–152 (2012).
Martinez, J. L. Antibiotics and antibiotic resistance genes in natural environments. Science 321, 365–367 (2008).
Yuan, Q.-B. et al. Redistribution of intracellular and extracellular free & adsorbed antibiotic resistance genes through a wastewater treatment plant by an enhanced extracellular DNA extraction method with magnetic beads. Environ. Int. 131, 104986 (2019).
Charpentier, X., Polard, P. & Claverys, J. P. Induction of competence for genetic transformation by antibiotics: convergent evolution of stress responses in distant bacterial species lacking SOS? Curr. Opin. Microbiol. 15, 570–576 (2012).
Parnanen, K. et al. Evaluating the mobility potential of antibiotic resistance genes in environmental resistomes without metagenomics. Sci. Rep. 6, 35790 (2016).
Qian, X. et al. Reducing antibiotic resistance genes, integrons, and pathogens in dairy manure by continuous thermophilic composting. Bioresour. Technol. 220, 425–432 (2016).
Hammerum, A. M. et al. Detection of sul1, sul2 and sul3 in sulphonamide resistant Escherichia coli isolates obtained from healthy humans, pork and pigs in Denmark. Int. J. Food Microbiol. 106, 235–237 (2006).
Giufre, M. et al. Escherichia coli of human and avian origin: detection of clonal groups associated with fluoroquinolone and multidrug resistance in Italy. J. Antimicrob. Chemother. 67, 860–867 (2012).
Leverstein-van Hall, M. A. et al. Dutch patients, retail chicken meat and poultry share the same ESBL genes, plasmids and strains. Clin. Microbiol. Infect. 17, 873–880 (2011).
Levy, S. B. Emergence of antibiotic-resistant bacteria in the intestinal flora of farm inhabitants. J. Infect. Dis. 137, 689–690 (1978).
Muloi, D. et al. Are food animals responsible for transfer of antimicrobial-resistant Escherichia coli or their resistance determinants to human populations? A systematic review. Foodborne Pathog. Dis. 15, 467–474 (2018).
Colomer-Lluch, M., Jofre, J. & Muniesa, M. Antibiotic resistance genes in the bacteriophage DNA fraction of environmental samples. PLoS ONE 6, e17549 (2011).
Zeng, X. & Lin, J. Factors influencing horizontal gene transfer in the intestine. Anim. Health Res. Rev. 18, 153–159 (2017).
Yuan, Q. B., Guo, M. T. & Yang, J. Monitoring and assessing the impact of wastewater treatment on release of both antibiotic-resistant bacteria and their typical genes in a Chinese municipal wastewater treatment plant. Environ. Sci. Proc. Imp. 16, 1930–1937 (2014).
Rysz, M., Mansfield, W. R., Fortner, J. D. & Alvarez, P. J. Tetracycline resistance gene maintenance under varying bacterial growth rate, substrate and oxygen availability, and tetracycline concentration. Environ. Sci. Technol. 47, 6995–7001 (2013).
Ji, X. et al. Antibiotic resistance gene abundances associated with antibiotics and heavy metals in animal manures and agricultural soils adjacent to feedlots in Shanghai; China. J. Hazard. Mater. 235–236, 178–185 (2012).
Zhang, J. et al. Impacts of addition of natural zeolite or a nitrification inhibitor on antibiotic resistance genes during sludge composting. Water Res. 91, 339–349 (2016).
Seiler, C. & Berendonk, T. U. Heavy metal driven co-selection of antibiotic resistance in soil and water bodies impacted by agriculture and aquaculture. Front. Microbiol. 3, 399 (2012).
Zhang, Q. Q., Ying, G. G., Pan, C. G., Liu, Y. S. & Zhao, J. L. Comprehensive evaluation of antibiotics emission and fate in the river basins of china: source analysis, multimedia modeling, and linkage to bacterial resistance. Environ. Sci. Technol. 49, 6772–6782 (2015).
Ma, L. et al. Metagenomic assembly reveals hosts of antibiotic resistance genes and the shared resistome in pig, chicken, and human feces. Environ. Sci. Technol. 50, 420–427 (2016).
Rosewarne, C. P., Pettigrove, V., Stokes, H. W. & Parsons, Y. M. Class 1 integrons in benthic bacterial communities: abundance, association with Tn402-like transposition modules and evidence for coselection with heavy-metal resistance. FEMS Microbiol. Ecol. 72, 35–46 (2010).
Wang, N. et al. A comprehensive analysis on spread and distribution characteristic of antibiotic resistance genes in livestock farms of Southeastern China. PLoS ONE 11, e0156889 (2016).
Cai, J. S., Huang, J. Y. & Lai, Y. K. 3D Au-decorated Bi2MoO6 nanosheet/TiO2 nanotube array heterostructure with enhanced UV and visible-light photocatalytic activity. J. Mater. Chem. A 5, 16422–16422 (2017).
Guo, M. T., Yuan, Q. B. & Yang, J. Ultraviolet reduction of erythromycin and tetracycline resistant heterotrophic bacteria and their resistance genes in municipal wastewater. Chemosphere 93, 2864–2868 (2013).
Maclean, M., MacGregor, S. J., Anderson, J. G. & Woolsey, G. Inactivation of bacterial pathogens following exposure to light from a 405-nanometer light-emitting diode array. Appl. Environ. Microbiol. 75, 1932–1937 (2009).
Graves, A. K. et al. Distribution of ten antibiotic resistance genes in E. coli isolates from swine manure, lagoon effluent and soil collected from a lagoon waste application field. Folia Microbiol. (Praha) 56, 131–137 (2011).
Diehl, D. L. & LaPara, T. M. Effect of temperature on the fate of genes encoding tetracycline resistance and the integrase of class 1 integrons within anaerobic and aerobic digesters treating municipal wastewater solids. Environ. Sci. Technol. 44, 9128–9133 (2010).
Pal, C., Bengtsson-Palme, J., Kristiansson, E. & Larsson, D. G. J. The structure and diversity of human, animal and environmental resistomes. Microbiome 4, 54 (2016).
Sanganyado, E. & Gwenzi, W. Antibiotic resistance in drinking water systems: Occurrence, removal, and human health risks. Sci. Total Environ. 669, 785–797 (2019).
Marti, R. et al. Impact of manure fertilization on the abundance of antibiotic-resistant bacteria and frequency of detection of antibiotic resistance genes in soil and on vegetables at harvest. Appl. Environ. Micro. 79, 5701–5709 (2013).
Wu, G. D. et al. Linking long-term dietary patterns with gut microbial enterotypes. Science 334, 105–108 (2011).
De Filippo, C. et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural. Afr. Proc. Natl Acad. Sci. 107, 14691–14696 (2010).
David, L. A. et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 505, 559 (2014).
Yatsunenko, T. et al. Human gut microbiome viewed across age and geography. Nature 486, 222 (2012).
Derrien, M. & van Hylckama Vlieg, J. E. T. Fate, activity, and impact of ingested bacteria within the human gut microbiota. Trends Microbiol. 23, 354–366 (2015).
Ben, Y. et al. Human health risk assessment of antibiotic resistance associated with antibiotic residues in the environment: a review. Environ. Res. 169, 483–493 (2018).
Gibson, M. K., Crofts, T. S. & Dantas, G. Antibiotics and the developing infant gut microbiota and resistome. Curr. Opin. Microbiol. 27, 51–56 (2015).
Dethlefsen, L. & Relman, D. A. Incomplete recovery and individualized responses of the human distal gut microbiota to repeated antibiotic perturbation. Proc. Natl Acad. Sci. 108, 4554–4561 (2011).
Jakobsson, H. E. et al. Short-term antibiotic treatment has differing long-term impacts on the human throat and gut microbiome. PLoS ONE 5, e9836 (2010).
Lerner, A., Matthias, T. & Aminov, R. Potential effects of horizontal gene exchange in the human gut. Front. Immunol. 8, 1630 (2017).
Krezalek, M. A. et al. Can methicillin-resistant Staphylococcus aureus silently travel from the gut to the wound and cause postoperative infection? Modeling the “Trojan Horse Hypothesis”. Ann. Surg. 267, 749–758 (2018).
Van Schaik, W. The human gut resistome. Philos. Trans. R. Soc. Lond., B, Biol. Sci. 370, 20140087 (2015).
Becker, K., Ballhausen, B., Kahl, B. C. & Kock, R. The clinical impact of livestock-associated methicillin-resistant Staphylococcus aureus of the clonal complex 398 for humans. Vet. Microbiol. 200, 33–38 (2017).
Groeger, S. E. & Meyle, J. Epithelial barrier and oral bacterial infection. J. Periodontol. 2000 69, 46–67 (2015).
Iwasaki, A. & Medzhitov, R. Control of adaptive immunity by the innate immune system. Nat. Immunol. 16, 343–353 (2015).
Graveland, H., Wagenaar, J. A., Bergs, K., Heesterbeek, H. & Heederik, D. Persistence of livestock associated MRSA CC398 in humans is dependent on intensity of animal contact. PLoS ONE 6, e16830 (2011).
Martinez, J., Dabert, P., Barrington, S. & Burton, C. Livestock waste treatment systems for environmental quality, food safety, and sustainability. Bioresour. Technol. 100, 5527–5536 (2009).
Tao, C. W. et al. Evaluation of five antibiotic resistance genes in wastewater treatment systems of swine farms by real-time PCR. Sci. Total Environ. 496, 116–121 (2014).
Huang, X. et al. Removal of antibiotics and resistance genes from swine wastewater using vertical flow constructed wetlands: Effect of hydraulic flow direction and substrate type. Chem. Engi. J. 308, 692–699 (2017).
Liu, L. et al. Elimination of veterinary antibiotics and antibiotic resistance genes from swine wastewater in the vertical flow constructed wetlands. Chemosphere 91, 1088–1093 (2013).
Graham, J. P. & Nachman, K. E. Managing waste from confined animal feeding operations in the United States: the need for sanitary reform. J. Water Health 8, 646–670 (2010).
Song, W. et al. Effects of different swine manure to wheat straw ratios on antibiotic resistance genes and the microbial community structure during anaerobic digestion. Bioresour. Technol. 231, 1–8 (2017).
Pu, C. et al. Impact of direct application of biogas slurry and residue in fields: In situ analysis of antibiotic resistance genes from pig manure to fields. J. Hazard. Mater. 344, 441–449 (2018).
Sun, W., Qian, X., Gu, J., Wang, X. J. & Duan, M. L. Mechanism and effect of temperature on variations in antibiotic resistance genes during anaerobic digestion of dairy Manure. Sci. Rep. 6, 30237 (2016).
Sun, W., Gu, J., Wang, X., Qian, X. & Peng, H. Solid-state anaerobic digestion facilitates the removal of antibiotic resistance genes and mobile genetic elements from cattle manure. Bioresour. Technol. 274, 287–295 (2019).
Munir, M. & Xagoraraki, I. Levels of antibiotic resistance genes in manure, biosolids, and fertilized soil. J. Environ. Qual. 40, 248–255 (2011).
Kyselkova, M. et al. Tetracycline resistance genes persist in soil amended with cattle feces independently from chlortetracycline selection pressure. Soil Biol. Biochem. 81, 259–265 (2015).
Joy, S. R. et al. Fate and transport of antimicrobials and antimicrobial resistance genes in soil and runoff following land application of swine manure slurry. Environ. Sci. Technol. 47, 12081–12088 (2013).
Zhang, J. et al. Effects of graphene oxide on the performance, microbial community dynamics and antibiotic resistance genes reduction during anaerobic digestion of swine manure. Bioresour. Technol. 245, 850–859 (2017).
Awasthi, M. K. et al. Application of metagenomic analysis for detection of the reduction in the antibiotic resistance genes (ARGs) by the addition of clay during poultry manure composting. Chemosphere 220, 137–145 (2019).
Cui, E., Wu, Y., Zuo, Y. & Chen, H. Effect of different biochars on antibiotic resistance genes and bacterial community during chicken manure composting. Bioresour. Technol. 203, 11–17 (2016).
Peng, S., Li, H., Song, D., Lin, X. & Wang, Y. Influence of zeolite and superphosphate as additives on antibiotic resistance genes and bacterial communities during factory-scale chicken manure composting. Bioresour. Technol. 263, 393–401 (2018).
Guo, A. et al. Effects of superabsorbent polymers on the abundances of antibiotic resistance genes, mobile genetic elements, and the bacterial community during swine manure composting. Bioresour. Technol. 244, 658–663 (2017).
Guo, M. T., Yuan, Q. B. & Yang, J. Insights into the amplification of bacterial resistance to erythromycin in activated sludge. Chemosphere 136, 79–85 (2015).
Huang, X. et al. Performance of vertical up-flow constructed wetlands on swine wastewater containing tetracyclines and tet genes. Water Res. 70, 109–117 (2015).
Chen, J. et al. Removal of antibiotics and antibiotic resistance genes from domestic sewage by constructed wetlands: Optimization of wetland substrates and hydraulic loading. Sci. Total Environ. 565, 240–248 (2016).
Erik, K. & Knudsen, B. Development of antibiotic resistance and options to replace antimicrobials in animal diets. Proc. Nutr. Soc. 60, 291–299 (2007).
Tang, K. L. et al. Restricting the use of antibiotics in food-producing animals and its associations with antibiotic resistance in food-producing animals and human beings: a systematic review and meta-analysis. Lancet Planet. Health 1, e316–e327 (2017).
Borgen, K. et al. Continuing high prevalence of VanA-type vancomycin-resistant Enterococci on Norwegian poultry farms three years after avoparcin was banned. J. Appl. Microbiol. 89, 478–485 (2000).
Lopatkin, A. J. et al. Persistence and reversal of plasmid-mediated antibiotic resistance. Nat. Commun. 8, 1689 (2017).
Ghosh, C., Sarkar, P., Issa, R. & Haldar, J. Alternatives to conventional antibiotics in the era of antimicrobial resistance. Trends Microbiol. 27, 323–338 (2019).
Suresh, G. et al. Alternatives to antibiotics in poultry feed: molecular perspectives. Crit. Rev. Microbiol. 44, 318–335 (2018).
Pourabedin, M. & Zhao, X. Prebiotics and gut microbiota in chickens. FEMS Microbiol. Lett. 362, fnv122 (2015).
Samanta, A. K. et al. Assessment of fecal microflora changes in pigs supplemented with herbal residue and prebiotic. PLoS ONE 10, e0132961 (2015).
Şenel, S. & McClure, S. J. Potential applications of chitosan in veterinary medicine. Adv. Drug Deliv. Rev. 56, 1467–1480 (2004).
Yuan, Q. B., Guo, M. T. & Yang, J. Fate of antibiotic resistant bacteria and genes during wastewater chlorination: Implication for antibiotic resistance control. PLoS ONE 10, e0119403 (2015).
Shi, P. et al. Metagenomic insights into chlorination effects on microbial antibiotic resistance in drinking water. Water Res. 47, 111–120 (2013).
Mathieu, J., Yu, P., Zuo, P., Da Silva, M. L. B. & Alvarez, P. J. J. Going viral: emerging opportunities for phage-based bacterial control in water treatment and reuse. Acc. Chem. Res. 52, 849–857 (2019).
Che, Y. et al. Mobile antibiotic resistome in wastewater treatment plants revealed by Nanopore metagenomic sequencing. Microbiome 7, 44 (2019).
Spencer, S. J. et al. Massively parallel sequencing of single cells by epicPCR links functional genes with phylogenetic markers. ISME J. 10, 427–436 (2016).
Stalder, T., Press, M. O., Sullivan, S., Liachko, I. & Top, E. M. Linking the resistome and plasmidome to the microbiome. ISME J. 13, 2437–2446 (2019).
U.S Food & Drug Administration. The National Antimicrobial Resistance Monitoring System (NARMS). https://www.fda.gov/animal-veterinary/antimicrobial-resistance/national-antimicrobial-resistance-monitoring-system (2018).
Chereau, F., Opatowski, L., Tourdjman, M. & Vong, S. Risk assessment for antibiotic resistance in South East Asia. BMJ 358, j3393 (2017).
Weinstein, R. A., Bonten, M. J. M., Austin, D. J. & Lipsitch, M. Understanding the spread of antibiotic resistant pathogens in hospitals: mathematical models as tools for control. Clin. Infect. Dis. 33, 1739–1746 (2001).
Locatelli, C. et al. Occurrence of methicillin-resistant Staphylococcus aureus in dairy cattle herds, related swine farms, and humans in contact with herds. J. Dairy Sci. 100, 608–619 (2017).
Dubin, K. A. et al. Diversification and evolution of vancomycin-resistant Enterococcus faecium during intestinal domination. Infect. Immun. 87. https://doi.org/10.1128/IAI.00102-19 (2019).
Angulo, F. J., Nargund, V. N. & Chiller, T. C. Evidence of an association between use of anti-microbial agents in food animals and anti-microbial resistance among bacteria isolated from humans and the human health consequences of such resistance. J. Vet. Med. B 51, 374–379 (2004).
Pärnänen, K. M. M. et al. Antibiotic resistance in European wastewater treatment plants mirrors the pattern of clinical antibiotic resistance prevalence. Sci. Adv. 5, eaau9124 (2019).
Finley, R. L. et al. The scourge of antibiotic resistance: the important role of the environment. Clin. Infect. Dis. 57, 704–710 (2013).
Grosso-Becerra, M. V. et al. Pseudomonas aeruginosa clinical and environmental isolates constitute a single population with high phenotypic diversity. BMC Genomics 15, 318 (2014).
Ekwanzala, M. D., Dewar, J. B., Kamika, I. & Momba, M. N. B. Systematic review in South Africa reveals antibiotic resistance genes shared between clinical and environmental settings. Infect. Drug Resist. 11, 1907–1920 (2018).
Mandel, G. N. The future of biotechnology litigation and adjudication. Pace Envtl. L. Rev. 23, 83–112 (2005).
Zhang, R., Wang, X., Gu, J. & Zhang, Y. Influence of zinc on biogas production and antibiotic resistance gene profiles during anaerobic digestion of swine manure. Bioresour. Technol. 244, 63–70 (2017).
Tien, Y. C. et al. Impact of dairy manure pre-application treatment on manure composition, soil dynamics of antibiotic resistance genes, and abundance of antibiotic-resistance genes on vegetables at harvest. Sci. Total Environ. 581–582, 32–39 (2017).
Awasthi, M. K. et al. The behavior of antibiotic resistance genes and their associations with bacterial community during poultry manure composting. Bioresour. Technol. 280, 70–78 (2019).
Ma, Z. et al. Long-term low dissolved oxygen accelerates the removal of antibiotics and antibiotic resistance genes in swine wastewater treatment. Chem. Eng. J. 334, 630–637 (2018).
ECDC (European Centre for Disease Prevention and Control), EFSA (European Food Safety Authority) and EMA (European Medicines Agency). ECDC/EFSA/EMA first joint report on the integrated analysis of the consumption of antimicrobial agents and occurrence of antimicrobial resistance in bacteria from humans and food-producing animals. EFSA J. 13, 4006 (2015).
Dammel, E. E. & Schroeder, E. D. Density of activated sludge solids. Water Res. 25, 841–846 (1991).
Kowalski, Z., Makara, A. & Fijorek, K. Changes in the properties of pig manure slurry. Acta Biochim. Pol. 60, 845–850 (2013).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (51608260) and State Key Laboratory of Pollution Control and Resource Reuse Foundation (NO. PCRRF16029), and by the U.S. National Science Foundation (NSF) Awards OISE:1545756 and 1820015.
Author information
Authors and Affiliations
Contributions
All authors contributed literature review, data analysis and intellectual input to this paper. Y.H. and Q.Y. drafted the first version with help from R.S. and content guidance from P.A. N.S., J.M., L.S., and P.A. provided critical input in trend analysis and research needs.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
He, Y., Yuan, Q., Mathieu, J. et al. Antibiotic resistance genes from livestock waste: occurrence, dissemination, and treatment. npj Clean Water 3, 4 (2020). https://doi.org/10.1038/s41545-020-0051-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41545-020-0051-0
This article is cited by
-
Staphylococcus aureus and biofilms: transmission, threats, and promising strategies in animal husbandry
Journal of Animal Science and Biotechnology (2024)
-
Mitigation potential of antibiotic resistance genes in water and soil by clay-based adsorbents
npj Materials Sustainability (2024)
-
Characterization of resistance and virulence factors in livestock-associated methicillin-resistant Staphylococcus aureus
Scientific Reports (2024)
-
Duplicated antibiotic resistance genes reveal ongoing selection and horizontal gene transfer in bacteria
Nature Communications (2024)
-
Puzzle: taking livestock tracking to the next level
Scientific Reports (2024)