Abstract
Algal cultivations are strongly influenced by light and dark cycles. In this study, genome-scale metabolic models were applied to optimize nutrient supply during alternating light and dark cycles of Chlorella vulgaris. This approach lowered the glucose requirement by 75% and nitrate requirement by 23%, respectively, while maintaining high final biomass densities that were more than 80% of glucose-fed heterotrophic culture. Furthermore, by strictly controlling glucose feeding during the alternating cycles based on model-input, yields of biomass, lutein, and fatty acids per gram of glucose were more than threefold higher with cycling compared to heterotrophic cultivation. Next, the model was incorporated into open-loop and closed-loop control systems and compared with traditional fed-batch systems. Closed-loop systems which incorporated a feed-optimizing algorithm increased biomass yield on glucose more than twofold compared to standard fed-batch cultures for cycling cultures. Finally, the performance was compared to conventional proportional-integral-derivative (PID) controllers. Both simulation and experimental results exhibited superior performance for genome-scale model process control (GMPC) compared to traditional PID systems, reducing the overall measured value and setpoint error by 80% over 8 h. Overall, this approach provides researchers with the capability to enhance nutrient utilization and productivity of cell factories systematically by combining genome-scale models and controllers into an integrated platform with superior performance to conventional fed-batch and PID methodologies.
Similar content being viewed by others
Introduction
Microalgae represent promising microorganisms for transforming renewable resources and inorganic carbon sources into biomass, biofuel precursors, and high-value products1. One of the major challenges in making algal-derived products competitive against existing products in the market is developing sustainable bioprocesses that maximize productivity while also minimizing nutrient costs. Another challenging aspect of algal cultures is the presence of alternating light and dark cycles, which can result in major changes in algal physiology and metabolism2. Under photoautotrophic conditions, algae will grow in the light and lose biomass in the dark due to the release of carbon through respiration3. This can be problematic if the goal is to maximize biomass or cellular components. An alternative to this approach is to employ organic carbon sources, such as glucose, to drive biomass production during dark cycles4. However, most algae bioreactors operate predominantly in fed-batch mode and thus feeding can be challenging during dark cycles due to difficulties in estimating the amount of glucose organics to be added5,6. As a result, traditional fed batch is not an ideal control strategy to achieve efficient nutrient utilization for periodic alternating light and dark cycles due to the presence of unutilized glucose remaining in the bioreactor during the light cycles or the premature depletion of glucose before the end of the dark cycle.
In order to address this issue, we applied a genome-scale model (GSM) predictive control (MPC) strategy to optimize glucose and nitrate feeding for the model alga Chlorella vulgaris. In this system, during autotrophic growth, energy from light was used to capture carbon dioxide (CO2) to support algal growth, whereas glucose was added to supply energy to sustain algal growth during dark cycles. Over the past twenty years, constraint-based modeling has been developed and implemented to gain insights into the metabolism and the metabolic flux within different microorganisms7. GSMs have been constructed for multiple algal species including Chlamydomonas8, Chlorella9, and Nannochloropsis10. Recent studies have attempted to couple GSMs with dynamic flux balance analysis to optimize feeding strategies in order to improve ethanol production in Saccharomyces cerevisiae and Escherichia coli cultures in silico11,12. Another study used a GSM to optimize media supply and maintain C. vulgaris biomass13. The majority of these studies utilized a single model to optimize nutrient feeding, however, cells dynamically change their biomass composition under different culture conditions14. Previously, our group developed a genome-scale metabolic model with dynamic biomass compositions based on experimentally determined biomass measurements and corresponding omics data15. We then applied this dynamic model to predict nutrient requirements under autotrophic, heterotrophic, and nitrogen-limited conditions16.
MPC has become a widely used strategy to optimize biomanufacturing by using a model to predict cellular behavior, which is a highly nonlinear process. To utilize this approach, kinetic models have been applied with a nonlinear MPC algorithm to optimize fixation of CO2 in C. vulgaris cultures17. Another study applied an artificial neural network to build data-driven models and regulate light intensity for a microalgae culture18. Both studies demonstrated the robustness of this MPC strategy but primarily utilized kinetic equations modeling nutrient transport from extracellular environments without detailed representation of the entire metabolism. Indeed, a previous large-scale outdoor culture study indicated that the significant barrier to algal bioproduction was the lack of understanding of microalgal biology for optimal biomass generation19. To improve our knowledge of metabolism, GSMs have been constructed for multiple algal species by our group and others and used as tools to improve strain development, to identify metabolic bottleneck and to understand microbiome interactions, among other applications20,21. In this regard, GSMs have been used to predict metabolic demands under different culture conditions such as photoautrophic and heterotrophic environments, depletion of nitrogen source or for multiple species interaction22. This expansive description of cellular metabolism provides a valuable knowledge base that can also be used as a powerful tool for controlling cellular performance in biomanufacturing processes.
Therefore, in this study we incorporated separate photoautotrophic (iCZPA-T1) and heterotrophic (iCZH-T1) genome-scale metabolic models for light and dark cycles, respectively, in order to optimize metabolic pathways and utilization of nutrient supplies and compared their performance in terms of production of biomass, fatty acids (FAs), and lutein to standard autotrophic and heterotrophic cultivations. The metabolic models were then used to control nutrient supply in both open-loop and closed-loop configurations, which represents a useful approach to efficiently channel diverse nutrients to algal cellular components and enhance production of algal-derived bioproducts during day-night cycles. To enhance control, we adapted the parameters in the closed-loop controller based on feedback measurements of biomass and nutrients to improve model predictions for bioreactor operations. This approach of closed-loop genome model process control (GMPC) demonstrated significant improvements in nutrient utilization efficiencies compared to conventional fed-batch approaches. Finally, we demonstrated superior performance for this GMPC system compared to conventional PID systems, illustrating the value of this methodology for improving biomanufacturing processes.
Results and discussion
Advantages of using genome-scale model predictions on C. vulgaris cultures under light/dark cycles
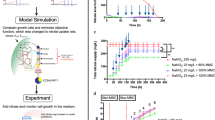
Previously, a collection of genome-scale metabolic models (GSMs) of C. vulgaris has been constructed to reflect different growth phenotypes (iCZ946-PAT1 to PAT6 and iCZ946-HT1 to HT5 in Supplementary Table 1)9,15 and our group has successfully demonstrated the application of iCZ946-PAT1, HT1, and PAT5 models to optimize nutrient supply for autotrophic, heterotrophic and nitrogen limited conditions in 150 mL cultures, respectively16. Here, we expanded on this methodology to apply photoautotrophic (iCZ946-PAT1) and heterotrophic (iCZ946-HT1) models, in sequence to predict nutrient requirements under alternating light and dark cycles, which mimic conditions present in outdoor cultivation systems (Fig. 1a). During these cycles, glucose concentration, nitrate concentration, and biomass as OD750 were measured every 8 or 16 h. The OD750 measurements were converted to biomass concentration, which served as constraints in genome-scale metabolic models to predict nutrient requirements to support a fixed growth rate under alternating autotrophic and heterotrophic conditions. The GSM iCZH-T1 was applied to predict glucose and nitrate requirements during heterotrophic growth, which averaged around 0.023 h−1 under dark conditions and the corresponding glucose and nitrate levels were added to the cultures accordingly based on model predictions. In this way, the alternating day-night cultures received the appropriate glucose and nitrate feeds, based on the model predictions, for growth during the dark periods. Alternatively, for the light cycle, the model was switched to the photoautotrophic model iCZPA-T1 (see Supplementary Table 1) in order to predict nitrate feeding requirements, which averaged around 0.019 h−1 during the 16-hour light conditions (Fig. 1a).
a Flowchart of model predictions over light/dark cycles. b Cell growth. c Glucose uptake in all three conditions. d Glucose feed and glucose concentration in the media under 16 h light (A)/ 8 h dark (H) cycles. e Nitrate uptake in all three conditions. f Nitrate feed and nitrate concentration in the media under 16 h light (A)/8 h dark (H) cycles. g Biomass concentration, fatty acid titer and lutein titer. h Biomass yield, Fatty acid yield and lutein yield on glucose. The data represents the mean ± SD for n = 3 (biological triplicates).
The optical density following glucose and nitrate feeding based on model predictions was then compared to growth under 24-h light autotrophic (24 h light (A)) and under 24-h dark heterotrophic (24 h dark (H)) conditions without any model predictive feeding (Fig. 1b). For heterotrophic conditions, excess glucose (10 g/L) and nitrate (250 mg/L) were added to ensure no limitations during the growth period. While no glucose was present in the autotrophic cultures, the cells still grew by fixing CO2 and reached OD750 of approximately 1.0 with 250 mg/L initial nitrate added (red line in Fig. 1b, c). Alternatively, the optical density for the alternating light–dark cycling (16 h light (A)/8 h dark (H)) was comparable to that obtained by heterotrophic conditions with both reaching OD750 of 1.9 or higher by 120 h (Fig. 1b). Previous studies found that culturing C. vulgaris under heterotrophic conditions can improve its growth under subsequent autotrophic conditions23. Indeed, the activation of Rubisco protein under autotrophic conditions can also support higher growth rates under subsequent heterotrophic conditions24. These studies demonstrated the synergistic benefits from alternating light and dark cycles and helped to explain the comparable growth profiles we observed under alternating light and dark cycles (blue line in Fig. 1b) and completely dark heterotrophic conditions (green line in Fig. 1b). Alternatively, C. vulgaris grew more slowly under pure autotrophic conditions (red line in Fig. 1b).
For the 24 h dark (H) conditions, C. vulgaris consumed approximately 3200 mg/L glucose over 120 h (Fig. 1c). For the alternating light and dark cycle case (16 h light (A)/8 h dark (H)), an initial feed of 100 mg/L glucose was added to facilitate growth during the first dark cycle. Glucose levels in the media were measured before we supplemented glucose. C. vulgaris consumed most of the initial glucose, reducing the concentration to 16 mg/L within the first 8 h (Fig. 1d). At this point, conditions were cycled back to light and no additional glucose was added until 24 h. At 24 h, a bolus of 56 mg/L glucose was added to support heterotrophic growth based on model-predicted requirements for the next 8 h. After 32 h, the added glucose was indeed nearly consumed by the end of the second dark cycle. Glucose supplemented at the start of each dark cycle was then consumed completely over the dark cycles at 48, 72, and 96 h. Overall, the glucose consumption was reduced from 3200 mg/L for the heterotrophic cultures to 800 mg/L for the light/dark cycles within 120 h, a reduction of nearly 75% (Fig. 1c).
Nitrate consumption was also compared in the three different cultures (Fig. 1e). C. vulgaris consumed 120 mg/L nitrate and 300 mg/L in the autotrophic and heterotrophic cultures, respectively, over 120 h. Alternatively, for the light and dark cycling case, the basal 25 mg/L nitrate was exhausted in the media by 48 h (Fig. 1f). Subsequently, the GSMs iCZPA-T1 and iCZH-T1 were applied to estimate the nitrate required at the beginning of the light and dark periods for each cycle. This strategy resulted in no significant excess of nitrate remaining in the media at the end of both periods (Fig. 1f). As a result, around 230 mg/L nitrate was consumed by 120 h in alternating light/dark cycles, which was 23% less than the nitrate utilized in the dark heterotrophic case (Fig. 1e).
Two major microalgae-derived products of biotechnological interest, lutein, and FA, were analyzed for three different cultures in terms of total titer (mg/L in Fig. 1g) and yield per glucose (mg/g glucose in Fig. 1h). At the completion of the experiment (120 h), without light stimulation, the lutein titer was the lowest under 24 h dark (H) conditions (Fig. 1g), however, the FA and biomass concentrations were the highest at approximately 36 and 570 mg/L. The slower growth rate at the 24-h light (A) conditions resulted in the lowest FA titer and biomass concentrations. For the 16 h light (A)/8 h dark (H) conditions, FA and biomass concentrations were intermediate between the values we obtained in autotrophic and heterotrophic conditions. Interestingly, given the combinatorial effect of high biomass accumulation from heterotrophic conditions and high lutein content from photoautotrophic light cycles, the highest lutein titer of 2.2 mg/L was achieved under alternating light and dark cycles (Fig. 1g). Previous studies indeed found that light is an important factor to stimulate lutein production in microalgae25. Overall lutein, FA and biomass yields per gram of consumed glucose increased significantly between three and eightfold under alternating light and dark cycles as compared to heterotrophic conditions (Fig. 1h). During the light and dark cycling, C. vulgaris only needed the glucose supply for 8 h of the dark period every 24 h, which led to more efficient glucose utilization.
In this way, our results showed that total biomass, lutein, and FA titers were all higher for light–dark cycling compared with autotrophic cultures (Fig. 1g) and product yields per gram of glucose were all higher for light cycling than heterotrophic cultures (Fig. 1h). Previous omic26 and fermentation studies27 typically compared mixotrophic with autotrophic or heterotrophic conditions because they did not have the capability to control nutrient supply accurately under alternating light and dark cycles. However, both metabolic pathways are important for C. vulgaris. Autotrophic metabolism provides algae with the capability to fix carbon dioxide and synthesize high amount of carotenoids such as lutein in order to protect the free radicals generated during photosynthesis. Heterotrophic metabolism provides algae with the capacity to utilize organic carbon for efficient biomass and FA generation. By taking advantages of both metabolic capacities, our nutrient control approach clearly exemplifies the benefits of genome-scale metabolic models to optimize nutrient supply for maximizing biomass and product yields under light and dark cycles.
Comparing the open-loop genome-scale model process control with fed-batch control on C. vulgaris cultures
After demonstrating the capability of using these genome-scale metabolic models to control nutrient supply against autotrophic and heterotrophic systems, we next compared the performance of cultures using genome-scale model process control (GMPC) with traditional fed-batch cultures for alternating photoautotrophic light and heterotrophic dark cycles. The experiment was performed in a 2 L bioreactor system with computer-controlled glucose and nitrate supply using the MatlabTM software (MathWorks) to run GSMs and control two peristaltic pumps. The standard (non-model controlled) fed-batch bioreactor was initially supplied with 500 mg/L glucose with additional glucose supplied to the bioreactor periodically in 500 mg/L bolus increments as the glucose was exhausted (Fig. 2a). Alternatively, an open-loop model control system relied on Matlab-based GSM predictions to control the nutrients required for growth without any inputs of measurements from the system. This algorithm runs by setting the initial biomass concentration (X0), initial nitrate concentration (N0), and glucose concentration (G0), which are three controlled variables in the system, along with fixed photoautotrophic and heterotrophic growth rates, which were based on previous test runs. The computer then runs the GSM to predict the nitrate and glucose requirements, which dictated manipulated variables (FG, FN) to be supplied automatically every 8 and 16 h for a period of 4 days.
a Flowchart of standard fed-batch and open-loop GMPC cultures. b Cell growth. c Growth rate comparison between GMPC (Experiment) and GMPC (Prediction). d Glucose supply during the cultures. e Glucose level in the media. f Biomass yield on glucose. The data represents the mean ± SD for n = 3 (technical triplicates). *P ≤ 0.05 (GraphPad unpaired t test).
C. vulgaris grew similarly in both standard fed-batch culture and GMPC culture with the biomass concentration reaching around 500 mg/L at 96 h (Fig. 2b). However, a significant deviation was observed between the GMPC prediction (green) and experimental results (blue) after 80 hours of culture. Measured growth of C. vulgaris was significantly lower than model simulation, indicating an inconsistency between model predictions and experimental results during this time period. A growth rate comparison between GMPC prediction and experimental results for individual growth periods, including heterotrophic and autotrophic cycles, is shown in Fig. 2c. The experimental growth rate of autotrophic cultures (blue bars in Fig. 2c) gradually declined from 0.038 h−1 at 32–48 h to 0.030 h−1 at 56–72 h and eventually to 0.014 h−1 at 80–96 h, which was 60% lower than the model prediction of approximately 0.035 h−1 (green bars in Fig. 2c). Previous studies have observed that progressive increase in biomass will block light penetration and thus alter algal growth28, which may explain the gradual decline in the algal growth rate and resulting deviations away from model predictions, as this effect was not considered in our open-loop model predictive control. Furthermore, the heterotrophic growth rate was around 0.04 h−1 at 48–56 h and 0.032 h−1 at 72–80 h, which, in contrast, was higher than the model prediction of around 0.023 h−1 at these time points (Fig. 2c). Previous investigations have indicated that CO2 fixation is still active up to 18 h after transferring from light-facilitated photoautotrophic to heterotrophic conditions24, which may explain the higher growth rate observed in our measurements compared to the predictions under purely heterotrophic conditions.
For the GMPC case, 36 mg/L glucose was supplied initially based on model predictions of the amount to be consumed over the 8 h heterotrophic growth followed by additional supplements at 24, 48, and 72 h (Fig. 2d). However, glucose was not consumed completely in GMPC cultures at 32, 56, and 80 h with 40, 70, and 100 mg/L glucose, respectively, was left in the media, indicating limitations for these open-loop model predictions (Fig. 2e). In the standard fed-batch cultures, a bolus of 500 mg/L glucose was fed to the standard fed-batch culture at 0 h and completely utilized by C. vulgaris at around 72 h, followed by the addition of another 500 mg/L glucose supplement at that time (Fig. 2d). As a result, C. vulgaris was likely consuming some glucose and CO2 simultaneously during the light cycle, resulting in mixotrophic conditions for both the GMPC case and standard fed-batch cultures. Indeed, we observed declines in the glucose levels of C. vulgaris during the light cycles in both cases (white sections in Fig. 2e). This may explain the similar cell growth curves of C. vulgaris between standard fed-batch cultures and GMPC cultures (Fig. 2b). However, even though C. vulgaris consumed glucose in both conditions, the biomass yield per gram of glucose was still 12% higher for the GMPC group (0.63 gBiomass/gglucose) compared to the fed-batch group (0.57 gBiomass/gglucose) (p < 0.05) (Fig. 2f).
The extra glucose measured (Fig. 2e) and the variation in growth predictions (Fig. 2b) suggest that there are deficiencies in the model’s capacity to predict growth and glucose consumption in this open-loop control system. In addition, the performance of open-loop GMPC in terms of biomass yield on glucose was only moderately better than fed-batch cultures, likely in part because the glucose supply was not controlled appropriately under dark cycles for the GMPC conditions. Consequently, a closed-loop feedback control loop, which was adapted to compensate for limitations in the model’s ability to describe growth at higher algal densities and mixotrophic conditions was implemented next as an improved GMPC framework.
Closed-loop GMPC achieved better performance than standard fed-batch control
To further improve GMPC, we incorporated measured data to build a closed-loop model prediction algorithm that incorporates biomass (Xm), nitrate (Nm), and glucose (Gm) levels measured every 8 h (Fig. 3a). The measurements were then used as inputs into the model (green box in Fig. 3a). Unlike the open-loop system in Fig. 2, we considered CO2 fixation by active Rubisco protein in heterotrophic cycles, which was previously observed for algal two-stage cultivations (photoautotrophic phase-heterotrophic phase)24. For the heterotrophic cycles, the calculated growth rate (μC), calculated glucose demand (FG,C), and calculated nitrate demand on a per L basis (FN,C) during the 8-h period were determined based on measured inputs (Xm, Gm, Nm). Next, the model and algorithm optimizer (green and blue boxes in Fig. 3b) then estimate the fractions of photoautotrophic growth and heterotrophic growth in order to determine the feed under heterotrophic (dark) conditions based on the algorithm shown in Fig. 3b. The algal cells were assumed to operate under two different types of metabolism in the simulations for the dark cycle. One fraction of algal cells was assumed to grow strictly heterotrophically, as represented by model iCZH-T1. In addition, a certain fraction (a) of algal biomass was assumed to grow mixotrophically and thus fixes CO2 during the dark cycle as suggested in previous publications24. The current photoautotrophic genome-scale model iCZPA-T1 was not constructed to function when the light intensity (PRISM_solar_litho) is set as zero in the model. In this simulation, we therefore set the light intensity in the iCZPA-T1 model to a minimum for the current simulations in dark periods of the cycle. As a result, three equations were added to consider this combined metabolic operation and its impact on growth rate, glucose consumption rate, and nitrate consumption rate. The algorithm optimizes six variables including autotrophic growth rate (μA), autotrophic nitrate demand (FNA), autotrophic biomass percentage (a), heterotrophic growth rate (μH), heterotrophic glucose demand (FHG), and heterotrophic nitrate demand (FNG) to minimize the difference between model simulations and experimental growth rate as well as glucose and nitrate demand for the most recent 8-h dark cycle. Those optimized parameters were then used to dictate overall glucose demand (FG) and nitrate demand on a per L basis (FN) in the next 8 h, which were then controlled by the pumps. Based on the predictions, the control pump supplies glucose and nitrate to the bioreactor.
Alternatively, in the photoautotrophic phase, a differential equation for cell mass accumulation with respect to time was incorporated, which includes a term to describe the logarithmic decay of cell growth rate that occurs at increasing biomass concentrations due to light shading (Fig. 3c)28. This equation was built based on our experimental biomass measurements from a separate autotrophic culture run. The calculated growth rate was then used in iCZPA-T1 to predict and optimize nitrate supply during the light cycles.
The GMPC culture was then compared with a standard fed-batch culture similar to the conditions used in the open-loop experiment (Fig. 4a). Unlike the open-loop system, algal growth for the GMPC culture (blue line in Fig. 4a) was slower than the standard fed-batch culture (red line in Fig. 4a) after 56 h and eventually reached 360 mg/L for GMPC conditions versus 610 mg/L for fed-batch cultures at 96 h due to the greater stringency of glucose control during dark periods. Furthermore, the growth prediction for the closed-loop system (green line) was more closely aligned to the experimental growth rate for the GMPC cultures after 80 h (Fig. 4a) compared with the growth prediction (green line in Fig. 2b) and experimental findings for the open-loop system (blue line in Fig. 2b). Previous studies indicated the success of model predictive control is contingent on a robust process model and on-line measurements29,30. Indeed, in our closed-loop system, the model predictive algorithm was modified based on experimental measurements of cell density, glucose, and nitrate for both autotrophic and heterotrophic conditions in order to predict nutrient requirements for every cycle for the closed-loop system.
The growth rates between simulation and experimental results were compared for individual time periods of the cycling photoautotrophic and heterotrophic cultures (Fig. 4b). Both the model predictions and experimental growth rates changed dynamically over different heterotrophic and autotrophic cycles. The model predictions (green bars in Fig. 4b) for the growth rate during the dark cycles varied between 0.014 h−1 (24–32 h) and 0.031 hour−1 (48–56 h) over the course of the 96 hours experiment. For the experiment, the growth rates (blue bars in Fig. 4b) also changed across the different time periods with a max around 0.032 h−1 and a min around 0.009 h−1. Meanwhile, the model predictions for growth during the light cycles gradually declined from 0.030 h−1 at 8–24 h and eventually falling to 0.019 h−1 at 80–96 h. The experimental growth rates followed the same trend, decreasing from 0.037 h−1 at 32–48 h to 0.015 h−1 at 80–96 h, which again reflects the light shading effect on algal cultures.
Importantly, a total of 910 mg/L glucose was fed to the standard fed-batch as compared to 300 mg/L glucose for GMPC cultures by 96 h (Fig. 4c), a 3-fold reduction in the total glucose consumed. Furthermore, most of the glucose fed to GMPC cultures was consumed almost completely during the dark cycles, with a minimum of 6 mg/L remaining at 78 h and a maximum of 40 mg/L left at 32 h in GMPC cultures (Fig. 4d). For the fed-batch cultures, 440 mg/L glucose was consumed completely at 72 h, leading to the need for a second glucose feed around 500 mg/L. Due to the efficient glucose utilization occurring during the dark cycles of this closed-loop control system, the biomass yield on glucose increased dramatically by 2.2-fold (122%) from 0.67 g/g in standard fed-batch cultures to 1.49 g/g in the closed-loop GMPC system (Fig. 4e). In contrast, the open-loop GMPC system only had a modest 10.6% increase in biomass yield on glucose for the open-loop system (Fig. 2f).
Overall, the closed-loop GMPC demonstrated more accurate controller performance than the open-loop GMPC system. This closed-loop GMPC system relies on glucose and nitrate off-line measurement and those two measurements can take around 30 min. To address this technical challenge, other more rapid nutrient and metabolite measurement tools could be integrated such as in situ Raman spectrometry for metabolite measurements31,32. Alternatively, off gas analysis can be used to characterize cell metabolism toward biomass accumulation or lipid synthesis33 for future versions of GSM control.
GMPC is highly robust in comparison to a PID control during dark heterotrophic periods
After demonstrating the advantages of closed-loop model prediction and its associated higher efficiency of biomass productivity with respect to glucose fed, the model predictive controller was compared to a standard PID controller in silico and experimentally. Using SimulinkTM, a kinetic model consisting of four ODE equations was incorporated in order to describe changes in biomass, nutrient levels, and medium volume during the heterotrophic dark periods in a bioreactor (Supplementary Fig. 2). Genome-scale metabolic models were then used to determine the relationship between growth rate, glucose uptake rate, and nitrate uptake rate as described previously16. Next a PID controller and an GMPC controller were used to control glucose and nitrate levels separately in the bioreactor (Fig. 5a and Supplementary Fig. 2). In order to simulate the performance of the PID and GMPC controllers, a setpoint change was introduced to glucose and nitrate control around 20 and 35 h, respectively (Fig. 5b–d). The glucose setpoint was decreased from 40 to 20 mg/L (blue lines in Fig. 5b, c) and the nitrate setpoint was decreased from 20 to 10 mg/L (blue lines in Fig. 5d, e). Both PID and GMPC controllers were simulated to control glucose supply and nitrate supply every hour. The simulated glucose and nitrate levels exhibited damped oscillations when using a PID controller, a common response for this controller type (yellow lines in Fig. 5b, d). In contrast, employing the GMPC controller eliminated the damping issues and enabled the glucose and nitrate level to more rapidly reach values near the target levels (yellow lines in Fig. 5c, e). In the PID controller, the damping amplitude of glucose level was around 0–20 mg/L (Fig. 5b). In comparison, in the GMPC controller, the error in the glucose levels was around 0–5 mg/L (Fig. 5c). In the same vein, the nitrate level also exhibited damping oscillations around 0–20 mg/L after the setpoint changed to 10 mg/L at 35 h in the PID control system (Fig. 5d) while the error in the GMPC controller was less than 1 mg/L after 44 h (Fig. 5e). The total error between the setpoint and the value was around 50% and 80% lower for the glucose and nitrate levels in the GMPC simulations compared to PID controller systems following the setpoint change. Meanwhile, the glucose supply and nitrate supply increased gradually in the GMPC controller (red lines in Fig. 5c, e), while the nutrient supplied oscillated widely in the PID controller because the PID controller could not predict nutrient requirements. Instead, the amount added (red lines in Fig. 5b, d) was based on the error and the three different gains in the controller.
Step changes in the set point of the glucose were also tested experimentally in 2 L bioreactors to compare the effectiveness of both PID control and closed-loop GMPC on glucose concentration under heterotrophic dark conditions as a test case. The initial biomass was around 400 mg/L and nitrate was held at substantial levels (data not shown), such that nitrate concentration was not a limiting factor in this experiment. The glucose setpoint was changed from 90 to 60 mg/L at 4 h (Fig. 6). The PID controller gains were tuned on MatlabTM to achieve optimal performance with proportional gain (Kp), integral gain (Ki) and derivative gain (Kd) equal to 1.91, 1.27, and 0.10, respectively (Fig. 6a). The glucose levels were measured every hour and the data was fed to both the PID controller and the closed-loop GMPC controller with a pure heterotrophic model since light and dark cycles were not presented. The feedback signal could compensate for the modeling errors and also help to reject the disturbance in the GMPC controller. For the PID controller, the glucose levels exhibited damping oscillations around 90 mg/L from 0 to 4 h with amplitude of the oscillations around 30–60 mg/L, which was 33–66% of the setpoint level (Fig. 6a). After the setpoint changed to 60 mg/L at 4 h, the glucose level was still oscillating around the setpoint. However, for the closed-loop GMPC controller, the glucose level was controlled around 90 mg/L from 0 to 4 h with the largest error around 10 mg/L at 4 h (11% of setpoint in Fig. 6b). After the setpoint change, the glucose level gradually decreased and was stably controlled. The overall error between glucose measured value and setpoint between 0 and 8 h was 203 mg/L for PID and 43 mg/L for closed-loop GMPC, a decrease of nearly 80% for the model based controller approach.
Overall, both SimulinkTM simulations and experimental results demonstrated that the GMPC approach provided more robust and precise control than traditional PID controllers. While the model could anticipate the future behavior of the fermentation and take appropriate control action, the PID controller did not have this capability resulting in oscillations and overshoot behavior in both simulations and experiments. Thus, our study demonstrates how GMPC systems can serve as a bridge between genome-scale metabolic modeling and control algorithms. Since the cultivation conditions can change and affect algal cellular metabolism, our system connected feedback measurements with genome-scale metabolic models and achieved more efficient nutrient utilization and higher product yields for dynamic algal cultivation conditions. In this way, genome-scale metabolic models can be effectively utilized to improve biomanufacturing of microalgae and other industrially important microbial cell factories.
Conclusions
Fed-batch cultivation and PID controllers have been widely used in bioprocess development. Unfortunately, fed-batch cultivation often results in poor nutrient control and wasted nutrients and conventional PID control can lead to oscillating cell behaviors and poor performance under dynamic conditions. In this study, we have utilized the power of genome-scale metabolic models to predict and control glucose and nitrate supply for C. vulgaris cultures under light and dark cycles and compared this approach to conventional autotrophic and heterotrophic processes. Our results first showed that utilizing genome scale models to track and limit glucose and nitrate feeding led to higher titers of biomass, FAs, and lutein than autotrophic conditions and more efficient glucose utilization and higher product yields than heterotrophic conditions. Next, implementing these models into an open loop system modestly improved performance. However, implementing a complete closed-loop system that incorporates glucose and nitrate feed optimizer for dark cycles and accounts for shading during light cycles increased biomass yield 122% more than normal fed-batch cultures. Finally, both computational simulations and experimental results demonstrated that this genome-based MPC system exhibits superior controller performance compared to conventional PID methods.
Methods
Algal strain and cultivation conditions
Green microalgae C. vulgaris UTEX 395 was obtained from the Culture Collection of Algae at the University of Texas at Austin and maintained on sterile agar plates (1.5% w-v) containing Bold’s Basal Medium (BBM). Liquid cultures were inoculated with a single colony in 12.5 mL of sterile BBM. Cells were transferred to 150 mL cultures or 2 L bioreactors at 25 °C using BBM. The growth of the cultures was monitored by measuring optical density (OD) at 750 nm. All the 150 mL cultures were done in biological triplicates at a flow rate of 13 mL/min and 2 L bioreactor cultures were done in technical triplicates at a flow rate of 400 mL/min in this study. Autotrophic cultures were grown with 5% CO2 under 30 μE m−2 s−1 fluorescence illumination for 125 mL cultures and 100 μE m−2 s−1 fluorescence illumination for 2 L cultures Heterotrophic cultures were grown with ambient air (0.04% CO2) under complete dark conditions. For alternating light and dark cycles, autotrophic conditions were used for light sections and heterotrophic conditions were used for dark sections.
Measurement of biomass, glucose, nitrate, FA and lutein
Liquid cultures were harvested using a high-speed centrifuge (Beckman J2–21, Baltimore, USA) at 4000 × g for 10 min. The pellets were stored at −80 °C and lyophilized for 24 h at −40 °C under freeze-dried machine. The lyophilized algal dry biomass was weighted gravimetrically using an analytical balance. Cell culture supernatant samples were stored in 4 °C fridge for glucose and nitrate analysis. The glucose concentration was measured using YSI 2700 biochemistry analyzer (Yellow Springs, OH). The nitrate concentration was measured using nitrite/nitrate Colorimetric Assay Kit (Cayman 780001).
FAME production followed the procedure provided by Dong et al.34. The dry biomass samples (6–7 mg) were mixed with 0.1 mL heptadecanoic acid (5 mg/mL in methanol) (internal standard), 1 mL of chloroform/ methanol and 1.5 mL of HCl/ MeOH (5% v/v). The mixture was heated at 85 °C for 1 h. Then, the mixture was mixed with 1 mL hexane and 5 mL saturated NaCl. The solution was centrifuged at 6000 rpm for 10 min to collect hexane phase. Then, FAMEs were analyzed using an Agilent’s gas chromatography (GC) system with discharge ionization detection equipped with a capillary column (Stabilwax-DA, 30 m 0.25 mm ID, film thickness 0.25 mm). GC inlet was set at 250 °C and the injections were in a volume of 1 μL. The temperature program started at 50 °C and then increased to 170 °C at a rate of 20 °C min−1, with a plateau for 1 min. After this plateau, the temperature increased from 170 to 220 °C at a rate of 4 °C min−1 and then kept constant for 14 min. The total analysis time was 35 min. Helium was used as carrier gas.
Lutein extraction followed the procedure provided by Yuan et al.35. The dried algae pellets (5–10 mg) were homogenized using a mortar and pestle with 4 mL extraction solvent, the mixture of dichloromethane (25%) and methanol (75%), for 2 min and 2 times. The extraction solution was centrifuged at 10,000×g for 10 min and kept in dark in −20 °C. The solution was filtered before HPLC analysis. The mobile phases are eluent A (dichloromethane: methanol: acetonitrile: water, 5.0:85.0:5.5:4.5, v/v) and eluent B (dichloromethane: methanol: acetonitrile: water, 25.0:28.0:42.5:4.5, v/v).
Genome-scale metabolic model prediction and bioreactor setup
For small-scale 150 mL cultures, biomass concentrations were input into the model to predict nutrient supplies and added to cell cultures manually. For 2 L bioreactor cultures, open-loop and closed-loop algorithms were written in MatlabTM language.
The iCZ946 model, including six different biomass compositions for autotrophic conditions (PAT1-PAT6) and five different biomass compositions for heterotrophic conditions (HT1-HT5), was obtained from Zuniga et al.15 (Supplementary Table 1). GSM simulations were performed using the Gurobi Optimizer Version 5.6.3 (Gurobi Optimization Inc., Houston, Texas) solver in MatlabTM (The MathWorks Inc., Natick, MA) with the COBRA Toolbox.
The experimental setup is shown in Supplementary Fig. 1. Biomass concentration (X), glucose level (G), and nitrate level (N) were chosen as controlled variables in the culture and those data were collected every 8–16 h manually. The manipulated variables were glucose demand (FG) and nitrate demand on a per L basis (FN) for 8-h period. Two pumps were used to control both variables automatically by MatlabTM through Arduino chip. All the control algorithms were run on MatlabTM and the codes are provided in Supplementary information.
The SimulinkTM simulation is shown in Fig. 5a and Supplementary Fig. 2. The blue box in Fig. 5a describes the bioreactor behavior. Four equations were built inside the blue box as shown in Supplementary Fig. 2. The inputs were FG and FN. The outputs were biomass, nitrate level, glucose level and volume. Only nitrate levels and glucose levels were fed into the PID and GMPC controller. For the proportional-integral-derivative (PID) controller, the proportional gain (Kp), integral gain (Ki) and derivative gain (Kd) equal to 1.91, 1.27, and 0.1, respectively. The PID controller and GMPC controller were used to control glucose supply and nitrate supply every hour in both simulation and experiment. Changes in the setpoint for glucose were introduced to see how both PID and GMPC responded to those changes.
Summary of equations for 2L bioreactor cultures
Open-loop system
Initial biomass levels (x0), glucose levels (G0) and nitrate levels (N0) were measured as described above and used as inputs into the open-loop system.
Three equations shown below were used to predict biomass growth, nitrate consumption rate, and glucose consumption rate in the open-loop system. The growth rates under light and dark cycles were determined based on previous experimental data. After that, the growth rates were constrained in the autotrophic and heterotrophic GSMs, respectively to determine nutrient exchange rates (rN and rG) under light and dark cycles. The methods for using growth rate to estimate nutrient exchange rates have been described previously in Chen et al.36. We assumed a rapid switch to a new operational steady state following the transition between light and dark cycles.
Based on the results, two pumps will pump glucose and nitrate every 8 h during the day and night cycles. The bolus cellular nitrate demand (FN) and the cellular glucose demand (FG) in mg/liter fed were determined based on the following two equations.
The peristaltic pumps will pump nitrate and glucose into the bioreactor (Tpump,N, Tpump,G) based on nitrate and glucose demand on a per liter basis (FN, FG (mg/L)), feed stock concentration (CN, CG (mg/mL)), volume of the bioreactor (V (L)) and pump volumetric flow rate speed (Q (mL/s)).
Closed-loop system
Initial biomass levels (x0), glucose levels (G0) and nitrate levels (N0) were measured and used as inputs into the closed-loop system.
During the experiment, biomass levels (xm), glucose levels (Gm) and nitrate levels (Nm) were msured and used as inputs into the closed-loop system.
Light cycles
For the light cycle, two equations were built to describe and predict biomass accumulation rate and nitrate consumption rate. Unlike the open loop system, the light shielding effect was considered and the growth rate would decrease as the biomass concentration increased as described in the equation below and shown in Fig. 3c. The GSM was used to predict nutrient exchange rate (rN) based on the measured growth rate. Nitrate demand on a per liter basis (FN (mg/L)) was fed to the bioreactor every 8 h during the light cycle.
Dark cycles
For the dark cycles, three model equations were built to predict biomass accumulation rate, nitrate consumption rate and glucose consumption rate as listed below and shown in Fig. 3b. In the biomass equation, we assumed a fraction of heterotrophic biomass, a, was derived from autotrophic metabolism and the simulated growth rate was μA. Meanwhile, some biomass was derived through heterotrophic metabolism with the simulated growth rate, μH. The nutrient exchange rates (rNA, rNH, rGH) were determined by inputting simulated growth rates into the autotrophic and heterotrophic GSMs respectively.
where μA is simulation growth rate from autotrophic metabolism, μH is the growth rate from heterotrophic metabolism, rNA is nitrate exchange rate from autotrophic metabolism, rNH is the nitrate exchange rate from heterotrophic metabolism, rGH is the glucose exchange rate from heterotrophic metabolism.
The bolus glucose demand and nitrate demand on a per liter basis (FN, model, FG,model (mg/L)) can be determined based on the following equations:
The calculated growth rate (μc) and calculated nutrient demand (FNC and FGC) were determined from feedback measurements (Xm, Gm, Nm) at t and t − 8 using the following equations:
Next, we applied a fitting objective function (J) to minimize the difference between calculated values and simulated (model) values in order to estimate the optimal parameter values (a, μA, μH, rNA, rNH, rGH) for dictating the actual nitrate and glucose feeds to the bioreactor.
The actual bolus nitrate demand (FN) and the glucose demand (FG) were thus determined by using values obtained from this fitting objective function. Based on these estimations, two separate pumps would pump glucose and nitrate every 8 h at the beginning of the dark cycle period.
The peristaltic pumps will pump nitrate and glucose into the bioreactor (Tpump,N, Tpump,G) based on nitrate and glucose demand on a per liter basis (FN, FG (mg/L)), feed concentration (CN, CG (mg/mL)), volume of the bioreactor (V (L)) and pump volumetric flow rate speed (Q (mL/s)).
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Code availability
The codes that support the findings of this study are provided in supplementary information.
References
Rosenberg, J. N., Oyler, G. A., Wilkinson, L. & Betenbaugh, M. J. A green light for engineered algae: redirecting metabolism to fuel a biotechnology revolution. Curr. Opin. Biotechnol. 19, 430–436 (2008).
Shene, C., Asenjo, J. A. & Chisti, Y. Metabolic modelling and simulation of the light and dark metabolism of Chlamydomonas reinhardtii. Plant J. 96, 1076–1088 (2018).
Kato, Y. et al. Light/dark cycling causes delayed lipid accumulation and increased photoperiod-based biomass yield by altering metabolic flux in oleaginous Chlamydomonas sp. Biotechnol. Biofuels 12, 39 (2019).
Cheirsilp, B. & Torpee, S. Enhanced growth and lipid production of microalgae under mixotrophic culture condition: effect of light intensity, glucose concentration and fed-batch cultivation. Bioresour. Technol. 110, 510–516 (2012).
Zheng, Y. et al. High-density fed-batch culture of a thermotolerant microalga Chlorella sorokiniana for biofuel production. Appl. Energy 108, 281–287 (2013).
Shi, X. M., Jiang, Y. & Chen, F. High-yield production of lutein by the green microalga Chlorella protothecoides in heterotrophic fed-batch culture. Biotechnol. Prog. 18, 723–727 (2002).
Bordbar, A., Monk, J. M., King, Z. A. & Palsson, B. O. Constraint-based models predict metabolic and associated cellular functions. Nat. Rev. Genet. 15, 107–120 (2014).
Chang, R. L. et al. Metabolic network reconstruction of Chlamydomonas offers insight into light‐driven algal metabolism. Mol. Syst. Biol. 7, 1–13 (2011).
Zuñiga, C. et al. Genome-scale metabolic model for the green alga Chlorella vulgaris utex 395 accurately predicts phenotypes under autotrophic, heterotrophic, and mixotrophic growth conditions. Plant Physiol. https://doi.org/10.1104/pp.16.00593 (2016).
Loira, N. et al. Reconstruction of the microalga Nannochloropsis salina genome-scale metabolic model with applications to lipid production. BMC Syst. Biol. 11, 66 (2017).
Chang, L., Liu, X. & Henson, M. A. Nonlinear model predictive control of fed-batch fermentations using dynamic flux balance models. J. Process Control 42, 137–149 (2016).
Jabarivelisdeh, B., Findeisen, R. & Waldherr, S. Model predictive control of a fed-batch bioreactor based on dynamic metabolic-genetic network models. IFAC-PapersOnLine 51, 34–37 (2018).
Juneja, A. & Murthy, G. S. Model predictive control coupled with economic and environmental constraints for optimum algal production. Bioresour. Technol. 250, 556–563 (2018).
Ogbonna, J. C. & Tanaka, H. Night biomass loss and changes in biochemical composition of cells during light/dark cyclic culture of Chlorella pyrenoidosa. J. Ferment. Bioeng. 82, 558–564 (1996).
Zuniga, C. et al. Predicting dynamic metabolic demands in the photosynthetic eukaryote Chlorella vulgaris. Plant Physiol. https://doi.org/10.1104/pp.17.00605 (2017).
Li, C.-T. et al. Utilizing genome-scale models to optimize nutrient supply for sustained algal growth and lipid productivity. NPJ Syst. Biol. Appl. 5, 1–11 (2019).
Tebbani, S., Lopes, F., Filali, R., Dumur, D. & Pareau, D. Nonlinear predictive control for maximization of CO2 bio-fixation by microalgae in a photobioreactor. Bioprocess Biosyst. Eng. 37, 83–97 (2014).
Hu, D. W., Liu, H., Yang, C. L. & Hu, E. Z. The design and optimization for light-algae bioreactor controller based on Artificial Neural Network-Model Predictive Control. Acta Astronaut 63, 1067–1075 (2008).
White, R. L. & Ryan, R. A. Long-term cultivation of algae in open-raceway ponds: lessons from the field. Ind. Biotechnol. 11, 213–220 (2015).
Gu, C., Kim, G. B., Kim, W. J., Kim, H. U. & Lee, S. Y. Current status and applications of genome-scale metabolic models. Genome Biol. 20, 1–18 (2019).
O’Brien, E. J., Monk, J. M. & Palsson, B. O. Using genome-scale models to predict biological capabilities. Cell 161, 971–987 (2015).
Colarusso, A. V., Goodchild-Michelman, I., Rayle, M. & Zomorrodi, A. R. Computational modeling of metabolism in microbial communities on a genome-scale. Curr. Opin. Syst. Biol. 26, 46–57 (2021).
Han, F. et al. Enhancement of microalgal biomass and lipid productivities by a model of photoautotrophic culture with heterotrophic cells as seed. Bioresour. Technol. 118, 431 (2012).
Xiong, W., Gao, C., Yan, D., Wu, C. & Wu, Q. Double CO2 fixation in photosynthesis–fermentation model enhances algal lipid synthesis for biodiesel production. Bioresour. Technol. 101, 2287–2293 (2010).
Xiao, Y. et al. Photosynthetic accumulation of lutein in Auxenochlorella protothecoides after heterotrophic growth. Mar. Drugs 16, 283 (2018).
Vidotti, A. D. et al. Analysis of autotrophic, mixotrophic and heterotrophic phenotypes in the microalgae Chlorella vulgaris using time-resolved proteomics and transcriptomics approaches. Algal Res. 51, 102060 (2020).
Park, J.-E., Zhang, S., Han, T. H. & Hwang, S.-J. The contribution ratio of autotrophic and heterotrophic metabolism during a mixotrophic culture of Chlorella sorokiniana. Int. J. Env. Res. Public Health 18, 1353 (2021).
Huesemann, M. et al. A validated model to predict microalgae growth in outdoor pond cultures subjected to fluctuating light intensities and water temperatures. Algal Res. 13, 195–206 (2016).
Mears, L., Stocks, S. M., Sin, G. & Gernaey, K. V. A review of control strategies for manipulating the feed rate in fed-batch fermentation processes. J. Biotechnol. 245, 34–46 (2017).
Sommeregger, W. et al. Quality by control: Towards model predictive control of mammalian cell culture bioprocesses. Biotechnol. J. 12, 1600546 (2017).
Chen, G., Hu, J., Qin, Y. & Zhou, W. Viable cell density on-line auto-control in perfusion cell culture aided by in-situ Raman spectroscopy. Biochem. Eng. J. 172, 108063 (2021).
Lee, H. L., Boccazzi, P., Gorret, N., Ram, R. J. & Sinskey, A. J. In situ bioprocess monitoring of Escherichia coli bioreactions using Raman spectroscopy. Vib. Spectrosc. 35, 131–137 (2004).
Xu, J., Liu, N., Qiao, K., Vogg, S. & Stephanopoulos, G. Application of metabolic controls for the maximization of lipid production in semicontinuous fermentation. Proc. Natl Acad. Sci. USA 114, E5308–E5316 (2017).
Dong, T. et al. Direct quantification of fatty acids in wet microalgal and yeast biomass via a rapid in situ fatty acid methyl ester derivatization approach. Appl. Microbiol. Biotechnol. 99, 10237–10247 (2015).
Yuan, J.-P., Chen, F., Liu, X. & Li, X.-Z. Carotenoid composition in the green microalga Chlorococcum. Food Chem. 76, 319–325 (2002).
Chen, Y. et al. An unconventional uptake rate objective function approach enhances applicability of genome-scale models for mammalian cells. NPJ Syst. Biol. Appl. 5, 1–11 (2019).
Acknowledgements
This work was supported by the U.S. National Science Foundation EFRI program (Grant number: 1332344) and CBET program (Grant number: 1804733) and the Department of Energy (Grant number: DE-SC0019388).
Author information
Authors and Affiliations
Contributions
C.T.L., R.E., C.Z., Y.C., and M.J.B. contributed to conception and design of the experiment. C.T.L., R.E., and K.W.H. conducted the experiments. C.T.L., R.E., and M.J.B. analyzed the data. C.T.L., R.E., C.Z., K.Z., and M.J.B. drafted the paper. All authors read and approved the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, CT., Eng, R., Zuniga, C. et al. Optimization of nutrient utilization efficiency and productivity for algal cultures under light and dark cycles using genome-scale model process control. npj Syst Biol Appl 9, 7 (2023). https://doi.org/10.1038/s41540-022-00260-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41540-022-00260-7