Abstract
Since publishing our original reports on the safety and immunogenicity of a polyvalent DNA prime-protein boost HIV vaccine (PDPHV) which elicited high titer antibody responses with broad specificity, neutralizing activities to multiple HIV-1 subtypes, as well as poly-functional T cell responses, accumulated findings from other HIV vaccine studies indicated the important roles of Ig isotype distribution, Fc medicated functions and the persistence of memory immune responses which were not studied in previous PDPHV related reports. The current report provides further detailed characterization of these parameters in human volunteers receiving the PDPHV regimen. Antibody responses were assessed using IgG isotype and gp70-V1V2-binding ELISAs, peptide arrays, and antibody-dependent cellular cytotoxicity (ADCC) assays. B cell ELISPOT was used to detect gp120-specific memory B cells. Our results showed that the gp120-specific antibodies were primarily of the IgG1 isotype. HIV-1 envelope protein variable regions V1 and V2 were actively targeted by the antibodies as determined by specific binding to both peptide and V1V2-carrying scaffolds. The antibodies showed potent and broad ADCC responses. Finally, the B cell ELISPOT analysis demonstrated persistence of gp120-specific memory B cells for at least 6 months after the last dose. These data indicate that broadly reactive binding Abs and ADCC responses as well as durable gp120-specific memory B cells were elicited by the polyvalent heterologous prime-boost vaccination regimens and showed great promise as a candidate HIV vaccine.
Similar content being viewed by others
Introduction
Development of a safe and effective vaccine is crucial for the control of the HIV pandemic. After the moderate success of the heterologous viral vector prime-protein boost approach in the RV144 trial in Thailand1, the HIV vaccine field continues to explore the various combinations of prime and boost modalities to improve the immunogenicity of preventive HIV vaccine candidates2,3,4,5.
Antibodies are known to be the key elements in vaccine-induced protection against a wide range of human infectious diseases, but the protective mechanisms are diverse. In recent years, it became clear that candidate HIV vaccines may elicit immune protection via Fc-mediated antibody functions6,7,8,9,10. In particular, detailed biomarker analysis of the RV144 trial showed that the gp70-V1V2-specific antibody responses and Fc-mediated antibody functions inversely correlated with the risk of infection while there were no broadly neutralizing antibodies (bNAbs) detected in protected volunteers6,11. This is an important finding because many previous HIV vaccine studies have only focused on bNAbs12,13,14.
The importance of Fc-mediated antibody functions was also reported from the study results in NHP models and HIV infected patients. ADCC responses were reported to inversely correlate with virus set point in acute SIV infection15 and in vaccinated animals following SHIV challenge16,17. In HIV-1 infection, a direct role for ADCC responses was shown in controlling virus replication by delaying overt disease18,19,20. HIV mother-to-child transmission (MTCT) studies also demonstrated that passively acquired Ab mediating ADCC responses could reduce mortality in HIV infected infants21, and higher pre-existing ADCC responses against exposure strains associated with less likelihood of HIV-1 MTCT and lower morbidity in infected infants22. Therefore, one of the major tasks in the HIV vaccine field is to further improve on the ADCC responses achieved by RV1445.
RV144 used the ALVAC prime-protein boost vaccine approach which belongs to the heterologous prime-boost strategy23. Another heterologous prime-boost approach is the DNA prime-protein boost which has been studied by our team in the last two decades24,25,26,27,28 including our first HIV vaccine clinical trial DP6-00129.
Using DNA-encoded gp120 immunogens to prime the host immune system with the matched gp120 proteins boost is a promising approach leading to high titers of functional antibodies and cell-mediated immune responses29,30,31. More importantly, the polyvalent antigen formulation has been shown effective in eliciting antibody responses across different clades of HIV-1 in the DP6-001 trial. The gp120-specific serum IgG responses were robust and broadly cross-reactive against gp120 antigens from a wide range of major HIV-1 clades and the neutralizing activities from volunteers’ immune sera were also cross-reactive against pseudotyped viruses expressing Env antigens from clades of A, B, C, and AE29,31. Furthermore, the mAbs isolated from DP6-001 volunteers showed broad binding to both autologous and heterologous Env antigens and mediated potent ADCC response32. Since the publication of initial reports on the overall immunogenicity of DP6-001 vaccine, much new information has been learned in the HIV-1 vaccine field such as the roles of IgG isotypes and the identification of V1V2 region as the possible target for ADCC responses11,33. In the current report, data from new studies are presented to provide more detailed assessment of the additional humoral responses including IgG isotypes, recognition profiles of HIV-1 Env V1V2 region, ADCC response and the gp120-specific peripheral memory B cell responses from these volunteers. By including the previously reported neutralizing antibody data29,31, the polyvalent DNA prime-protein boost regimen shows its potential to elicit poly-functional antibody responses against HIV-1.
Results
Sources of volunteer samples
The human samples used in the current studies were collected from the previously reported DP6-001 study which was a phase 1 clinical trial testing safety and immunogenicity of a polyvalent DNA-prime/protein-boost preventive HIV vaccine29,30. The DNA vaccine components included five plasmids encoding gp120 Env proteins from HIV-1 clades A, B, C, and AE (two variants used for clade B) and one plasmid encoding Gag protein from HIV-1 clade C. The protein vaccine includes five CHO-produced recombinant gp120 Env proteins that exactly match those expressed by the Env DNA vaccines, mixed with the QS-21 adjuvant at the time of injection. The volunteers received three doses of the DNA vaccine at weeks 0, 4, and 12 followed by protein boosts at weeks 20 and 28. Group A received the DNA vaccines intradermally and Group B intramuscularly (both in saline) without electroporation or other facilitated DNA delivery, while both groups received protein vaccines intramuscularly along with QS-21.
IgG isotype analyses
We previously reported the dynamics of the overall gp120-binding IgG against the mixture of all five autologous gp120 proteins included in the vaccine formulation29. The individual total IgG titers at 2 weeks after the last immunization are shown in Supplementary Fig. 1. Both intradermal and intramuscular DNA priming immunizations resulted in median 2 × 105 binding titers to the vaccine-matched gp120 proteins.
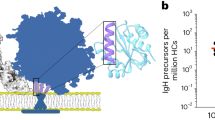
In the current study, we further explored the distribution of IgG isotypes elicited by the DP6-001 vaccine (Fig. 1a–c). Most volunteers were positive for IgG1, IgG3, and IgG4 antibodies against the consensus gp140-ConS antigen, while IgG2 antibodies were detected only in 10% (Group A) or 27% (Group B) of volunteers (Fig. 1c). When the magnitude of the response with each isotype was measured, the concentrations of IgG1 antibodies were found to be the highest among the four IgG isotypes and ranged between 103 and 104 ng/mL (Fig. 1a, b). IgG3 and IgG4 concentrations were 100-1000 fold lower than IgG1 concentrations. Most of the vaccinees also developed gp120-specific IgA responses (Fig. 1c), but the magnitude of IgA responses was not determined.
gp120 regions targeted by antibodies
Next, we explored which regions of gp120 were primarily targeted by the serum antibodies elicited by vaccination. Binding to pools of linear peptides derived from each variable or conserved region of the consensus Group M gp120 protein was measured for each volunteer. Mean OD values at 1:1000 serum dilution for each group are shown in Fig. 2. Consistent with previous reports of gp120-elicited antibodies, variable region 3 (V3) was the dominant target for antibody responses, and constant regions 1 and 5 (C1 and C5) were also targeted. However, we also observed substantial binding to peptide pools derived from V1 and V2 regions, binding to which was previously identified as an inverse correlate of risk in the RV144 vaccine efficacy study6,11,33. There were no significant differences in antibody specificity between the two study groups (Fig. 2a and Fig. 2b).
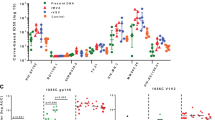
To better dissect the specificity of antibody targeting and its relationship with a breadth of recognition of diverse HIV-1 strains, peptide arrays were created corresponding to seven strains covering the wide diversity of circulating HIV-1 (Fig. 3). Antibodies from sera of vaccinated volunteers bound to peptides from all seven strains in conserved regions C1, C2, and C5, while peptides from C3 and C4 were less effectively recognized, probably due to their internal position within the protein.
Among variable regions, peptides from V3 region were most broadly recognized, with clade D being the only non-recognized strain. Recognition of peptides from other variable regions was more variable and of lower magnitude than that of V3. Nevertheless, in V1 and V2 regions, sera from vaccinated volunteers bound to peptides from A2, JRFL, B33, and LN40 strains, as well as in some cases to peptides from C2, D, and E strains demonstrating the breadth of the response elicited by the polyvalent formulation of the vaccine. There is no significant difference between Group A and Group B (Fig. 3a and Fig. 3b).
The peptide-binding results indicated the breadth of V1V2 response in the vaccinees. In the inverse correlate of risk analysis of the RV144 study, the assay that identified V1V2 binding as a correlate relied on binding to the V1V2 region fused to a gp70 scaffold. Therefore, we explored the binding specificity of the sera against similar scaffolds carrying V1V2 regions from diverse strains of HIV-1 (Fig. 4a, b). When tested against scaffolds carrying V1V2 regions from the strains that were used in the vaccine (“autologous”), the binding was detected for all five V1V2-carrying scaffolds. The titers against reagents with V1V2 regions from strains not used in the vaccine (“heterologous”) were comparable to those against autologous V1V2 regions or in some cases were even higher. These results also demonstrated the high degree of breadth and magnitude of the responses elicited by polyvalent DP6-001 formulation.
Antibody responses were measured against gp70 scaffold V1V2 antigens from either autologous gp120 or heterologous gp120 using sera from Group A (a) or Group B (b). Antibody responses against the cyclic V2 (A244) were also measured (c). Each bar represents the group mean titer with sera from each volunteer group.
Finally, we measured antibody binding to cyclic V1V2 region from Case-A2 strain, another reagent responses to which were identified as an inverse correlate of risk in the RV144 study, and found high binding titers (>103) in both ID (Group A) and IM (Group B) DNA priming groups (Fig. 4c).
Durability of gp120-specific memory B cell response
Previous study demonstrated that the DP6-001 vaccine elicited durable antibody titers that only slightly declined over the 6-month period after the last vaccination29. To follow up on this finding we explored the development and dynamics of gp120-specific memory B cell response in the vaccinees. PBMC samples from pre-bleed, two weeks after each vaccination and at 6 months after the last dose were stimulated with interleukin-2 and a polyclonal activator, R848, to induce memory B cells to differentiate into antibody-secreting cells. Total antibody-secreting, as well as antigen-specific B cells, were then quantified by B cell ELISPOT (Fig. 5). DNA priming alone only minimally elicited gp120-specific memory B cells. However, after protein boost, gp120-specific memory B cells were greatly expanded with the peak of 300-400 cells per million PBMCs, which is consistent with the robust antibody response at those time points as reported29. After 6 months, the number of gp120-specific memory B cells decreased slightly but remained at ~200 cells per million PBMC, which indicated a long-term B cell memory after the initial response to vaccination.
A mixture of five autologous gp120 proteins same as used in DP6-01 vaccine was coated on the plates. a ELISPOT readouts in duplicate from one volunteer sample collected at different time points of vaccination course are shown. b The dynamics of gp120-specific memory B cell responses is shown as the group average in spot/1 million cells with standard error.
ADCC responses
We first assessed ADCC response against target cells coated with various HIV-1 gp120 proteins (Fig. 6 and Supplementary Table 1). When tested against the gp120 proteins autologous to the gp120 antigens used as part of the DNA/protein vaccine formulation, we observed the highest response rate against cells coated with clade AE gp120 (93TH976) (7 out of 9 tested plasma samples from Group B and 8 out of 9 from Group A) followed by the response rate against the clade A gp120 (92UG037) (6 out of 9 tested serum samples for both Groups A and B). The response rate against the other two autologous clade B gp120 proteins (B715 and Bal) was lower, with three and five volunteers having a response, respectively.
ADCC-GTL assay was to detect ADCC against target cells coated either with the autologous gp120 proteins (a) or with the heterologous gp120 proteins (b). The numbers of positive responders and total volunteers are shown above the plots. The cut-off line is shown and the negative responders below the cut-off level were not included in the magnitude analysis.
When tested against the gp120 proteins heterologous to the gp120 antigens used in the vaccine formulation, the ADCC response rate was overall lower with 3 or 4 volunteers out of 9 showing a positive response to gp120 antigens from clades B (MN), C (TV1), and AE (CM235). Lastly, only one Group A and one Group B vaccine recipient had detectable ADCC response heterologous clade AE gp120 antigen (A244) (Supplementary Table 1).
When positive samples were considered, the magnitude of ADCC responses was high against A2, B715, and Bal (autologous, Fig. 6a) and MN and TV1 (heterologous, Fig. 6b) with mean titers over 1:10,000. ADCC Ab titers below 1:10,000 were detectable against AE (autologous, Fig. 6a) and CM235 (heterologous, Fig. 6b). In general, there is no significant magnitude difference between Group A and Group B, except for ADCC activities against CM235 which showed a high degree of variation among serum samples from volunteers in Group A but minimal variation in volunteers from Group B (Fig. 6b).
Similar studies were done to measure ADCC activities against target cells infected by infectious molecular clones (IMC) from clades B (Bal-IMC), C (TV1-IMC, 1086c-IMC), and AE (CM235-IMC), all of them different from the viral variants included in the vaccine. In this study, IL-15-treated PBMCs were used as effector cells (33). The frequency of positive ADCC responses against all four IMCs were shown in Supplementary Table 2. The ADCC response rate was >44% (4–8 serum samples out of 9 serum samples tested (Fig. 7)). The ADCC Ab titers against IMC infected cells ranged from 1:100 to close to 1:10,000. There was no difference in positive response rates or magnitude of responses between Group A and Group B.
For both ADCC against gp120-coated target cells and ADCC against IMC infected target cells, negative responders with ADCC activities below cut-off were not included for magnitude analysis.
Discussion
The current report expands the previous findings of high antibody titers elicited in phase 1 clinical trial DP6-001 of the polyvalent DNA prime-protein boost vaccine by demonstrating the breadth of these responses and their functionality.
We show that elicited antibodies have binding characteristics that have previously been identified as protective in an efficacy study RV1441,6,11,33. Specifically, V1V2 peptides and V1V2-based reagents from a wide diversity of HIV-1 clades were recognized by vaccinee sera. Our previously published data showed that the same vaccinee sera have neutralization activity against a wide range of HIV-1 variants from diverse clades29. These results indicate that antibodies elicited by DP6-001 formulation are able to recognize broad HIV-1 envelopes.
The new IgG isotype analysis included in the current report demonstrated that the majority of antibody responses are of IgG1 isotype, the predominant isotype observed in response to viral infections and vaccines. Our study also showed a high rate of IgG3 responses similar to the rate of IgG1 responses, however, the titers of IgG3 were lower than those for IgG1. Both IgG1 and IgG3 are known to have a high affinity for FcγRs and are potent activators of Fc-mediated functional activity. Literature showed that antibodies directed against the V1V2 region of gp120, in particular the IgG1 and IgG3 subclass mediating ADCC response, seem to play a predominant role in protection against HIV-1 acquisition34. On the other hand, the response rate of IgG2 and IgG4 were low in our study, similar to what were observed in other studies.
In the current report, we measured total IgG responses against V1V2, but not for different subtypes of Ig. The measurement of IgG3 against V1V2 could be performed in future studies. The rate of IgA responses among both study groups were also measured but not for their magnitude. We didn’t analyze the ADCC with IgA levels in the current study. The competition of IgA with ADCC responses in RV144 was demonstrated using monoclonal Ab. It is difficult to adjust the concentration of polyclonal IgA to compete IgG functions without assessing the specificity of the responses.
In recent years, based on results from both human and animal studies it became clear that candidate HIV-1 vaccines may elicit immune protection via Fc-mediated antibody functions6,7,8,9,10. In particular, detailed biomarker analysis of RV144 trial and HVTN 505 showed that Fc-mediated antibody functions inversely correlated with the risk of infection in this trial6,11,35 and with viral load set point35. In the current study, potent and broad ADCC activity was shown in DP6-001 immune sera, a response that was shown inversely correlated with the risk of infection in the RV144 study.
Our study demonstrated that DP6-001 volunteer sera had ADCC responses against IMC infected targeted cells while most of the previous HIV-1 vaccine studies were mainly directed ADCC against gp120-coated target cells36. Presumably, ADCC against IMC infected cells would target more on conformational epitopes while ADCC against gp120-coated targes may aim at CD4-induced epitopes7,37,38.
The broad reactivity observed in neutralizing antibodies29, gp120-binding, V1V2-binding, and ADCC assays supports the concept of polyvalency for HIV-1 vaccine, an approach that has not been sufficiently explored in the HIV vaccine field. The polyvalent formulation studied here resulted in responses that go beyond the vaccine-autologous strains and extend to diverse variants from different HIV-1 clades.
Finally, we demonstrate that gp120-specific memory B cells were generated in the course of vaccination and these cells persisted in vaccinated volunteers for at least 6 months after the last vaccination. These long-lasting, high-level memory responses elicited by the PDPHV regimen may not need a late boost as had been conducted in the RV305 study. This finding is consistent with the previously described sustained antibody titers29. High level and persisting antigen-specific memory B cell responses should be the results of effective germinal center B cell activation stimulated by DNA immunization39. In line with this finding, we previously reported that DNA immunization can improve the production of high-affinity antibodies40. While the HVTN lab published the ELISPOT method to monitor B cells in HIV-1 infected patients41, the current report would be the first to show the persistent Env-specific memory B cell responses induced by an HIV-1 vaccine.
The DP6-001 is the first-in-human study of the polyvalent DNA prime protein-boost vaccine29. The results indicate that this approach is able to induce humoral responses with high response rates, high titers and breadth, functional significance, and long durability. Manufacturing polyvalent DNA and protein vaccines may generate certain programmatic challenges, but the vaccine industry is very experienced in producing polyvalent vaccines. Several new batches of our PDPHV were manufactured in the last few years. The second generation of this vaccine has been recently tested in the HVTN 124 trial by the HIV Vaccine Trials Network which will provide further information about this approach.
Methods
Serum and PBMC samples from DP6-001 vaccinees
The serum and PBMC samples were from a closed phase I trial DP6-001 (NCT00061243)29. All participants gave written informed consent. The serum and PBMC samples were collected according to protocols approved by the institutional review board (IRB) at the University of Massachusetts Medical School (UMMS), USA29
Groups A or B of DP6-001 received three DNA immunizations at Days 0, 28, and 84, by either ID or IM needle injections, respectively, with a dose of 1.2 mg DNA containing five gp120 and one Gag plasmids formulated in 0.2 ml saline as previously reported29. Protein boosts contained a fixed dose of five recombinant gp120 proteins (0.375 mg) with adjuvant QS-21 administered twice at Days 140 and 196. The 5-valent gp120 recombinant proteins matched the polyvalent gp120 DNA vaccines. The serum samples used in the current report were collected on Day 0 and Day 210. The PBMC samples used in this study were collected on Day 0 and 14 days after each immunization, and at the study closeout.
Proteins and peptides
Recombinant gp120, and gp70-V1V2 proteins and peptides were used to evaluate the Env-specific antibody and B cell responses, including the autologous gp120 proteins from clades A (92UG037, A2), B (92US715, B715, and Bal), C (96ZM651, CZM), and AE (93TH976), and the heterologous gp120 proteins from clades B (MN), C (TV1) and AE (CM235 and A244); and gp140-ConS. Ten gp70-V1V2 proteins covered HIV-1 clades A (92UG037), B (92US715, Bal, JRFL, and Case-A2), C (96ZM965 and 93MW965.26), D (92UG021.16), and AE (93TH976 and consensus), and gp70 backbone protein was included as a negative control.
The Group M consensus gp120 peptides (15-mer with 11-overlap) were from NIH AIDS Reagent Program. Twelve overlap peptide pools were prepared to cover each gp120 region for ELISA assays: C1-N (14 peptides), C1-C (14 peptides), V1 (7 peptides), V2 (11 peptides), C2-N (11 peptides), C2-C (11 peptides), V3 (10 peptides), C3 (10 peptides), V4 (7 peptides), C4 (10 peptides), V5 (7 peptides) and C5 (6 peptides). The gp120 peptides printed on microarray slides were 15-mer with 11-overlap generated by JPT Peptide Technologies (Berlin, Germany) covering HIV-1 clade A (92UG037), B (JRFL, B33, LN4042), C (93MW965), D (92UG021), and AE (consensus). The cyclic V2 peptide (CSFNMTTELRDKKQKVHALFYKLDIVPIEDNTSSSEYRLINC) for clade AE (A244)33 was synthesized by EZBiolab (Carmel, IN).
Enzyme-linked immunosorbent assay (ELISA)
ELISAs were performed in 96-well microtiter plates. For gp120-specific antibody detection, the plates were coated with the mixture of five autologous gp120 antigens (92UG037, 92US715, Bal, 96ZM651, and 93TH976) at 1μg/ml in PBS (100 μL) at room temperature (RT) for 1 h. For gp70-V1V2-specific antibody detection, the plates were coated with individual gp70-V1V2 protein at 1 μg/ml in PBS (100 μL) at RT for 1 h. For detection of peptide-specific antibody responses, the plates were coated with peptides at 5 μg/mL at 4 °C overnight. After the relevant antigen coating, the ELISA was performed following previously established protocols29,31,43.
Linear epitope mapping by peptide microarray
The sera collected at 2 weeks after the second protein boost and pre-bleed as control were analyzed against a library of HIV-1 Env linear peptides using JPT microarray43. Briefly, the peptide microarray was performed using a Tecan HS4000 Hybridization WorkStation. Arrays were scanned using an Axon GenePix 4300 Scanner (Molecular Devices, Sunnyvale, CA). Images were analyzed using GenePix Pro 7 software (Molecular Devices).
Isotype binding antibodies
The Ig isotyping of Env-specific antibody responses in DP6-001 vaccinee sera was performed similarly to previously described11,44,45. HIV Env ConS gp140 antigen (Drs. Liao and Haynes, Duke University), purified IgG, IgG1, IgG2, IgG3, IgG4, and IgA proteins (Sigma-Aldrich, St. Louis, MO, USA, used as controls) were covalently coupled to carboxylated xMAP microspheres. HIV-specific antibody subclasses in serum samples were detected with biotin-conjugated mouse anti-human isotype antibodies. Detailed information can be found in the above references. All assays were run under Good Clinical Laboratory Practice compliant conditions.
Memory B cell ELISPOT
PBMCs from DP6-001 vaccinee was first treated to induce memory B cells to differentiate into antibody-secreting cells by a 5-day stimulation, similar to previous report41. Briefly, the PBMCs were cultured at 1 × 106 cells/ml in R-10 supplemented with 1 μg/ml R848 (Mabtech, Cincinnati, OH) and 10 ng/ml human IL-2 (Mabtech). The 96-well ELISPOT filter plates (Millipore) were coated with the mixture of five autologous gp120 antigens (A2, B715, Bal, Czm, and AE) at 10 μg/ml in PBS (100 μL) or goat anti-human IgG (Mabtech) at 4 °C overnight. After blocking, the stimulated PBMCs were added to the plates at 0.1 and 0.2 million cells/well, and incubated for 16 h. Plates were subsequentially incubated with biotinylated anti-human-IgG (Mabtech) for 2 h at RT, Streptavidin-ALP conjugate (Mabtech) for 1 h, and then developed using BCIP/NBT-plus substrate. The plates were washed with PBS between steps. The number of spots was scanned and analyzed using an automated ELISPOT counter.
ADCC assays
The HIV-specific antibody-dependent cell-mediated cytotoxicity (ADCC) activities in DP6-001 sera were evaluated using two types of assays that have been qualified for testing samples from clinical trials and detect responses related to two immunological spaces:36 (1) GranToxiLux (GTL)-ADCC assay against gp120 protein-coated target cells46,47, and (2) Luciferase-based ADCC (ADCC-Luc) assay against HIV-1 infectious molecular clone (IMC) infected target cells48. FACS gating strategy is provided in Supplementary Fig. 2.
The ADCC‐GTL assay was performed based on well-established protocols46,47. The gp120-coated CEM.NKRCCR5 CD4+ T cell line was used as the target cells. Briefly, individual gp120 protein-coated target cells were labeled with TFL4 and NFL1 (both from OncoImmunin, Gaithersburg, MD). Then, 104 target cells per well were added to 96‐well V‐bottom plates and incubated with the Granzyme B (GzB) substrate (OncoImmunin) and effector cells for 5 min at RT. Effector cells were PBMC obtained from an HIV-seronegative donor heterozygous for FcγR3A at position 158 (158 F/V) and for FcγR2A at position 131 (131H/R)49,50,51. PBMC was obtained by leukapheresis to collect enough cells for completion of the study with a single donation52, minimizing any potential for variability in the effector cell populations to influence the study outcome. PBMC were used at an effector cell to target cell ratio of 30:1. The antibodies were then added and tested after four-fold serial dilutions starting at 1:100; the plates were incubated an additional 15 min at RT, then for 1 h at 37 °C 5% CO2 following centrifugation for 1 min at 300 g. The plates were then incubated for 30 min at 4 °C and subsequently washed three times with 1% FBS PBS wash buffer. Well contents were then re‐suspended in 150 µl wash buffer and acquired directly with BD Fortessa flow cytometer (BD Biosciences, San Jose, CA) within 4 h using the High Throughput Sampler (HTS, BD Biosciences). Flow cytometry data analysis was performed using FlowJo 9.9.4 software (FlowJo, LLC., Ashland OR). Data are reported as the maximum proportion of cells positive for proteolytically active granzyme B (GzB) out of the total viable target cell population (maximum %GzB activity) after subtracting the background activity observed in wells containing effector and target cells in the absence of plasma. ADCC endpoint titers were determined by interpolating the last sera dilution above the previously established positive cutoff for this assay (8% GzB activity) using GraphPad Prism, version 7.0b software (GraphPad Software, Inc., La Jolla, CA) and were reported as reciprocal dilution.
ADCC activity was also determined by a luciferase (Luc)‐based assay based on well-established protocols47,48. CEM.NKRCCR5 target cells infected with HIV‐1 IMC encoding Renilla luciferase were incubated with PBMC obtained by leukapheresis to collect enough cells for completion of the study with a single donation as described for the GTL assay. PBMC effector cells treated or untreated with IL-15 and antibodies in ½ area opaque flat-bottom plates for 30 min at RT in duplicate wells. The plates were then centrifuged for 1 min at 300 g and subsequently incubated for an additional 5.5 h at 37 °C with 5% CO2. ADCC activity, reported as percent specific killing, was calculated from the change in Relative Light Units (RLU; ViviRen luciferase assay; Promega, Madison, WI) resulting from the loss of intact target cells in wells containing effector cells, target cells, and antibody samples compared to amounts in control wells containing target cells and effector cells alone according to the following formula: percent specific killing = [(number of RLU of target and effector well—number of RLU of the test well)/number of RLU of target and effector well] × 100. ADCC endpoint titers were determined by interpolating the last sera dilution above the positive cut-off for this assay (10% specific killing) using GraphPad Prism, version 7.0b software after subtracting the background activity observed for matched pre-vaccination samples, and were reported as reciprocal dilution.
Statistical analyses
One-way ANOVA was used to compare the average natural log-transformed Env-specific antibody magnitudes, IgG isotype concentrations, ADCC titers, and the number of B cells between groups A and B. The p-value < 0.05 (p < 0.05) was considered as a significant difference.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data that support the findings from this study are available from the corresponding author on reasonable request.
References
Rerks-Ngarm, S. et al. Vaccination with ALVAC and AIDSVAX to prevent HIV-1 infection in Thailand. N. Engl. J. Med. 361, 2209–2220 (2009).
Hu, X. et al. DNA vaccine-induced long-lasting cytotoxic T cells targeting conserved elements of human immunodeficiency virus Gag are boosted upon DNA or recombinant modified vaccinia Ankara vaccination. Hum. Gene Ther. 29, 1029–1043 (2018).
Viegas, E. O. et al. Optimizing the immunogenicity of HIV prime-boost DNA-MVA-rgp140/GLA vaccines in a phase II randomized factorial trial design. PLoS One 13, e0206838 (2018).
Malherbe, D. C. et al. Combination adenovirus and protein vaccines prevent infection or reduce viral burden after heterologous clade C simian-human immunodeficiency virus mucosal challenge. J. Virol. https://doi.org/10.1128/JVI.01092-17 (2018).
Excler, J. L. & Kim, J. H. Novel prime-boost vaccine strategies against HIV-1. Expert Rev. Vaccines 18, 765–779 (2019).
Haynes, B. F. et al. Immune-correlates analysis of an HIV-1 vaccine efficacy trial. N. Engl. J. Med. 366, 1275–1286 (2012).
Pollara, J. et al. HIV-1 vaccine-induced C1 and V2 Env-specific antibodies synergize for increased antiviral activities. J. Virol. 88, 7715–7726 (2014).
Bonsignori, M. et al. Antibody-dependent cellular cytotoxicity-mediating antibodies from an HIV-1 vaccine efficacy trial target multiple epitopes and preferentially use the VH1 gene family. J. Virol. 86, 11521–11532 (2012).
Bradley, T. et al. Pentavalent HIV-1 vaccine protects against simian-human immunodeficiency virus challenge. Nat. Commun. 8, 15711 (2017).
Ferrari, G., Pollara, J., Tomaras, G. D. & Haynes, B. F. Humoral and innate antiviral immunity as tools to clear persistent HIV infection. J. Infect. Dis. 215, S152–S159 (2017).
Yates, N. L. et al. Vaccine-induced Env V1-V2 IgG3 correlates with lower HIV-1 infection risk and declines soon after vaccination. Sci. Transl. Med 6, 228ra239 (2014).
Mascola, J. R. et al. Protection of macaques against vaginal transmission of a pathogenic HIV-1/SIV chimeric virus by passive infusion of neutralizing antibodies. Nat. Med. 6, 207–210 (2000).
Ko, S. Y. et al. Enhanced neonatal Fc receptor function improves protection against primate SHIV infection. Nature 514, 642–645 (2014).
Pegu, A. et al. A meta-analysis of passive immunization studies shows that serum-neutralizing antibody titer associates with protection against SHIV challenge. Cell Host Microbe 26, 336–346. e333 (2019).
Sun, Y. et al. Antibody-dependent cell-mediated cytotoxicity in simian immunodeficiency virus-infected rhesus monkeys. J. Virol. 85, 6906–6912 (2011).
Xiao, P. et al. Multiple vaccine-elicited nonneutralizing antienvelope antibody activities contribute to protective efficacy by reducing both acute and chronic viremia following simian/human immunodeficiency virus SHIV89.6P challenge in rhesus macaques. J. Virol. 84, 7161–7173 (2010).
Florese, R. H. et al. Contribution of nonneutralizing vaccine-elicited antibody activities to improved protective efficacy in rhesus macaques immunized with Tat/Env compared with multigenic vaccines. J. Immunol. 182, 3718–3727 (2009).
Wren, L. H., Stratov, I., Kent, S. J. & Parsons, M. S. Obstacles to ideal anti-HIV antibody-dependent cellular cytotoxicity responses. Vaccine 31, 5506–5517 (2013).
Wren, L. & Kent, S. J. HIV Vaccine efficacy trial: glimmers of hope and the potential role of antibody-dependent cellular cytotoxicity. Hum. Vaccin. 7, 466–473 (2011).
Smalls-Mantey, A. et al. Antibody-dependent cellular cytotoxicity against primary HIV-infected CD4+ T cells is directly associated with the magnitude of surface IgG binding. J. Virol. 86, 8672–8680 (2012).
Milligan, C., Richardson, B. A., John-Stewart, G., Nduati, R. & Overbaugh, J. Passively acquired antibody-dependent cellular cytotoxicity (ADCC) activity in HIV-infected infants is associated with reduced mortality. Cell Host Microbe 17, 500–506 (2015).
Thomas, A. S. et al. Pre-existing infant antibody-dependent cellular cytotoxicity associates with reduced HIV-1 acquisition and lower morbidity. Cell Rep. Med. 2, 100412 (2021).
Lu, S. Heterologous prime-boost vaccination. Curr. Opin. Immunol. 21, 346–351 (2009).
Xu, G. et al. Intramuscular delivery of a cholera DNA vaccine primes both systemic and mucosal protective antibody responses against cholera. Vaccine 27, 3821–3830 (2009).
Wang, S. et al. Heterologous HA DNA vaccine prime–inactivated influenza vaccine boost is more effective than using DNA or inactivated vaccine alone in eliciting antibody responses against H1 or H3 serotype influenza viruses. Vaccine 26, 3626–3633 (2008).
Gil, A. et al. DNA vaccine prime followed by a boost with live attenuated virus significantly improves antigen-specific T cell responses against human cytomegalovirus. Hum. Vaccin. Immunother. 9, 2120–2132 (2013).
Suguitan, A. L. Jr et al. Influenza H5 hemagglutinin DNA primes the antibody response elicited by the live attenuated influenza A/Vietnam/1203/2004 vaccine in ferrets. PLoS One 6, e21942 (2011).
Li, W., Wang, S. & Lu, S. Pilot study on the use of DNA priming immunization to enhance Y. pestis LcrV-Specific B cell responses elicited by a recombinant LcrV protein vaccine. Vaccines (Basel) 2, 36–48 (2013).
Wang, S. et al. Cross-subtype antibody and cellular immune responses induced by a polyvalent DNA prime-protein boost HIV-1 vaccine in healthy human volunteers. Vaccine 26, 3947–3957 (2008).
Bansal, A. et al. Multifunctional T-cell characteristics induced by a polyvalent DNA prime/protein boost human immunodeficiency virus type 1 vaccine regimen given to healthy adults are dependent on the route and dose of administration. J. Virol. 82, 6458–6469 (2008).
Vaine, M. et al. Profiles of human serum antibody responses elicited by three leading HIV vaccines focusing on the induction of Env-specific antibodies. PLoS One 5, e13916 (2010).
Costa, M. R. et al. Fc receptor-mediated activities of Env-specific human monoclonal antibodies generated from volunteers receiving the DNA prime-protein boost HIV vaccine DP6-001. J. Virol. 90, 10362–10378 (2016).
Zolla-Pazner, S. et al. Vaccine-induced IgG antibodies to V1V2 regions of multiple HIV-1 subtypes correlate with decreased risk of HIV-1 infection. PLoS One 9, e87572 (2014).
Kim, J. H., Excler, J. L. & Michael, N. L. Lessons from the RV144 Thai phase III HIV-1 vaccine trial and the search for correlates of protection. Annu Rev. Med. 66, 423–437 (2015).
Neidich, S. D. et al. Antibody Fc effector functions and IgG3 associate with decreased HIV-1 risk. J. Clin. Invest. 129, 4838–4849 (2019).
Huang, Y. et al. Diversity of antiviral IgG effector activities observed in HIV-infected and vaccinated subjects. J. Immunol. 197, 4603–4612 (2016).
Ferrari, G. et al. An HIV-1 gp120 envelope human monoclonal antibody that recognizes a C1 conformational epitope mediates potent antibody-dependent cellular cytotoxicity (ADCC) activity and defines a common ADCC epitope in human HIV-1 serum. J. Virol. 85, 7029–7036 (2011).
Veillette, M. et al. Interaction with cellular CD4 exposes HIV-1 envelope epitopes targeted by antibody-dependent cell-mediated cytotoxicity. J. Virol. 88, 2633–2644 (2014).
Hollister, K. et al. The role of follicular helper T cells and the germinal center in HIV-1 gp120 DNA prime and gp120 protein boost vaccination. Hum. Vaccin. Immunother. 10, 1985–1992 (2014).
Vaine, M., Wang, S., Hackett, A., Arthos, J. & Lu, S. Antibody responses elicited through homologous or heterologous prime-boost DNA and protein vaccinations differ in functional activity and avidity. Vaccine 28, 2999–3007 (2010).
Walsh, P. N. et al. Optimization and qualification of a memory B-cell ELISpot for the detection of vaccine-induced memory responses in HIV vaccine trials. J. Immunol. Methods 394, 84–93 (2013).
Vaine, M. et al. Two closely related Env antigens from the same patient elicited different spectra of neutralizing antibodies against heterologous HIV-1 isolates. J. Virol. 85, 4927–4936 (2011).
Wang, S. et al. Screening of primary gp120 immunogens to formulate the next generation polyvalent DNA prime-protein boost HIV-1 vaccines. Hum. Vaccin. Immunother. 13, 2996–3009 (2017).
Tomaras, G. D. et al. Initial B-cell responses to transmitted human immunodeficiency virus type 1: virion-binding immunoglobulin M (IgM) and IgG antibodies followed by plasma anti-gp41 antibodies with ineffective control of initial viremia. J. Virol. 82, 12449–12463 (2008).
Yates, N. L. et al. Multiple HIV-1-specific IgG3 responses decline during acute HIV-1: implications for detection of incident HIV infection. AIDS 25, 2089–2097 (2011).
Pollara, J. et al. High-throughput quantitative analysis of HIV-1 and SIV-specific ADCC-mediating antibody responses. Cytometry. A 79, 603–612 (2011).
Pollara, J. et al. Application of area scaling analysis to identify natural killer cell and monocyte involvement in the GranToxiLux antibody dependent cell-mediated cytotoxicity assay. Cytometry. A 93, 436–447 (2018).
Fisher, L. et al. Vaccine-induced antibodies mediate higher antibody-dependent cellular cytotoxicity after interleukin-15 pretreatment of natural killer Effector Cells. Front Immunol. 10, 2741 (2019).
Bruhns, P. et al. Specificity and affinity of human Fcgamma receptors and their polymorphic variants for human IgG subclasses. Blood 113, 3716–3725 (2009).
Koene, H. R. et al. Fc gammaRIIIa-158V/F polymorphism influences the binding of IgG by natural killer cell Fc gammaRIIIa, independently of the Fc gammaRIIIa-48L/R/H phenotype. Blood 90, 1109–1114 (1997).
Nimmerjahn, F. & Ravetch, J. V. Fcgamma receptors as regulators of immune responses. Nat. Rev. Immunol. 8, 34–47 (2008).
Garcia, A. et al. Leukopak PBMC sample processing for preparing quality control material to support proficiency testing programs. J. Immunol. Methods 409, 99–106 (2014).
Acknowledgements
Research reported in this publication was partially supported by NIH/NIAID grants R01 AI065250, HIVRAD P01 AI082274, IPCAVD U19 AI082676, R21/R33 AI087191, Duke Center for AIDS Research AI064518, and Bill & Melinda Gates Foundation, and P01 A1120756.
Author information
Authors and Affiliations
Contributions
G.F., G.D.T., and S.L. designed studies and drafted the manuscript. S.W., N.L.Y., J.P., S.S.O., D.H., G.H., and W.L. executed various experiments. Y.V. participated in data processing and manuscript drafting.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, S., Yates, N.L., Pollara, J. et al. Broadly binding and functional antibodies and persisting memory B cells elicited by HIV vaccine PDPHV. npj Vaccines 7, 18 (2022). https://doi.org/10.1038/s41541-022-00441-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41541-022-00441-9