Abstract
Vaccine-enhanced disease (VED) occurs as a result of vaccination followed by infection with virulent Mycoplasma pneumoniae. To date VED has prevented development of an efficacious vaccine against this significant human respiratory pathogen. Herein we report that vaccination of BALB/c mice with M. pneumoniae lipid-associated membrane proteins (LAMPs) induces lung lesions consistent with exacerbated disease following challenge, without reducing bacterial loads. Removal of lipid moieties from LAMPs prior to vaccination eliminates VED and reduces bacterial loads after infection. Collectively, these data indicate that lipid moieties of lipoproteins are the causative factors of M. pneumoniae VED.
Similar content being viewed by others
Introduction
Mycoplasma pneumoniae (Mp) is a highly contagious and widespread human respiratory pathogen causing over 2 million cases of community-acquired pneumonia (CAP) and ~100,000 adult hospitalizations annually in the United States1. Additionally, innate resistance to commonly used β-lactam antibiotics, increasing resistance to macrolide antibiotics, and mortality in certain demographic groups establish Mp as a high-risk pathogen capable of causing acutely severe disease2,3,4,5,6,7. Critically, no vaccine against Mp is available, as experimental vaccine-enhanced disease (VED) after infection has stymied further development.
M. pneumoniae VED was first reported in human volunteers in the 1960s. In these studies, prisoners and military recruits were vaccinated with formalin-inactivated or a serially passaged live-attenuated strain of Mp, then subsequently challenged with a virulent strain of the bacterium. While some protection was observed, many vaccinated individuals in both studies exhibited more severe clinical symptoms than those receiving a placebo control8,9. Recent studies by our group (and others) have since recapitulated VED in an animal model by utilizing live-attenuated or crude extract Mp vaccine candidates, respectively10,11,12,13. In these murine models, VED is characterized by more severe histopathology post-challenge, when compared to sham vaccinated, Mp challenged mice. VED has also been observed in vaccine candidates against other Mycoplasma species, making the identification of the causative factors a crucially important task for vaccine development against these atypical pathogens.
Results and discussion
Lipoproteins are the most abundant components of the lipid- associated membrane protein (LAMP) fraction
To identify the specific causative factor(s) for Mp VED, we utilized TX-114 phase partitioning to fractionate Mp proteins into three fractions: a LAMP hydrophobic detergent phase, an aqueous phase (Aq) harboring mostly hydrophilic cytosolic proteins, and an insoluble phase (Ins) containing the insoluble triton shell and a spontaneously forming, phospholipid-rich precipitate14. An initial pilot study indicated that vaccination of BALB/c mice with LAMPs resulted in VED after challenge with virulent Mp. We performed proteomic analysis to reveal lipoproteins, elongation factors, chaperones and chaperonins, and cytadherence proteins (previously found to induce strong antibody responses15,16,17) as the most abundant immunogenic and antigenic components of the LAMP fraction (Fig. 1a–f). Of these, lipoproteins are the most abundant (31.79%) (Fig. 1a) and include twenty-two lipoproteins representing all six Mp lipoprotein families (Fig. 1b, c). Family 2 lipoproteins, paralogs of the immunodominant Mycoplasma gallisepticum nucleotide-binding virulence factor MslA18,19, were the most abundant LAMP lipoproteins, making up 43.22% of the lipoprotein fraction (Fig. 1c). Lipoproteins not belonging to a multigene family were the second most abundant lipoproteins in the lipoprotein fraction (32.83%), followed by Family 4 lipoproteins (11.73%), Family 3 lipoproteins (5.53%) then Family 6 lipoproteins (4.36%) (Fig. 1c). Members of Family 5 lipoproteins (which are specific to Mp) were also identified in the fraction, albeit at minute relative quantities (0.67%) (Fig. 1c). Mycoplasma lipoprotein lipid moieties are known to be potent immuno-stimulators, inducing expression of inflammatory cytokines such as TNF-α, IL-6, and IL-1β following the recognition of their lipid moieties by Toll-like receptor complexes20. Given that these cytokines are frequently associated with immunopathology, we hypothesized that sensitization by Mp lipoprotein lipid moieties during vaccination induces VED upon challenge with virulent Mp.
a Breakdown of the major immunogenic/antigenic components of the LAMPs fraction as percentage of spectral hits. b Breakdown of the major immunogenic/antigenic components of the LAMPs fraction into number of unique genes represented in the fraction. Percent breakdown of c lipoproteins, d chaperones and chaperonins, e elongation factors, and f cytadherence-associated proteins represented in the LAMPs fraction.
Lipoprotein lipid moieties are responsible for VED
To test our hypothesis, TX-114-derived Mp fractions were treated with exogenous lipoprotein lipase to generate delipidated fractions (dLAMPs, dAq, and dIns), which could be tested for immuno-stimulation in vitro and as vaccine candidates in vivo. Successful delipidation was assessed via a TLR2 bioassay. In murine macrophages, LAMPs stimulated higher levels of TNF-α than Aq and Ins fractions. Delipidation significantly reduced LAMP-stimulated TNF-α production, notably to levels lower than those stimulated by Aq or Ins fractions (Fig. 2b). Surprisingly, delipidation also reduced Aq fraction-stimulated TNF-α production, suggesting that some lipoproteins may have sequestered to the Aq phase during TX-114 phase partitioning. Delipidation did not affect the ability of the Ins fraction to stimulate TNF-α production in vitro (Fig. 2b).
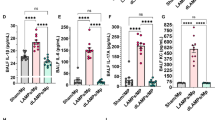
a Illustration of experimental timeline. b Supernatant TNF-α levels produced by murine J774A.1 macrophages stimulated by intact or lipase treated M. pneumoniae fractions. Each individual point represents an individual biological replicate. c Group clinical scores of mice showing clinical signs such as piloerection, hunching, nose bulge, orbital tightening and lethargy as a function of time after prime and boost injections. Group clinical scores were derived by adding +1 per sign displayed per animal. Data from one individual experiment with n = 12 animals per group. d Histopathological lung lesion scores and e bacterial loads recovered from vaccinated-then-challenged animals. Data for (d), and (e) are pooled from five independent experiments, with each point representing data from an individual animal. f Representative histopathological lung images of sham vaccinated/sham challenged animals (0), sham vaccinated/M. pneumoniae challenged animals (1.5), LAMPs vaccinated/M. pneumoniae challenged animals (3), and dLAMPs vaccinated/M. pneumoniae challenged animals (1.5). Arrows point to perivascular and peribronchiolar leukocytic infiltrates. Error bars indicate mean and standard error of the mean (SEM) for parametric data (b) and median and interquartile ranges (IQR) for non-parametric data (d, e) (*p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001).
To test the effect of lipoprotein delipidation on VED, native and delipidated Mp fractions (LAMPs, dLAMPs, Aq, dAq, Ins, and dIns) were used to intraperitoneally vaccinate mice according to the schedule illustrated in Fig. 2a with 50 μg of protein from each fraction per dose. Shortly after vaccination, mice receiving non-lipase-treated fractions began to display clinical signs consistent with mild septic shock (Supplementary Table 1, Supplementary Fig. 1), which we quantified as a group clinical score (Fig. 2c). While these clinical signs abated by 48-h post-vaccination, they point to potential immediate safety concerns for vaccine candidates containing intact lipoproteins. Notably, early clinical signs were absent in mice vaccinated with delipidated fractions. Given that in vivo data mirrored those obtained in vitro (Fig. 2b), early clinical signs may have resulted from the over-stimulation of peritoneal immune cells by the lipid moieties of native Mp lipoproteins.
In terms of VED, only mice vaccinated with LAMPs (containing native lipoproteins) exhibited histopathological lung lesions that were more severe than sham-vaccinated/challenged animals, consistent with LAMP-mediated VED (Fig. 2d, f). Furthermore, vaccination with LAMPs did not reduce bacterial loads, suggesting that the induced inflammation is solely pathologic and consistent with VED (Fig. 2e). Importantly, vaccination with delipidated LAMPs (dLAMPs) did not exacerbate lung pathology (Fig. 2d, f), demonstrating that the factors responsible for VED are the lipid moieties of Mp lipoproteins. Compared to controls, dLAMP-vaccinated mice also had significantly lower bacterial loads (Fig. 2e). These data suggest that the dLAMP vaccine preparation includes protective antigens that improve bacterial clearance and can reduce Mp-induced lung disease when freed of the VED effects associated with Mp lipoprotein lipid moieties.
Mice vaccinated with MPLA-SM adjuvanted dLAMPs and challenged had significantly reduced lung lesion scores when compared to sham vaccinated, Mp challenged mice (Supplementary Fig. 2A, C), but did not differ in bacterial loads recovered (Supplementary Fig. 2B). This was unsurprising as lesion scores did not always correlate with bacterial loads. These data further indicate that the choice of adjuvant will be critical in developing a vaccine that reduces bacterial loads while also protecting the host from unnecessary pulmonary inflammation.
Conclusions
The identification of Mp lipoprotein lipid moieties as the cause of Mp VED is a breakthrough for the field, overcoming a major roadblock for Mp vaccine development. The findings presented here may be broadly applicable to other Mycoplasma species for which vaccine induced disease exacerbation has been reported. Vaccination of calves with inactivated whole cell vaccines and partially purified membrane proteins of Mycoplasma bovis resulted in increased pulmonary pathology upon virulent challenge21,22. Additionally, vaccination of cattle with inactivated whole cell Mycoplasma mycoides subsp. mycoides SC and a subunit lipoprotein Q (LppQ) vaccine also were reported to exacerbate the effects of Contagious Bovine Pleuropneumonia (CBPP) in challenged animals23. These studies did not identify the lipid moieties of lipoproteins or any other factors as the causative factors of VED in these cases. Given the parallels of these studies with our data, however, it is attractive to speculate that the lipid moieties of the lipoproteins included in these vaccine candidates may have been the maladaptive factors responsible for disease exacerbation. Indeed, vaccination of cattle with the N terminus of LppQ (which includes the region where the lipid moieties attach) failed to be protective and was also associated with adverse events24. It is important, however, to note that more studies need to be conducted to demonstrate the maladaptive effect of lipoproteins in ruminant mycoplasmosis vaccines.
In terms of M. pneumoniae, our data indicate that caution should be used when considering the inclusion of native Mycoplasma lipoproteins in future vaccine formulations against this significant human pathogen. Furthermore, the findings here may be broadly applicable to other bacterial pathogens for which VED is observed, and suggest that the potential role of lipoproteins in this phenomenon should not be easily dismissed.
Methods
TX-114 phase partitioning of Mp proteins
TX-114 partitioning25 of M. pneumoniae proteins was performed as detailed in the supplementary methods (see Extended Methods). Fractions were ethanol precipitated and resuspended in TS buffer (15 mM Tris, 150 mM Nacl, pH 7.6)via sonication (Sonics VCX-500 ultrasonicator, Newtown, CT), then quantified by a Qubit 2.0 Fluorometer (Thermofisher, Waltham, MA) protein assay kit.
Proteomic analysis of Mp LAMPs
Purified LAMPs were prepared using a slightly modified filter-aided sample preparation (FASP) method26 described in the supplementary methods (see Extended Methods). Purified peptides were injected onto a PepMap RSLC C18 column (Thermo Scientific) and separated using reversed phase gradient on a Dionex Ultimate 3000 RSLC UPLC instrument (Thermo Scientific) coupled directly to a Q Exactive HF mass spectrometer (Thermo Scientific) via electrospray ionization. MaxQuant (v1.6.1.0) was used to search raw files against a custom Mp proteome database (CP010538_faa) using the Andromeda search engine27 and for peptide and protein quantification using the LFQ algorithm. Scaffold v4.9 (Proteome Software, Inc.) was used for visualization and analysis, and gene designations were identified through comparison to the Mycoplasma pneumoniae (T00006) KEGG genome database with 95–100% sequence identity and lowest E-value used as cut-offs. Multigene Mp lipoproteins were categorized into the 6 numerically named families based on previously established criteria28. Lipoproteins not belonging to a multigene family were categorized as “No Defined Family”.
Lipoprotein lipase digestion and macrophage inflammatory bioassay
Precipitated Mp TX-114 fractions (Insoluble: Ins, Aqueous: Aq, and Detergent: LAMPs) were treated with Lipoprotein Lipase from Burkholderia sp. (EC 3.1.1.34; Sigma Aldrich, St. Louis, MO) to generate the delipidated fractions (respectively). The efficiency of lipoprotein lipase treatment was assessed via TLR-2 Bioassay/Macrophage Inflammatory Assay29 described in the supplementary methods (see Extended Methods).
Bacterial strains and animal studies
Mycoplasma pneumoniae strain PI1428 was utilized for all aspects of this study. All animal experiments were conducted in accordance with approved Institutional Animal Care and Use Committee protocol (A17–034) as described in the supplementary methods (see Extended Methods).
Statistical analyses
Results were analyzed via a non-parametric one-way ANOVA on ranks (Kruskal–Wallis) with a Dunn’s post hoc test or a parametric ordinary one-way ANOVA with a Tukey’s post hoc test for multiple pairwise comparisons between groups. Analysis of data with only two groups was conducted utilizing a one tailed Mann–Whitney U test for nonparametric data (α = 0.05). All data were analyzed using the GraphPad Prism software, version 8.02 (GraphPad Software, La Jolla, California, USA).
Reporting summary
Further information on experimental design is available in the Nature Research Reporting Summary linked to this article.
Data availabiliy
The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE30 partner repository with the dataset identifier PXD016814. All other data are included in this manuscript and supplementary file.
References
Waites, K. B. et al. Mycoplasma pneumoniae from the respiratory tract and beyond. Clin. Microbiol. Rev. 30, 747–809 (2017).
Bajantri, B., Toolsie, O., Venkatram, S. & Diaz-Fuentes, G. Mycoplasma pneumoniae pneumonia: walking pneumonia can cripple the susceptible. J. Clin. Med. Res. 10, 891–897 (2018).
Marrie, T. J. Mycoplasma pneumoniae pneumonia requiring hospitalization, with emphasis on infection in the elderly. Arch. Intern Med. 153, 488–494 (1993).
Niederman, M. S. & Brito, V. Pneumonia in the older patient. Clin. Chest Med. 28, 751–771 (2007).
Khoury, T. et al. Increased rates of intensive care unit admission in patients with Mycoplasma pneumoniae: a retrospective study. Clin. Microbiol Infect. 22, 11–714 (2016).
Liu, X. et al. Drug resistance mechanisms of Mycoplasma pneumoniae to macrolide antibiotics. Biomed. Res Int. https://doi.org/10.1155/2014/320801 (2014).
Bebear, C., Pereyre, S. & Peuchant, O. Mycoplasma pneumoniae: susceptibility and resistance to antibiotics. Future Microbiol. 6, 423–431 (2011).
Smith, C. B., Friedewald, W. T. & Chanock, R. M. Inactivated Mycoplasma pneumoniae vaccine. Evaluation in volunteers. J. Am. Med. Assoc. 199, 353–358 (1967).
Smith, C. B., Chanock, R. M., Friedewald, W. T. & Alford, R. H. Mycoplasma pneumoniae infections in volunteers. Ann. N. Y. Acad. Sci. 143, 471–483 (1967).
Szczepanek, S. M. et al. Vaccination of BALB/c mice with an avirulent Mycoplasma pneumoniae P30 mutant results in disease exacerbation upon challenge with a virulent strain. Infect. Immun. 80, 1007–1014 (2012).
Kurai, D. et al. Mycoplasma pneumoniae extract induces an IL-17-associated inflammatory reaction in murine lung: implication for mycoplasmal pneumonia. Inflammation 36, 285–93 (2013).
Kurata, S. et al. Role of IL-17A and IL-10 in the antigen induced inflammation model by Mycoplasma pneumoniae. BMC Microbiol. 14, 156 (2014).
Chu, H. W. et al. Repeated respiratory Mycoplasma pneumoniae infections in mice: effect of host genetic background. Microbes Infect. 8, 1764–1772 (2006).
Proft, T. & Herrmann, R. Identification and characterization of hitherto unknown Mycoplasma pneumoniae proteins. Mol. Microbiol. 13, 337–348 (1994).
Regula, J. T. et al. Defining the mycoplasma cytoskeleton: the protein composition of the Triton X-100 insoluble fraction of the bacterium Mycoplasma pneumoniae determined by 2-D gel electrophoresis and mass spectrometry. Microbiology 147, 1045–1057 (2001).
Beghetto, E., De Paolis, F., Montagnani, F., Cellesi, C. & Gargano, N. Antobody response to Mycoplasma pneumoniae: protection of host and influence on outbreaks? Microbes Infect. 11, 66–73 (2009).
Nuyttens, H., Cyncynatus, C., Renaudin, H., Pereyre, S. & Bebear, C. Identification, expression and serological evaluation of the recombinant ATP synthase beta subunit of Mycoplasma pneumoniae. BMC Microbiol. 10, 216 (2010).
Szczepanek, S. M. et al. Identification of lipoprotein MslA as a neoteric virulence factor of Mycoplasma gallisepticum. Infect. Immun. 78, 3475–3483 (2010).
Masukagami, Y. et al. The Mycoplasma gallisepticum virulence factor lipoprotein MslA is a novel polynucleotide binding protein. Infect. Immun. 81, 3220–3226 (2013).
Zuo, L. L., Wu, Y. M. & You, X. X. Mycoplasma lipoproteins and toll-like receptors. J. Zhejiang Univ. Sci. B. 10, 67–76 (2009).
Rosenbusch, R. Test of an inactivated vaccine against Mycoplasma bovis respiratory disease by transthoracic challenge with an abscessing strain. Abstracts of the 12th International Organization of Mycoplasmology Conference, Sydney, Australia, 185 (1998).
Bryson, D., Ball, H. J., Brice, N., Pollock, D. S. & Forster, F. Pathology of induced Mycoplasma bovis pneumonia in experimentally vaccinated calves. Mycoplasmas Ruminants: Pathogenicity, Diagnostics, Epidemiol. Mol. Genet. 3, 128–132 (1998).
Nicholas, R. A. J. et al. An inactivated whole cell vaccine and LppQ subunit vaccine appear to exacerbate the effects of CBPP in adult cattle. Report of the 3rd Meeting of the FAO-OIE-AU/IBAR-IAEA Consultative Group on Contagious Bovine Pleuropneumonia (CBPP). 91–97 (2004).
Mulongo, M. et al. Vaccination of cattle with the LppQ-N’ sub-unit of Mycoplasma mycoides subsp. mycoides results in type III immune complex disease upon experimental infection. Infect. Immun. 83, 1992–2000 (2015).
Bordier, C. Phase separation of integral membrane proteins in Triton X-114 solution. J. Biol. Chem. 25;256, 1604–1607 (1981).
Wisniewski, J. R., Zougman, A., Nagaraj, N. & Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 6, 359–362 (2009).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b- range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–72 (2008).
Hallamaa, K. M., Browning, G. G. & Tang, S. L. Lipoprotein multigene families in Mycoplasma pneumoniae. J. Bacteriol. 188, 5393–5399 (2006).
Bartosh, T. J. & Ylostalo, J. H. Macrophage inflammatory assay. Bio. Protoc. 4, https://doi.org/10.21769/bioprotoc.1180 (2014).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res https://doi.org/10.1093/nar/gky1106 (2019).
Acknowledgements
We gratefully acknowledge the quantitative proteomics analysis conducted by Dr. Jeremy L. Balsbaugh, Director of the Proteomics & Metabolomics Facility, a component of the Center for Open Research Resources and Equipment at the University of Connecticut. We also wish to thank Dr. Jessica Canter and Morgan Hunte for technical assistance with mouse studies. This study was funded in part by funds made available by the Center of Excellence for Vaccine Research (S.J.G.) and through an UConn OVPR START Grant (S.M.S.).
Author information
Authors and Affiliations
Contributions
A.B.M. conceptualized, designed and performed experiments, analyzed data and wrote the manuscript. T.D.G. helped perform experiments and analysis. E.R.T., S.J.G., and S.M.S. conceptualized the project and participated in study design. S.J.G. and S.M.S. supervised the project. All authors contributed equally to manuscript editing.
Corresponding authors
Ethics declarations
Competing interests
E.R.T., S.J.G., and S.M.S are listed inventors on a U.S. Provisional Patent Application filed based on the data presented herein.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mara, A.B., Gavitt, T.D., Tulman, E.R. et al. Lipid moieties of Mycoplasma pneumoniae lipoproteins are the causative factor of vaccine-enhanced disease. npj Vaccines 5, 31 (2020). https://doi.org/10.1038/s41541-020-0181-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41541-020-0181-x
This article is cited by
-
Insight into the Pathogenic Mechanism of Mycoplasma pneumoniae
Current Microbiology (2023)
-
B cells oppose Mycoplasma pneumoniae vaccine enhanced disease and limit bacterial colonization of the lungs
npj Vaccines (2022)
-
Vaccination with Mycoplasma pneumoniae membrane lipoproteins induces IL-17A driven neutrophilia that mediates Vaccine-Enhanced Disease
npj Vaccines (2022)
-
Contagious Bovine and Caprine Pleuropneumonia: a research community’s recommendations for the development of better vaccines
npj Vaccines (2020)