Abstract
The ever-increasing need for high-quality drinking and process waters, and growing public awareness about possible contamination, drive efforts for the further development of automated control of water treatment plants. For example, membrane filtration processes and reverse osmosis in particular are generally regarded as a safe barrier for inorganic, organic, and microbial contamination. Yet, to ensure the final water quality and to increase the confidence of the end-user, intensive and preferably online monitoring should be further implemented as an early-warning tool to control membrane integrity and to prevent microbial regrowth in the distributing network. In this paper, we test the applicability of flow cytometry and cytometric fingerprinting for a full-scale water treatment plant. We demonstrate in a full-scale water treatment plant that flow cytometry can be used as online monitoring tool and that changes in water quality can be observed, which are not monitored by commonly used online quality parameters. Furthermore, we illustrate with ultrafiltration that process conditions impact the flow cytometric cell counts.
Similar content being viewed by others
Introduction
Bacteria in water are an important aspect of the water quality and may, when present in too high concentrations, lead to biofouling, microbiologically induced corrosion or even the spreading of pathogens. A close and online monitoring of the microbial quality of the water is thus a necessary tool to improve water quality and to reduce downstream costs or to mitigate health hazards. One of the main challenges is the unwanted growth of biofilms on surfaces known as biological fouling,1 which occurs mainly in recirculating systems such as cooling towers. Especially open cooling water systems provide a favorable environment for microorganisms because they scrub microorganisms from the air and concentrate the nutrients present in remaining water by evaporation, resulting in faster microbial growth.2 Biofilms can damage equipment through microbial induced corrosion (MIC), by clogging, and lead to an increased energy consumption due to decreased heat transfer.3,4 Biofouling and clogging leads to an increased pressure drop in ion exchangers and to increased resistance in membrane filters, which may also cause membrane breakthrough.5 Furthermore, pathogenic bacteria can nestle in these biofilms and contaminate the water through the natural shedding cycle of biofilms.6 A well-known example is the spreading of the pathogen Legionella pneumophila in the form of aerosols.7,8
Membrane processes such as microfiltration (MF), ultrafiltration (UF), nanofiltration (NF), and reverse osmosis (RO) are commonly used in water treatment, including reuse applications.9 MF is designed to retain most bacteria and suspended solids in the range of 0.1 to 5 μm. The other methods have pore sizes ranging between 10 to 100 nm (UF) to pore sizes smaller than 1 nm (NF and RO) and are, therefore, supposed to retain all bacteria. However, the passage of microorganisms through filtration membranes has been reported10 and research has demonstrated the existence of ultra-small bacteria in water.11,12 Also bacteria larger than the pore size are able to cross the membranes.10 Possible explanations are abnormalities in the membrane structure and oversized pores that were considerably larger than the manufacturers stated nominal pore size. Intensive use of membranes might lead to an enlargement of the pore sizes, and incompatible chemical cleaning (e.g., oxidative damage) can cause pore expansion as well.13,14 In addition, bacteria themselves can change in size in response to changing environmental conditions15 and may undergo size reduction in nutrient limited environments.16 The wall of most bacterial cells is not a rigid structure, but has a high flexibility and elasticity. This deformability, as well as the size of the bacteria, play an important role in their passage through filtration membranes.17 A last possibility is related to the breakthrough of membranes due to mechanical or chemical stress, resulting in the passage of unfiltered water.
Different techniques exist that characterize the aquatic bacteria. While the most commonly used method in drinking water is the heterotrophic plate count method (HPC), this method would be unsuitable for industrial applications due to the long incubation times and its labor intensive nature.18 A more convenient technique is the adenosine tri-phosphate (ATP) analysis, which provides an estimation of the active and viable biomass.19 The speed, robustness, easiness, and low cost make it a very appealing technique,20,21 yet ATP analysis is less frequently used than would be expected.19 A first issue is that ATP quantification is less precise for the low cell concentrations typically found water.19 Also, ATP measurements do not make the distinction between intra- and extracellular ATP. This can significantly alter the results as, in certain biological matrices, the extracellular ATP concentration can be several orders of magnitude higher than the intracellular ATP concentration.19,22 Finally, differences in species, cell sizes, and physiological states can alter the ATP concentration per cell making the conversion from ATP to biomass concentration difficult.23 Flow cytometry is a fast and robust method to determine bacterial concentration in liquids. With the use of appropriate dyes, it can determine viability and activity, similarly to ATP analysis. The single-cell resolution and the high throughput of the technique make it robust and by-pass the bias of converting the output to biomass concentration. In this paper, we applied online flow cytometry to monitor the microbial communities and dynamics in a full-scale water treatment plant. The water was monitored at different stages in the treatment plant. Our results illustrate the possible application of flow cytometry for microbial quality assurance and as tool to control membrane filtration processes.
Results
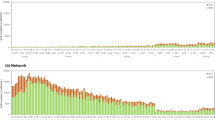
In the full-scale water treatment plant, the incoming surface water was monitored after passing through 300 µm self-cleaning strainers prior to the ultrafiltration step (Fig. 1). Cell concentrations fluctuated between 5.0*106 and 1.7*108 cells/mL (Fig. 2 b). After the first and the fourth day of measurement, two tailing peaks in cell density were observed. Besides the total cell concentration, also the Hill number diversity indices were calculated (D0, D1, and D2) according to Props et al.24 to characterize the dynamics in the microbial community composition. The two tailing peaks in cell concentration were reflected in all indices but most clearly in D2 that is most sensitive to the evenness of the abundance of the operational phenotypes (Supplementary Fig. 1). The Hill number diversity indices also showed dynamics, which were not clearly reflected by changes in the cell concentration (Fig. 2 a–d). Neither the peaks in the cell concentration measured by flow cytometry nor the peaks in feed water turbidity resulted in an increased conductivity (Fig. 2c). The conductivity measurements showed a large variability at the end of the experiments, which is also reflected in the flow cytometry data.
Online measurements on surface water. Flow cytometric Hill number diversity D2 expressed in arbitrary units (a), the corresponding bacterial concentration [cells/mL] (b), the turbidity [NTU] (c), and conductivity [µS/cm] (d) for surface water in function of time. The surface water was strained with 300 µm sieves prior sampling. Both the bacterial concentration and the diversity index D2 were calculated on the flow cytometry data. The sampling, staining, and incubation was fully automated. During the measurement period, the bacterial concentration increased twice. This increase was not observed in the either the turbidity or the conductivity that are typically used to monitor the water quality online. The diversity indices changed in function of the increased bacterial concentration but also revealed dynamics, which could not be explained by any of the measured parameters. It is hypothesized that this reflects the dynamics of the taxonomic microbial community composition24
In a second stage, the surface water was filtered by UF membranes with 100 nm pore size and the filtrate after UF was monitored for two different racks. The cell concentration after UF was approximately 3*104 cells/mL for rack 2 and 2.5*104 cells/mL for rack 3. Furthermore, a periodic pattern in the cell concentration was observed on both racks (Fig. 3c, d). For rack 2, a decrease in the concentrations was observed while for rack 3, an increase in the cell concentration was observed. These peaks coincide for both racks with a prolonged chemical cleaning of the membranes, which is performed by the recirculating a cleaning solution and characterized by an increased permeability (Fig. 3e, f). The cleaning procedure typically lasts 1 h and is done with a concentrated hypochlorite solution. UF rack 3 was chemically cleaned every 7 h, while UF rack 2 was only cleaned once daily. The reason for the different cleaning regimes is the age of the membrane units. The membrane lifetime of rack 3 is over 3 years while the membranes of rack 2 are only 1 year old. This is also reflected in the lower permeability of UF rack 3, in average 70 lmh/bar as compared to 100 lmh/bar for UF rack 2. Aside from chemical cleaning, short backwashes are programmed regularly as indicated by the frequent drops and surges in permeability every 30 min, for both racks. Yet, no effect of the backwashes was reflected in cell density. The cytometric Hill number diversity indices (D0, D1, and D2) reflected the decrease in cell density for rack 2 but also showed a decreasing trend during the measurement period. Also for rack 3, the influence of the cell density on the cytometric diversity indices could be observed though less clearly (Fig. 3a, b). Especially for diversity index D2 a clear change is observed during the measurement period, which is less visible for diversity index D0 (Supplementary Fig. 2).
Online measurements of UF filtrate. Flow cytometric Hill number diversity D2 expressed in arbitrary units (a, b) and the bacterial concentration [cells/mL] (c, d) of the UF filtrate in function of time. Both parameters were acquired with flow cytometry where the sampling, staining, and incubation was fully automated. Results are shown for UF rack 2 (left) and rack 3 (right). Both the cell concentration and the diversity D2 show different dynamics for rack 2 and rack 3. The main difference between the racks is their age and operation, which is reflected by the permeability [lmh/bar] (e, f). Rack 2 is relatively new and has a permeability of 100 lmh/bar while rack 3 is older and has a permeability of approximately 70 lmh/bar. Cleaning cycles for 1 h were performed daily for rack 2 and every 7 h for rack 3. Intermittently also backwashes were programmed
Similarly, the cell concentration and the Hill number diversity indices were monitored over time in the RO permeate water, which is used as cooling water, and in the recirculated cooling water return for reuse. A concentration of approximately 5*103 and 104 cells/mL was measured in the RO permeate water (Fig. 4). The overall bacterial concentration was slightly lower, though comparable to the cell concentrations found after UF. To confirm the results, samples were taken manually at different time points and resulted in a concentration of 8*102 cells/mL and 103 cells/mL. In the recirculating cooling water a concentration of 107 cells/mL was measured. For both RO and the recirculation water, no patterns were found in the cell concentrations. Furthermore, also no patterns could be discerned from the Hill number Diversity indices and no relationship could be established between the microbiological parameters and operational parameters (Supplementary Fig. 3).
Online measurements of RO permeate. Flow cytometric Hill number diversity D2 [A.U.] (a) and bacterial concentration [cells/mL] of the RO permeate in function of time (b). Both parameters were acquired with flow cytometry where the sampling, staining, and incubation was fully automated. The conductivity [µS/cm] of the water was monitored to assess water quality and membrane integrity (c)
Discussion
The bacterial concentration in the surface water measured after the strainers was, with a baseline concentration of 5.0*106 cells/mL, representative for surface water.25 A sudden increase and progressive decrease in the concentration was observed twice and as the water is pumped from a dock, boats roiling the water and the sludge near the treatment plant’s water inlet could cause these sudden changes in cell density. This could not be confirmed by the turbidity measurements alone as only the first peak in cell density coincides with peaks in turbidity. No clear explanation could be found for this discrepancy. This suggests that cell density is not always correlated to turbidity measurements, and that conductivity and turbidity alone are not sufficient as parameters for monitoring the incoming water quality of a membrane treatment plant. Total organic carbon (TOC) and dissolved organic carbon (DOC) measurements, for example, are important organic parameters to measure the load and fouling potential of the water, together with microbial water quality monitoring. DOC concentrations were measured periodically and showed seasonal fluctuations. In average, the concentration fluctuated between 6.07 ± 1.13 mg/L. Aside from cell concentrations, flow cytometry offers the possibility to calculate the Hill number diversity indices, which showed to be influenced by the cell concentration but which also reflected the dynamics in de microbial community composition as demonstrated by Props et al.24 Our results showed some dynamics in the microbial community, but mainly related to the sudden increase in cell concentration. In accordance to the publication of Prest et al.26 also the ratio of bacteria with high nucleic acid content (HNA) to low nucleic acid content (LNA) could be calculated for the waters. As demonstrated by Besmer et al.,27 this approach is complementary or equivalent to ours as illustrated by our results (Supplementary Fig. 4).
Subsequently the water was filtered by UF and concentrations of approximately 3*104 cells/mL for rack 2 and 2.5*104 cells/mL for rack 3 were measured. Though the pore size of the racks was 100 nm, not all bacteria in the water could be removed. As mentioned above, there are several possible explanations for the occurrence of bacteria after filtration. Aside from an impaired membrane integrity, also the presence of small bacteria in the surface water could cause the higher concentrations of bacteria. Luef et al.12 showed that ultra-small bacteria or ultramicrobiota can be found in ground water, Ghai et al.28 showed the same for sea water, and Wang et al.29 demonstrated their presence in surface water. The range and dispersion of these small bacteria is still unclear but Ghai, et al.28 estimated their relative abundance to be ~ 4% while Wang et al.29 estimated their relative abundance to be ~ 0.2% of the aquatic bacteria. Assuming that all bacteria after UF are ultra-small bacteria, their relative abundance would be ~ 1% for this surface water entering the treatment plant, which is in accordance with results cited above. The HNA to LNA ratio was also calculated and shows that the UF filtrate contained predominantly LNA bacteria, which is also in accordance to what could be expected from ultra-small bacteria (Supplementary Fig. 5). However, to proof this, confirmation with molecular techniques and microscopy is necessary. The membrane cleaning with a concentrated hypochlorite solution showed to have a different effect on the detection of bacteria for both membranes (Fig. 3b–e). While a decrease in the concentration was observed for rack 2, an increase in the cell concentration was observed for rack 3. Hypochlorite, an oxidizing agent, bleaches fluorochromes such as SG, making the bacteria undetectable with flow cytometry, which explains the results for UF rack 2. Despite the cleaning, filtration units get dirtier over time and an accumulation of organic compounds, inorganic precipitates, and biofilms reduces the overall efficiency of the filtration unit and increases the need for maintenance as for rack 3. Because of the higher degree of fouling on UF rack 3, we hypothesize that, during the chemical cleaning, the accumulated organics react much more with the hypochlorite than compared with the cleaning of a much younger and cleaner membrane unit like rack 2. As a consequence, less or no free chlorine is available to bleach SG or to kill bacteria. As bacteria detach from the membranes and the pipes during the chemical cleaning, even an increase in the cell concentration is visible in the recirculating cleaning solution.
In the RO permeate, the bacterial concentration fluctuated between 5*103 and 104 cells/mL, which is slightly lower, though comparable to the cell concentrations found after UF. The Hill number diversity indices or the HNA to LNA ratio also showed no noteworthy changes, which can be related to the short measurement period (Supplementary Figs. 3 and 6). In contrast to UF, it is unlikely that bacteria would pass through the membranes with pores smaller than 1 nm unless membranes are impaired. Dewettinck et al.30 and Kumar et al.31 reviewed the importance of membrane integrity monitoring for RO membranes and reported that challenge tests resulted in a log removal value (LRV) for bacteria between 2.9 and 5.4. Considering that the bacterial concentration in the RO concentrate can mount up to 106 CFU/mL,32 we estimate the bacterial concentration to be 108 cells/mL, assuming that only 1% of the bacteria can be cultured.20,32 The cell concentration we measured after RO would then result in a LRV of four, which is in accordance to what was reported before.31 An alternative hypothesis for the presence of bacteria after RO filtration could be related to the regrowth of bacteria in the distribution system due to the presence of organic carbon in the permeate and the subsequent shedding of bacteria from biofilms formed on the permeate side of the filter units. Indeed, periodic analysis of the DOC concentration in the RO permeate showed an average concentration of 0.47 ± 0.23 mg/L DOC. These results, however, do not indicate whether the DOC passed the RO membranes or is the result of leaching from the plastic RO permeate piping (Supplementary Fig. 5). Tang33 reported the presence of biofilms at the permeate side of RO membranes in dairy industry but, to our knowledge, no reports of biofilms on the permeate side of RO membranes have been published in the field of water treatment. To measure a concentration of 104 cells/mL in the bulk water, an estimated biofilm density of 107 cells/cm² is required considering that the RO module comprised 126 filter units, and assuming a biofilm growth rate of 0.03 per day.34 This biofilm density appears possible when comparing it to the biofilm density in drinking water distribution systems.35 Finally, a possible contamination of the sampling line and sampling port cannot be excluded but manually sampled controls resulted in similar concentrations and suggest the absence of contamination in, e.g., the tubing.
A bacterial concentration of approximately 107 cells/mL was measured for the recirculation water, which is likely caused by microbial regrowth after recirculation in the open cooling tower.
For all stages in the water treatment plant, flow cytometry showed that bacteria were present in concentrations above the expected level. Membrane processes commonly used for water production and reuse applications cannot be expected to produce pure and sterile water as small particles and molecules can pass through the membranes in several ways as discussed above. Routine membrane integrity tests are thus necessary to ensure the proper functioning of the membranes. Direct integrity tests such as the pressure decay test, which measures the rate of decline of pressure across a membrane require the membrane unit to be offline. Therefore, indirect methods are chosen for continuous online monitoring during production. Turbidity is an example of such an indirect test but we show here that it is not sufficient to monitor the microbial quality of the water. Based on our findings, we argue that cell concentrations, measured with flow cytometry, give additional information about the water quality, which other online techniques cannot provide. In this respect, flow cytometry could be used as a method to control process operation. Furthermore, it can serve as a tool to understand and control bacterial regrowth in the distribution networks and closed cooling circuits in order to ensure the final water quality for the end-user. Our results also showed that cytometric fingerprints, in the form of Hill number diversity indices, reflected the most important changes in cell concentrations but also showed additional dynamics, which are probably related the microbial community composition. The changes in community composition could not be confirmed in this paper. More variation should be expected over longer periods of time (i.e., months) as research has shown that fresh water communities also exhibit important seasonal community composition fluctuations.36
To conclude, we showed that flow cytometry is a good method to monitor fresh water communities online and automatically. We demonstrated in a full-scale water treatment plant that bacteria are present after every filtration step, even RO, and that bacterial concentrations in the water are directly related to process operations. For example, we showed that prolonged and chemical membrane cleaning can lead to an increase of bacterial concentrations in the recirculating cleaning solution, while backwashing did not affect the cell concentration in the filtrate water. As bacterial concentrations and the Hill number diversity indices gave a better insight in the microbiological dynamics than, e.g., turbidity measurements alone, we suggest that online flow cytometry could be used to indirectly monitor the membrane integrity and the changed microbial quality in the final water.
Methods
Online measurements
Online flow cytometry (FCM) measurements were performed at an industrial water treatment plant (Fig. 1). At this production site, four different water qualities are produced from brackish surface water (Rodenhuizendok, Port of Gent, Belgium). During a first measuring period of 6 days, surface water was monitored after pre-filtering through 300 μm strainers. During a second period of 7 days, the UF filtrate was measured continuously. The UF racks are built each with 40 Microza hollow fiber UNA modules (Pall) with a pore size of 100 nm. UF rack 1 and 2 have 1-year-old membrane modules, while UF rack 3 has membrane modules of more than 3 years. In addition to frequent backwashing, all UF racks are chemically cleaned by regularly recirculating a concentrated NaOCl solution during one hour. Based on the membrane permeability, the cleaning frequency is regulated by operator defined settings. In this case, the frequency for UF rack 2 was approximately every 24 h while for UF rack 3 it was every 7 h due to the lower average membrane permeability (older membrane age). RO1D produces 90 m³/h single-pass RO permeate, using BW30XFR-400/34i brackish water membranes (Dow) in an two-stage configuration. The system is working at an 75% recovery rate, resulting in a < 100 µS/cm demineralized water quality, fit-for-use as cooling water. The cooling tower has an 1500 m³ open buffer, susceptible to contamination. After seven to eight times thickening, the cooling water ( < 800 µS/cm) is continuously discharged at about 7.5 m³/h from the tower and reused again as feed water for the UF in the water treatment plant (cooling water return).
Flow cytometry
Instrumentation and staining
An OnCyt© staining robot (Oncyt, Switzerland) coupled to an BD Accuri C6 Plus flow cytometer (BD Biosciences), as described by Besmer et al.37 was used for continuous FCM measurements. Every 16 min, a fresh sample was taken and stained. An optimized staining protocol was used from Van Nevel et al.38 Bacteria were stained with 10 µL/mL of SYBR Green I (SG, Invitrogen, 100x diluted in DMSO from stock) for total cell counting. The samples were then incubated for 20 min at 37 °C inside the online robot to optimize the staining.
Data analysis
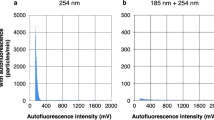
All data was extracted from the proprietary Accuri C6 CSampler software or FACSuite software in the flow cytometry standard (FCS 3.0) format and subsequently imported into R v3.4.039 through the functionality offered by the flowCore package v1.42.2.40 Data was first log transformed and then normalized by dividing all values by the maximum fluorescence intensity signal. No compensation was applied. Gating to reduce the background was performed in R studio using the flowCore package on both SG and SGPI stained samples. A 0.22 µm-filtered control was used to determine the position of the background. Based on this, a universal gate was constructed to remove as much background as possible. Next, a single-step discretization (‘‘binning’’) and Gaussian bivariate density estimation was performed on the selected parameters (green and red fluorescence, FSC-H and SSC-H) using the KernSmooth package v2.23.15.41 An equally spaced grid (binning grid) of 128 × 128 was fixed for each bivariate density estimation using the flowFDA package v1.0. All bivariate density estimations were concatenated to a one-dimensional feature vector, which we refer to as the fingerprint. Subsequently, phenotypic alpha diversity was calculated according to the publication of Props et al.24 where Hill number diversity indices are applied to describe the diversity of operational phenotypes within and between samples.
Data availability
Flow cytometry data is available on the open source Flow Repository database.42 Following dataset ID’s can be used: FR-FCM-ZYGV (surface water), FR-FCM-ZYZH (Ultrafiltration rack 2), FR-FCM-ZYH2 (Ultrafiltration rack 3), and FR-FCM-ZYHB (Reverse osmosis). The flowFDA package v1.0 available at https://github.com/lievenclement/flowFDA.
References
Chien, S. H. et al. Pilot-scale cooling tower to evaluate corrosion, scaling, and biofouling control strategies for cooling system makeup water. Rev. Sci. Instrum. 83, 024101 (2012).
Liu, Y. et al. Role of bacterial adhesion in the microbial ecology of biofilms in cooling tower systems. Biofouling 25, 241–253 (2009).
Meesters, K. P., Van Groenestijn, J. W. & Gerritse, J. Biofouling reduction in recirculating cooling systems through biofiltration of process water. Water Res. 37, 525–532 (2003).
Bott, T. R. Fouling in Heat Exchangers. (Elsevier Science B.V., 1995).
Mcdonogh, R., Schaule, G. & Flemming, H. C. The permeability of biofouling layers on membranes. J. Membr. Sci. 87, 199–217 (1994).
Wingender, J. & Flemming, H. C. Biofilms in drinking water and their role as reservoir for pathogens. Int. J. Hyg. Environ. Health 214, 417–423 (2011).
Dondero, T. J. et al. An outbreak of legionnaires’ disease associated with a contaminated air-conditioning cooling tower. New Engl. J. Med. 302, 365–370 (1980).
Keller, D. W. et al. Community outbreak of legionnaires’ disease: An investigation confirming the potential for cooling towers to transmit Legionella species. Clin. Infect. Dis. 22, 257–261 (1996).
Jiang, S. X., Li, Y. N. & Ladewig, B. P. A review of reverse osmosis membrane fouling and control strategies. Sci. Total Environ. 595, 567–583 (2017).
Ghayeni, S. B. S., Beatson, P. J., Fane, A. J. & Schneider, R. P. Bacterial passage through microfiltration membranes in wastewater applications. J. Membr. Sci. 153, 71–82 (1999).
Brown, C. T. et al. Unusual biology across a group comprising more than 15% of domain Bacteria. Nature 523, 208–U173 (2015).
Luef, B. et al. Diverse uncultivated ultra-small bacterial cells in groundwater. Nat. Commun. 6, 6372 (2015).
Goosen, M. F. A. et al. Fouling of reverse osmosis and ultrafiltration membranes: A critical review. Separ Sci. Technol. 39, 2261–2297 (2004).
Hai, F. I., Riley, T., Shawkat, S., Magram, S. F. & Yamamoto, K. Removal of pathogens by membrane bioreactors: A review of the mechanisms, influencing factors and reduction in chemical disinfectant dosing. Water-Sui 6, 3603–3630 (2014).
Chien, A. C., Hill, N. S. & Levin, P. A. Cell size control in bacteria. Curr. Biol. 22, R340–R349 (2012).
Egli, T. How to live at very low substrate concentration. Water Res. 44, 4826–4837 (2010).
Wang, Y. Y., Hammes, F., Duggelin, M. & Egli, T. Influence of size, shape, and flexibility on bacterial passage through micropore membrane filters. Environ. Sci. Technol. 42, 6749–6754 (2008).
De Roy, K., Clement, L., Thas, O., Wang, Y. & Boon, N. Flow cytometry for fast microbial community fingerprinting. Water Res. 46, 907–919 (2012).
Hammes, F., Goldschmidt, F., Vital, M., Wang, Y. & Egli, T. Measurement and interpretation of microbial adenosine tri-phosphate (ATP) in aquatic environments. Water Res. 44, 3915–3923 (2010).
Hammes, F. et al. Flow-cytometric total bacterial cell counts as a descriptive microbiological parameter for drinking water treatment processes. Water Res. 42, 269–277 (2008).
Velten, S., Hammes, F., Boller, M. & Egli, T. Rapid and direct estimation of active biomass on granular activated carbon through adenosine tri-phosphate (ATP) determination. Water Res. 41, 1973–1983 (2007).
Sakakibara, T., Murakami, S., Hattori, N., Nakajima, M.-o & Imai, K. Enzymatic treatment to eliminate the extracellular ATP for improving the detectability of bacterial intracellular ATP. Anal. Biochem. 250, 157–161 (1997).
Eydal, H. S. C. & Pedersen, K. Use of an ATP assay to determine viable microbial biomass in Fennoscandian Shield groundwater from depths of 3-1000 m. J. Microbiol. Methods 70, 363–373 (2007).
Props, R., Monsieurs, P., Mysara, M., Clement, L., & Boon, N. Measuring the biodiversity of microbial communities by flow cytometry. Methods Ecol. Evol. 7, 1376–1385 (2016).
Berney, M. et al. Rapid, cultivation-independent assessment of microbial viability in drinking water. Water Res. 42, 4010–4018 (2008).
Prest, E. I., Hammes, F., Kotzsch, S., van Loosdrecht, M. C. & Vrouwenvelder, J. S. Monitoring microbiological changes in drinking water systems using a fast and reproducible flow cytometric method. Water Res. 47, 7131–7142 (2013).
Besmer, M. D. et al. Laboratory-scale simulation and real-time tracking of a microbial contamination event and subsequent shock-chlorination in drinking water. Front. Microbiol. 8, 1900 (2017).
Ghai, R., Mizuno, C. M., Picazo, A., Camacho, A. & Rodriguez-Valera, F. Metagenomics uncovers a new group of low GC and ultra-small marine Actinobacteria. Sci. Rep. 3, 2471 (2013).
Wang, Y., Hammes, F., Boon, N. & Egli, T. Quantification of the filterability of fresh water bacteria through 0.45, 0.22, and 0.1 μm pore size filters and shape-dependent enrichment of filterable bacterial communities. Environ. Sci. Technol. 41, 7080–7086 (2007).
Dewettinck, T., Van Houtte, E., Geenens, D., Van Hege, K. & Verstraete, W. HACCP (Hazard Analysis and Critical Control Points) to guarantee safe water reuse and drinking water production—a case study. Water Sci. Technol. 43, 31–38 (2001).
Kumar, M., Adham, S. & DeCarolis, J. Reverse osmosis integrity monitoring. Desalination 214, 138–149 (2007).
Ridgway, H. F., Kelly, A., Justice, C. & Olson, B. H. Microbial fouling of reverse-osmosis membranes used in advanced wastewater-treatment technology—Chemical, bacteriological, and ultrastructural analyses. Appl. Environ. Microbiol. 45, 1066–1084 (1983).
Tang, X. Controlling Biofilm Development on Ultrafiltration and Reverse Osmosis Membranes Used in Dairy Plants. Doctor of philosophy thesis, Massey University (2011).
Boe-Hansen, R., Albrechtsen, H. J., Arvin, E. & Jorgensen, C. Bulk water phase and biofilm growth in drinking water at low nutrient conditions. Water Res. 36, 4477–4486 (2002).
Prest, E. I., Hammes, F., van Loosdrecht, M. C. M. & Vrouwenvelder, J. S. Biological stability of drinking water: Controlling factors, methods, and challenges. Front. Microbiol. 7, 45 (2016).
Pinto, A. J., Schroeder, J., Lunn, M., Sloan, W. & Raskin, L. Spatial-temporal survey and occupancy-abundance modeling to predict bacterial community dynamics in the drinking water microbiome. mBio 5, e01135–14 (2014).
Besmer, M. D. et al. The feasibility of automated online flow cytometry for in-situ monitoring of microbial dynamics in aquatic ecosystems. Front. Microbiol. 5, 265 (2014).
Van Nevel, S., Koetzsch, S., Weilenmann, H. U., Boon, N. & Hammes, F. Routine bacterial analysis with automated flow cytometry. J. Microbiol. Methods 94, 73–76 (2013).
R: A Language and Environment for Statistical Computing (Vienna, Austria, 2015).
Ellis B., Haaland P., Hahne F., Le Meur N., Gopalakrishnan N., Spidlen J., Jiang M. flowCore: flowCore: Basic structures for flow cytometry data. R package version 1.34.3. (2017).
Wand, M. KernSmooth: Functions for kernel smoothing for Wand & Jones (1995). R package version 2.23-15. (http://CRAN.R-project.org/package=KernSmooth) (2015).
Spidlen, J., Breuer, K., Rosenberg, C., Kotecha, N. & Brinkman, R. R. FlowRepository: A resource of annotated flow cytometry datasets associated with peer-reviewed publications. Cytom. Part A 81a, 727–731 (2012).
Acknowledgements
This work was supported by the project grant SB-131370 of the IWT Flanders and by the IMPROVED project, subvented by The interreg V “Vlaanderen-Nederland” program, a program for transregional collaboration with financial support from the European Regional Development Fund. More info: www.grensregio.eu (in Dutch). We would like to thank INDUSS and Water-link for providing access to the water treatment plant and particularly Stijn Merckx, Sven Versluys, Willem Verbanck, Kristin Van Hecke, and Nathalie Van Eeghem for their technical support. We would also like to thank Frederik Hammes and Michael Besmer (EAWAG, Switzerland) for the set-up of the OnCyt automated sampling and staining robot. Furthermore, we would like to thank A. Knezev (Het Waterlaboratorium, Haarlem, NL), and A. Verliefde (PaInT, Ghent University, Ghent, BE) for their valuable input.
Author information
Authors and Affiliations
Contributions
B.B., N.B., and B.D.G. designed the experiment and wrote the manuscript. B.B., A.N., and L.V. performed the experiments and B.B. and BDG analyzed the data and the related process operating conditions.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Buysschaert, B., Vermijs, L., Naka, A. et al. Online flow cytometric monitoring of microbial water quality in a full-scale water treatment plant. npj Clean Water 1, 16 (2018). https://doi.org/10.1038/s41545-018-0017-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41545-018-0017-7
This article is cited by
-
Development of gradient boosting-assisted machine learning data-driven model for free chlorine residual prediction
Frontiers of Environmental Science & Engineering (2024)
-
Artificial Intelligence-Based Optimization of Industrial Membrane Processes
Earth Systems and Environment (2021)
-
Online characterization of bacterial processes in drinking water systems
npj Clean Water (2020)
-
Computational flow cytometry of planktonic populations for the evaluation of microbiological-control programs in district cooling plants
Scientific Reports (2020)