Abstract
Evidence for epigenetic regulation playing a role in Parkinson’s disease (PD) is growing, particularly for DNA methylation. Approximately 90% of PD cases are due to a complex interaction between age, genes, and environmental factors, and epigenetic marks are thought to mediate the relationship between aging, genetics, the environment, and disease risk. To date, there are a small number of published genome-wide studies of DNA methylation in PD, but none accounted for cell type or sex in their analyses. Given the heterogeneity of bulk brain tissue samples and known sex differences in PD risk, progression, and severity, these are critical variables to account for. In this genome-wide analysis of DNA methylation in an enriched neuronal population from PD postmortem parietal cortex, we report sex-specific PD-associated methylation changes in PARK7 (DJ-1), SLC17A6 (VGLUT2), PTPRN2 (IA-2β), NR4A2 (NURR1), and other genes involved in developmental pathways, neurotransmitter packaging and release, and axon and neuron projection guidance.
Similar content being viewed by others
Introduction
Parkinson’s disease (PD) is the second most common neurodegenerative disorder in the US, and is characterized by progressive degeneration of dopaminergic neurons in the nigrostriatal pathway and the formation of α-synuclein-containing Lewy bodies1. An estimated 5–10% of PD cases are familial and several genes have been identified that cause these inherited forms of the disease2,3. The remaining ~90% of cases are likely due to a complex interaction between age, genes, and environmental factors. Given that epigenetic marks are dynamic with age, sensitive to the environment, and regulate gene expression throughout the lifespan, they are considered a potential mediator of the complex relationship between aging, genetics, the environment, and disease disease4,5.
Evidence for the role of epigenetic regulation in PD is growing, particularly for DNA methylation6,7,8,9,10,11,12. In particular, previous studies have reported differential DNA methylation at PD-related genes (MAPT, CYP2E1, and STX1B)13,14,15, and hypomethylation of the α-syn gene (SNCA) is associated with decreased protein levels in the substantia nigra (SN) and striatum of postmortem PD brain16,17,18,19,20,21,22,23. In addition to these gene-specific studies, genome-wide analyses of DNA methylation from postmortem PD brain tissue have identified a number of gene regions that show differential DNA methylation in PD brains24,25,26,27. Finally, a case-control study identified an association between a polymorphism in PD risk and DNA methyltransferase 3B (DNMT3B), which is responsible for establishing de novo patterns of DNA methylation during embryonic development28. Taken together, these multiple lines of evidence support the role for dysregulation of DNA methylation in the etiology of PD.
However, the existing genome-wide analyses of DNA methylation in PD have not adequately addressed the effects of sex or cell-type heterogeneity. Given that men and women show differences in PD risk, disease progression, and disease severity29,30,31, it is critical to include sex in experiments examining the potential role for the epigenome in PD. Furthermore, the majority of existing studies of epigenetics in PD have utilized either blood or bulk brain tissue samples24,26,32,33. Although DNA methylation changes in blood are useful for identifying biomarkers, they may not be representative of a pathogenic mechanism in the brain. Further, analyses from bulk brain tissue cannot address the cell-type specificity of identified changes, making it difficult to determine which cell types may be driving PD-related differential DNA methylation. In a step forward for the field, one recent paper investigated DNA methylation in an enriched neuronal population using fluorescence-activated cell sorting, but only explored DNA methylation at enhancer regions, not genome-wide27. Here, we report the results from a genome-wide analysis of DNA methylation in enriched neurons from PD brain stratified by sex.
In this study, we obtained human postmortem parietal cortex (n = 50 control, n = 50 Parkinson’s disease) from the Banner Sun Health Research Institute Brain Bank and enriched for neuronal populations using magnetic-activated cell sorting (MACS) for the neuronal marker, NeuN. Postmortem parietal cortex was used because this region develops pathology in the late stages of PD and is expected to still have robust populations of neurons, from which disease-affected gene regulatory marks can be measured. Genome-wide DNA methylation was measured using the Illumina EPIC BeadChip array paired with bisulfite treatment. Bisulfite treatment is a well-established method for measuring DNA methylation, but it actually cannot distinguish between DNA methylation and DNA hydroxymethylation34,35. As such, our dataset actually measures both DNA methylation and DNA hydroxymethylation without discriminating between these two epigenetic marks. Despite this important caveat, for simplicity’s sake and to match prior literature, we will discuss our results as changes in “DNA methylation.”
We also tested the hypothesis that PD is associated with accelerated epigenetic aging in the brain. Aging is the primary risk factor for PD, and previous work has shown that aging and PD progress via shared cellular mechanisms36,37,38,39. Multiple hypotheses have been proposed to explain the association between aging and the development of PD, including the ‘multiple hit’ hypothesis and the ‘stochastic acceleration hypothesis’37,40. In general, these hypotheses work under a similar biological model – that an accumulation of factors, both genetic and environmental, accelerate the normal pace of dopaminergic neuron loss with age, eventually exceeding a threshold for PD diagnosis37. However, the biological mechanism by which environment and genetics interact to accelerate age-related dopaminergic neuron loss remains unclear. Because the epigenome is dynamic with age and shows programmed age-related changes, it has been proposed that acceleration of these age-related changes can contribute to disease risk in the aged human41,42,43,44. Supporting this idea, studies have shown associations between accelerated epigenetic aging and a variety of disease states, including a study using blood samples from PD patients32,45. However, to our knowledge, no existing studies have investigated whether neurons show altered epigenetic aging in PD brains compared to control.
Results
Human brain tissue sample selection
De-identified tissue samples from control (n = 50) and Parkinson’s disease (n = 50) human brain samples were obtained from archival human autopsy specimens provided by the Banner Sun Health Research Institute (BSHRI), using BSHRI’s approved institutional review board (IRB) protocols. Further details about the BSHRI’s brain samples are available in a previous publication46. We selected PD patients with mid-stage disease (Braak stage = II–III), as defined by Lewy pathology47. The cohort of control brains consisted of patients who died from non-neurologic causes and whose brains had no significant neurodegenerative disease pathology. Subjects were split by sex (n = 63 males, n = 37 females), and data for the sample cohort are summarized in Table 1. For each subject (N = 100), the parietal cortex was obtained. This region develops pathology late in PD; in mid-stage PD, the parietal cortex is expected to still have robust populations of neurons (unlike the substantia nigra, where neuron loss occurs early in disease), providing an avenue to investigate pre-pathological changes in gene regulation.
Validation of neuronal enrichment
To validate the enrichment of NeuN+ nuclei via MACS prior to experimentation, we quantified total nuclei and NeuN+ nuclei for six validation samples using flow cytometry. A representative quantification plot is included in Fig. 1a. Plots for all six validation samples are available as a supplementary file (Supplementary Fig. 1). For the validation samples, the mean % NeuN+ was 92.0% (range = 89.2–95.2%). As a secondary validation, we estimated proportions of neuronal and glial cell types for all experimental MACS-sorted parietal cortex samples using the Cell Epigenotype Specific (CETS) package during EPIC array data processing. One sample from the male control group that was not one of the six with validation by flow cytometry had an estimated neuronal population of only 0.03% in the CETS analysis. This sample was removed prior to all downstream differential methylation testing. The average proportion of neurons in the remaining 99 samples was estimated to be 83.8% by CETS analysis (Fig. 1b).
a NeuN+ nuclei were identified using DAPI pulse area vs. width using the 405-450/45 channel for all nuclei (left) followed by NeuN-AlexaFluor488 bright events 488-525/40 (right). The percentage of NeuN+ nuclei is provided in the top-right corner. b Histogram of estimated neuronal proportion (vs. glial) for enriched PD NeuN+ nuclei population. Proportions of neuronal and glial cell types were estimated for MACS-sorted NeuN+ nuclei samples using the CETS R package during EPIC array data processing. Average proportion of neurons (vertical dotted line) was estimated to be 83.8% across all n = 99 samples included in analysis.
Differential testing for differentially methylated CpGs
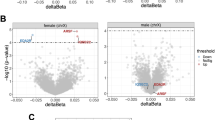
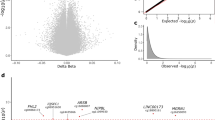
In our sex-stratified models for differential DNA methylation testing, we identified 3 differentially methylated CpGs (DMCs) in males and 87 significant DMCs in the female parietal cortex associated with PD (p-value < 9 × 10−8) (Additional files 1 and 2). This p-value cutoff was selected based on recommendations in a recent study of epigenome-wide association studies48. The sex-stratified DMC models included a sigma term to test for differences in the mean by disease state while accounting for potential differences in variability. The male DMCs annotated to 3 unique gene IDs and the female DMCs annotated to 85 unique gene IDs; two female DMCs annotated to the same gene. These results were sex-specific, and there was no overlap between any of the identified male and female DMCs by probe ID.
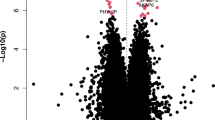
Here, we highlight the most significant DMCs and a DMC annotated to a gene of interest for PD. The most significant DMC in males (p-value = 1.38 × 10−9) was located within the PARK7 locus and showed male-specific hypomethylation in PD cortical neurons compared to control (Fig. 2a, b). Visualization of the identified DMCs in the Wash U Epigenome Browser with NIH ROADMAP Epigenomics chromatin state predictions show that this DMC is located in a CpG island flanking the transcription start site (Fig. 4a)49,50. In females, the most significant DMC was located within the gene body of the ATXN1 gene (p-value = 1.84 × 10−17). This ATXN1 DMC is located within a predicted enhancer associated with weak transcriptional activity and showed female-specific hypermethylation in PD cortical neurons compared to control (Figs. 2b, 4d)49,50. We also identified a female-specific hypomethylated DMC within the gene body of the SLC17A6 locus (p-value = 1.45 × 10−10) (Figs. 2c, d, 4b)49,50.
a In males, the most significant DMC was annotated to the PARK7 locus (cg08027640; p-value = 1.38 × 10−9). b There was no significant change in DNA methylation at cg08027640 in females. c There was no significant change in DNA methylation at cg19566879 in males. d In females, the most significant DMC was annotated to the ATXN1 locus (cg19566879; p-value = 1.84 × 10−17). e There was no significant change in DNA methylation at cg09150064 in males. f In females, a significant DMC was annotated to the SLC17A6 locus (cg09150064; p-value = 1.45 × 10−10). P-values indicate significant change in PD compared to control (cutoff for significance: p-value < 9 × 10−8).
Differential testing for differentially methylated regions
In our sex-stratified models for differential genomic region testing, we identified 258 significant differentially methylated regions (DMRs) in males and 214 significant DMRs in female parietal cortex by Parkinson’s disease (minimum smoothed FDR < 0.05) (Additional files 3 and 4). These DMRs annotated to 205 unique gene IDs in the males and 157 unique gene IDs in the females. Here, we highlight 3 DMRs of interest.
Comparing these two lists of DMR by chromosomal location, 5 regions showed at least partial overlap between the two sexes (Table 2), and one region—annotated to the PTPRN2 gene—showed exact, complete overlap in both males and females (Fig. 3a, b). This region of complete overlap (chr7:158093198–158093277) includes 3 CpGs—cg11293572, cg09992350, and cg27014435 – and is annotated to an intronic region within the PTPRN2 gene body (Fig. 4e)49,50. Although this region was a DMR in both males and females, it was hypermethylated in male brains and hypomethylated in female brains, so we have highlighted this gene (Table 2, Fig. 3). Additional DMCs were also identified with the PTPRN2 gene body (Fig. 4e)49,50. The other 4 regions of partial overlap – annotated to the ZIC1, GALNT15, KIAA0040, and GFPT2 genes – also showed opposite relationships between PD status and DNA methylation in males and females (Table 2).
Beta values (y-axis) represent DNA methylation across CpGs included in DMRs annotated to PTPRN2 (chr7: 158093198–158093277), SLC17A6 (chr11:22362708-22364961), and NR4A2 (chr2:157186666-157186681). a In males, PD brains (red) exhibited significant hypermethylation at PTPRN2 compared to control (black). b In females, PD brains (purple) exhibited significant hypomethylation at PTPRN2 compared to control (black). c In males, PD brains (red) did not exhibit significant differential methylation at SLC7A6 compared to control (black). d In females, PD brains (purple) exhibited significant hypomethylation at SLC17A6 compared to control (black). Significant DMC (cg09150064) indicated with box and asterisk. e In males, PD brains (red) exhibited significant hypermethylation at NR4A2 compared to control (black). f In females, PD brains (purple) did not exhibit significant differential methylation compared to control (black). Significant DMRs are indicated with minimum smoothed FDR values < 0.05. Thick solid and dashed lines represent smoothed means by group across DMRs.
Selected genes containing PD-associated DMCs and DMRs were visualized in the Wash U Epigenome Browser. Tracks included, from top to bottom: Gencode gene annotation, chromosomal location ruler, CpG context, DMCs and DMRs, location of EPIC probes, and chromatin state (NIH Roadmap Epigenomics chromHMM imputation for cortical neurospheres). a PARK7: A single DMC was identified in the predicted transcription start site. b SLC17A6: The identified DMR overlaps with a predicted bivalent promoter and inactive repressed polycomb region, while the DMC is located within the predicted repressed polycomb region. c NR4A2: A DMC was identified within a predicted bivalent promoter. d ATXN1: The identified DMC is located within an enhancer associated with weak transcription. e PTPRN2: Multiple DMC/DMRs were identified in this gene, with one located within the transcription start site and others within heterochromatin or areas of low activity49,50.
In addition to overlapping DMRs, we also identified a female-specific DMR annotated to the SLC17A6 gene, which included the DMC identified at this gene (Figs. 3d, 4b)49,50. We highlight this gene since it has previously been implicated in PD and was identified as containing most significant female-specific DMC (Fig. 2c, d). This DMR (chr11:22362708–22364961) spans a CpG island and chromatin annotated as a bivalent promotor region (Fig. 4b)49,50. The SLC17A6 DMR was not significant in males (Fig. 3c).
We also identified a male-specific DMR annotated to the NR4A2 gene (Fig. 3e), which we previously identified as a gene of interest for PD risk in a mouse model of developmental pesticide exposure51. This DMR (chr2:157186666–157186681) includes 2 CpGs—cg21226516 and cg11209121—that are located at an exon-intron boundary in the gene body of NR4A2 and within chromatin annotated as bivalent chromatin (Fig. 4c)49,50. The NR4A2 DMR was not significant in the female data (Fig. 3f). These results underscore that PD-related changes in cortical neuron DNA methylation are sex-specific.
Gene ontology pathway enrichment
The clusterProfiler R package was used to determine whether DMCs annotated to genes enriched for specific gene ontology biological process (GOBP) pathways. GOBP pathway enrichment analysis was not performed on male DMCs due to the low number of significant CpGs (n = 3) but was performed on the genes annotated to female hypomethylated DMCs and hypermethylated DMCs separately. Hypo- and hypermethylated DMCs were considered separately due to expected differences in associations between increased or decreased DNA methylation and gene regulation. Female hypermethylated DMCs did not show enrichment for any pathways, but female hypomethylated DMCs showed enrichment for 10 pathways, including several developmental pathways, neurotransmitter transport, neurotransmitter secretion, and signal release from synapse (Fig. 5a). Several differentially methylated genes were included in these GO term pathways, including SLC17A6 and PTPRN2 (Fig. 5b), the latter of which showed differential DNA methylation at the regional level in both males and females.
a Female hypomethylated DMCs showed enrichment for 10 pathways, including neurotransmitter transport, neurotransmitter secretion, and signal release from synapse. The x-axis is the number of genes in each pathway, color is used to represent FDR q-value (q-value), and size is used to represent GeneRatio, the ratio of differentially methylated genes in each GO term to the total number of genes input into the hypergeometric test. b Several differentially methylated genes were included in the enriched pathways, including PTPRN2, which also showed differential DNA methylation at the region level in both males and females. The size of each node is used to represent the number of genes in each GO term, and color represents the magnitude of decrease in DNA methylation (beta value) for each annotated probe. Connections between genes and GO terms represent inclusion of the gene in that GO term.
The clusterProfiler R package was also used to determine whether DMRs annotated to genes enriched for specific gene ontology biological process (GOBP) pathways. In both males and females, hypermethylated DMRs did not show any enrichment for GOBP terms. Meanwhile, for the hypomethylated DMRs, females showed significant enrichment for 6 GOBP terms (q-value < 0.05), including cell fate commitment and glucose homeostasis (Additional File 5). These enriched pathways for the female hypomethylated DMRs are collectively related to cell fate and metabolism. In male hypomethylated DMRs, 4 pathways approached significance (q-value = 0.06), including semaphorin-plexin signaling pathways involved in axon guidance and neuron projection guidance (Additional File 5).
Epigenetic clock analysis
In our epigenetic clock analysis, estimated epigenetic age significantly predicted chronological age in both sexes, with estimated age (“EpiAge”) significantly associated with chronological age at death (males: Beta coefficient = 1.1929, p-value = 8.97 × 10−9; females: Beta coefficient = 1.3515, p-value = 2.51 × 10−9). These results confirm that this is a well-calibrated epigenetic clock where there was an approximate 1 unit increase in estimated epigenetic age with a 1 unit increase in chronological age (Table 3). However, in our data, PD status did not modify trajectories of epigenetic aging in either sex (Fig. 6, Table 3).
Estimated epigenetic age showed a significant positive association with chronological age in males (left) and females (right). However, PD status did not significantly modify trajectories of epigenetic age. Control samples are shown as black dots, and PD samples are shown as red or purple triangles for males and females, respectively.
Discussion
In this study, we identified sex-specific, PD-associated DNA methylation changes in a total of 434 unique genes, including PARK7 (DJ-1), SLC17A6 (VGLUT2), PTPRN2 (IA-2β), NR4A2 (NURR-1), as well as other genes involved in developmental pathways, neurotransmitter packaging, and release, and axon/neuron projection guidance (Figs. 2–5). Our data did not show accelerated epigenetic aging in PD (Fig. 6, Table 3). This study expands on the existing literature through the utilization of nuclei-sorted cortical neurons, the inclusion of both sexes, and estimation of epigenetic age.
Sex-specificity of PD-associated differential DNA methylation
Sex is known to have significant effects on the epigenome, and recent work has shown sex-specific directionality of associations between DNA methylation and Alzheimer’s disease, suggesting that sex might also affect the relationship DNA methylation and other neurodegenerative diseases52. Given this fact, as well as the known sex differences in susceptibility and progression of PD by sex, we chose to stratify our analysis by sex, providing a genome-wide study of DNA methylation in PD that analyzed male and female samples separately29,30,31. To confirm that sex stratification was preferred, we compared p-value histograms between sex-adjusted and sex-stratified models, showing that the sex-stratified models provided more appropriate histograms (uniform with an overabundance of low p-values), whereas the combined model produced a non-uniform histogram with a skew towards high p-values (Supplementary Figure 2). This underscores the importance of stratifying data by sex rather than including as a covariate during analysis. While we expected to identify differences in DNA methylation in male and female samples, the near-total lack of overlap between sexes was striking, suggesting that epigenetic regulation may underlie the well-recognized sex differences in PD susceptibility and progression.
Male-specific PD-associated DNA hypomethylation at the PARK7 locus
The most significant DMC in males showed PD-related hypomethylation and was located in the PARK7 locus within a CpG island that spans the transcription start site (Figs. 2, 4). Mutations in the PARK7 locus, which encodes the DJ-1 protein, cause autosomal recessive, early-onset Parkinson’s disease and oxidized DJ-1 has been observed in brains of idiopathic PD patients53,54. Existing research shows that DJ-1 has multiple known neuroprotective functions—transcriptional regulator, molecular chaperone, and antioxidant—and downregulation of this locus is thought to play a role in both familial and idiopathic forms of PD55,56,57,58. Thus, it is possible that epigenetic regulation of this locus modifies expression in idiopathic PD.
Two previous studies examining the effects of PD status on DNA methylation at the PARK7 locus failed to identify a significant relationship59,60. However, these studies used bisulfite pyrosequencing to measure DNA methylation at regions in the PARK7 locus that do not overlap our DMC. In addition, one study was performed in blood leukocytes, which may not reflect DNA methylation in brain tissue. The other utilized a very small sample size (n = 5 per group) and used bulk postmortem brain tissue samples from substantia nigra, parietal cortex, and occipital cortex59,60. Given these differences, the existing data do not necessarily contradict our PARK7 results.
Female-specific PD-associated DNA hypermethylation at the ATXN1 locus
The most significant DMC in female subjects was located in a weak enhancer region within the ATXN1 gene, which showed PD-related hypermethylation (Figs. 2, 4). ATXN1 encodes Ataxin-1 (ATXN1), which is involved in transcriptional repression of a large number of target genes and is necessary for normal brain development and function61,62. Expanded polyglutamine repeats in this gene cause spinocerebellar ataxia type 1 (SCA1)61,63. Although the ATXN1 gene has not been previously studied in the context of PD, our data showing altered epigenetic regulation of the ATXN1 gene in PD brains opens new avenues of research. It is possible that epigenetic regulation at this locus leads to changes in the expression of ATXN1 and its target genes, affecting the susceptibility of cortical neurons to PD-related dysfunction and pathology.
Male-specific PD-associated changes in DNA methylation at NR4A2
In males, we also identified PD-related hypermethylation in an exon just downstream of an exon-intron boundary in the gene body of NR4A2 (Figs. 3, 4). NR4A2 encodes the nuclear receptor related-1 (NURR1) protein, a transcription factor critical for dopaminergic neuron development and maintenance. Previous research suggests that downregulation of the NR4A2 gene in the brain may contribute to PD pathogenesis64,65,66. Supporting this idea, in a previous animal model study, we showed that developmental exposure to the organochlorine pesticide dieldrin, which is known to be associated with increased PD risk, led to sex-specific changes in DNA methylation in the Nr4a2 gene body in mice51. These previous results, along with our present data, suggest that sex-specific epigenetic regulation at the NR4A2 gene may play a role in idiopathic PD risk.
Female-specific PD-associated DNA hypomethylation at the SLC17A6 locus
In females, we identified PD-related hypomethylation within the SLC17A6 gene body (Figs. 2–4). SLC17A6 encodes the vesicular glutamate transporter 2 (VGLUT2), which has been implicated in PD pathogenesis via multiple lines of evidence67. Studies using in vivo and in vitro models show increased VGLUT2 expression in dopaminergic (DA) neurons in response to the neurotoxicants, rotenone, 6-OHDA and MPP+68,69. Further, deletion of VGLUT2 in mice exacerbates the neurotoxic effects of 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine (MPTP) exposure70. Additional studies suggest that neurons that co-express VGLUT2 and VMAT2 show differentially vulnerability in PD and differences in neuronal structure71,72. Our analysis did not assess DNA methylation in DA neurons of the nigrostriatal pathway, which have been the subject of much of this work on VGLUT2 in the context of PD. However, the observed hypomethylation in the SLC17A6 gene in the parietal cortex suggests that this locus could show altered epigenetic regulation of gene expression during disease progression. Given the known neuroprotective role for VGLUT2, future studies should examine whether epigenetic changes at SLC17A6 are associated with decreased VGLUT2 expression and increased neuronal susceptibility.
Sex-specific PD-associated changes in DNA methylation at PTPRN2
In our regional analysis, one DMR—annotated to the PTPRN2 gene—showed exact, complete overlap in both males and females (Table 2). Although the identified PTPRN2 DMR was differentially methylated in both males and females, it was hypermethylated in brains from male PD patients and hypomethylated in brains from female PD patients (Fig. 3). PTPRN2 encodes Protein Tyrosine Phosphatase Receptor Type N2 (IA-2β), which is expressed on dense core and synaptic vesicles. Altered methylation of this gene has been previously implicated in PD in a study of longitudinal blood samples that showed an association between hypomethylation of a different CpG within the PTPRN2 gene in whole blood and faster motor progression in PD73. In addition, deletion of this gene and the related IA-2 protein in mice leads to reduced levels of norepinephrine, dopamine (DA) and serotonin in the brain and decreased release of DA, gamma-aminobutyric acid (GABA), and glutamate from synaptosomes74. One previous study found decreased expression of PTPRN2 in the SN of PD patients, while another found increased expression in DA neurons from PD patients with LRRK2 G2019S mutations75,76. Interestingly, given the known role of environmental exposures in PD and work in our lab investigating pesticide exposures and PD51,77, another study found an association between pesticide exposure and hypomethylation at a CpG in PTPRN2 in blood78. The opposite directionality of the relationship between PTRPN2 DNA methylation and PD status in males and females suggests that differential epigenetic regulation may be important in sex differences in PD.
Sex-specific pathways enriched for DNA methylation changes
In addition to our gene-level differences, pathway analysis of differentially methylated genes identified distinct enriched pathways in male and female subjects (Fig. 5). Female-specific differentially methylated genes were enriched for pathways including neurotransmitter transport, neurotransmitter secretion, and signal release from synapse (Fig. 5c). These pathways were not enriched in differentially methylated genes identified in male subjects. Instead, the semaphorin-plexin signaling pathways involved in axon guidance and neuron projection guidance were enriched in male-specific differentially methylated genes. These pathways included PLXNB1, PLXNC1, and PLXNB3, three genes that encode proteins in the plexin family, which act as transmembrane receptors for the semaphorins. The semaphorin-plexin signaling pathway is a key regulator of morphology and motility in many different cell types, including those that make up the nervous system, and is known to play a role in axon guidance79,80. As with our gene-specific results, the sex-specificity of these networks is striking and reinforces that epigenetic regulation in PD is highly sex-specific and may contribute to the known sex differences in PD.
PD is not associated with accelerated epigenetic aging
Over the past decade, recognition has been growing that the epigenome, especially DNA methylation, can be used to closely predict chronological age41,81,82,83. Based on this, researchers have established epigenetic clocks that use loci with the most predictable DNA methylation levels to provide estimates of epigenetic (or biological) age81. In a previous study using one of these epigenetic clocks, researchers found accelerated epigenetic aging in PD blood samples compared to control45. However, to our knowledge, no studies have tested whether epigenetic aging is accelerated in PD brain tissue. Here, we used an epigenetic clock specifically designed for human cortical tissue to estimate epigenetic age of control and PD parietal cortex samples84. In our analyses, we showed that our epigenetic clock was well-calibrated to predict chronological age at death, but found no evidence of significant epigenetic age acceleration by PD status in our cohort (Fig. 6). These data suggest that accelerated epigenetic aging may not play a role in PD development. Alternatively, the lack of relationship between epigenetic aging and disease status could also be a byproduct of the selected brain region—parietal cortex—which shows effects of PD pathology very late in disease. In contrast, neurons from brain regions that are impacted early in PD (i.e. midbrain) might demonstrate positive relationships between epigenetic aging and disease. Furthermore, the epigenetic aging algorithm used in this study was designed for human cortical tissue, but not specifically for neurons from parietal cortex, which could inflate the variability of our epigenetic age estimates and limit our ability to detect significant associations. Lastly, it is also possible that an association between disease and epigenetic age could occur in middle-age, but disappear at late ages, as has been shown previously for obesity-related epigenetic aging85. Samples from PD and control subjects across a broader range of ages would be needed to test this hypothesis.
Conclusions
Our study provides a genome-wide analysis of DNA methylation in an enriched neuronal population from both male and female PD brain tissue. Despite our study design’s strengths, there remain some limitations in our analysis that create opportunities for future studies. First, our results do not distinguish between DNA methylation and DNA hydroxymethylation, which show distinct genomic distributions and associations with gene expression in the brain, as well as differential responses to environmental exposures9,86,87. As such, DNA methylation and DNA hydroxymethylation may play discrete roles in neurodegenerative diseases9,87,88. It is critical for future studies to use modifications to the bisulfite conversion protocol, like oxidative BS or enzymatic conversion methods, that can distinguish between these marks. Second, because we used the Illumina EPIC array in this study, our data did not provide complete genomic coverage across the identified target genes. As a result, future work must follow up on these data with loci-specific analysis using BS-pyrosequencing or other targeted approaches to assess all cytosines at the identified loci. Third, the EPIC array does not provide a true “genome-wide” wide analysis of DNA methylation. This array was designed largely for cancer research, not brain-specific and neurodegenerative disease studies, and many genomic regions of interest to the PD field are not covered on the array. True genome-wide analysis requires whole-genome bisulfite sequencing (WGBS), but WGBS remains cost-prohibitive to scale up for a large cohort, making it unfeasible for this project. Although our study had a sample size smaller than recent recommendations published by Mansell et al after sex stratification, we experimentally controlled for multiple variables, including sex, race, and age, which were included as covariates in their estimates48. This increases our statistical power relative to the proposed minimum sample size, but future studies should include larger cohorts or be combined with publicly available data. Finally, we only analyzed the neuron-enriched population from our MACS-sorted samples. Given the growing body of evidence supporting the importance of non-neuronal cell types in PD, future work will aim to carry out similar analyses in neuron-depleted samples.
Despite our experimental limitations, this study provides a genome-wide analysis of DNA methylation in enriched neurons from male and female human postmortem brain tissue. Results from this work support the idea that epigenetic regulation is an important mechanism in PD pathogenesis and point to specific genes and pathways for further study.
Methods
Magnetic-activated cell sorting
De-identified parietal cortex samples were obtained from BSHRI. NeuN-positive (NeuN+) nuclei were enriched from 100 mg of flash frozen parietal cortex tissue using a two-stage magnetic-assisted cell sorting (MACS) method. First, 100 mg of frozen tissue were briefly thawed on ice and homogenized in a 2 mL, 1.4 mm ceramic bead tube (Thermo Fisher Scientific, Cat. # 15–340–153) with 1 mL of Nuclear Extraction Buffer (NEB) for 10 s at 4 m/s. NEB consisted of 0.32 M sucrose, 0.01 M Tris-HCl pH 8.0, 0.005 M CaCl2, 0.003 M MgCl2, 0.0001 M EDTA, and 0.1% Triton X-100, up to a stock volume of 1 L using water. Immediately prior to use, 0.001 M DTT was added to NEB. Homogenized samples were loaded into a 13 mL ultracentrifuge tube (BeckmanCoulter, Cat. # 331372) with 4 mL of NEB. Using a glass pipette, 7 mL of sucrose solution was pipetted down the side of each sample tube to create a sucrose gradient. Sucrose solution consisted of 1.8 M sucrose, 0.01 M Tris-HCl pH 8.0, 0.003 M MgCl2, up to a stock volume of 1 L using water. Immediately prior to use, 0.001 M DTT was added to NEB. After addition of sucrose, samples were spun at 4 °C, 24,000 rpm in the Sorvall Wx+ Ultracentrifuge in a swing bucket rotor (TH-641). Once the centrifugation was complete, the supernatant and debris layer found at the concentration gradient were both removed with the use of a vacuum, while being careful not to disturb the pellet containing the nuclei at the bottom of the tube. Next, 1 mL of primary antibody (anti-Neun 488—Millipore, Cat. # MAB377X) in MACS buffer was added to each nuclei pellet and placed on ice for 10 min. MACS buffer consisted of 0.5% Bovine Serum Albumin solution (Sigma-Aldrich, Cat. # A1595) in PBS pH 7.2 (Gibco, Cat. # 20012-027). Samples were then mechanically pipetted up and down 10–15 times to completely dissolve the nuclei pellet within the primary antibody-MACS buffer solution. This solution of nuclei was then transferred to a 2 mL tube and incubated for 60 min at 4 °C. After incubation, 40 µL of MACS Microbeads (anti-mouse IgG Microbeads - Miltenyi, Cat. # 130-048-401) were added to each sample. Samples were then inverted 4–5 times and incubated at 4 °C for 30 min. After incubation, nuclei were centrifuged at 300g for 10 min. Supernatant was then removed, and the nuclei were resuspended in 2 mL of MACS buffer and transferred to a MACS MS column (MS Separation columns—Miltenyi, Cat. # 130-042-201) that was pre-washed with MACS buffer and attached to the Miltyeni OctoMACS™ Separator. Positive selection of NeuN+ cells was then performed according to the standard MACS MS Columns protocol available from Miltenyi Biotec. After the first round of magnetic separation, NeuN+ nuclei were run through a separate, second MACS MS column to maximize cell type enrichment.
Enrichment validation using flow cytometry
Isolated nuclei were analyzed for flow cytometry on a CytoFlex S (Beckman Coulter), and data were analyzed using FlowJo V10. Single nuclei were identified by the presence of DAPI staining in a plot of 405–450/45-width signal vs 405–450/45-area signal. These nuclei were then analyzed for NeuN expression by looking for A488 fluorescence in the 488–525/40 channel. Determination of A488 positivity was made by comparing to a sample stained only with DAPI to define the background. Percent positivity for each sample was defined as the percentage of events in the NeuN-A488 + gate, divided by the total number of events identified as Nuclei.
DNA extraction, bisulfite treatment, and EPIC arrays
DNA was isolated from enriched NeuN+ nuclei using the Qiagen QIAamp DNA Micro Kit (Cat. # 56304), with some modifications to maximize yield. Given that samples were already homogenized during nuclei isolation, the sample lysis and incubation steps of the QIAamp DNA Micro Kit protocol were removed. Instead, 20 uL of proteinase K were added directly to each MACS eluate. Samples were then vortexed for 15 seconds and incubated at room temperature for 15 min. In addition, the optional carrier RNA was added to Buffer AL, the incubation time after the addition of 100% ethanol was increased to 10 min, the incubation time for the elution buffer was increased to 5 min, and the final elution step was repeated using 10 mM Tris-HCl pH 8.0.
Intact genomic DNA yield was quantified by Qubit fluorometry (Life Technologies). Bisulfite conversion was performed on 500 ng genomic DNA using the TrueMethyl Array kit (Cambridge Epigenetix). All conversion reactions were cleaned using SPRI-bead purification and eluted in Tris buffer. Following elution, BS-converted DNA was denatured and processed through the EPIC array protocol. The EPIC array contains ~850,000 probes that query DNA methylation at CpG sites across a variety of genomic features, including CpG islands, RefSeq genic regions, ENCODE open chromatin, ENCODE transcription factor binding sites, and FANTOM5 enhancer regions. To perform the assay, converted DNA was denatured with 0.4 N sodium hydroxide. Denatured DNA was then amplified, hybridized to the EPIC bead chip, and an extension reaction was performed using fluorophore-labeled nucleotides per the manufacturer's protocol. Array BeadChips were scanned on the Illumina iScan platform.
EPIC array data processing
After scanning on the iScan platform, BeadChip IDAT files were imported into R and processed using an in-house bioinformatics pipeline. This pipeline combined the minfi (version 1.22.1), ChAMP (version 2.14.0), posibatch (version 1.0), ENmix (version 1.12.4), CETS (version 3.03), and dplyr (version 1.0.8) packages in R. Quality control was assessed for internal control probes using the ENmix plotCtrl function. Failed probes were identified as those with a detection p-value > 0.01 in any sample. Failed probes were then removed from downstream analyses when detection p-value was > 0.01 in >5% of samples. No samples exceeded the selected threshold of failed probes (>10%) to warrant removal prior to analysis. Cross-reactive probes and probes containing SNPs were removed based upon previous identification89,90. After removal of technical artifacts, dye-bias correction was performed with ssNoob within minfi91. The proportion of neuronal vs. glial cells in each sample was estimated with CETS92. Samples with estimated glial cell to neuronal cell proportion > 0.90 were removed from analysis. This removed one sample from the male data set, leaving n = 62 males. Batch effects were assessed using the ChAMP package93, and then a beta value matrix for all samples was corrected for batch and positional effects using the posibatch R package94. After beta value estimation, we filtered out samples with mean DNA methylation beta value < 0.01. This beta value cutoff was instituted due to increased variability and decreased interpretability of beta values at such low levels. As a final step, we removed probes that were not annotated to a gene in the Illumina EPIC array manifest to ensure the interpretability of results after modeling. After all filtering steps, 552,332 probes remained for differential DNA methylation testing in the sex-stratified datasets. Code for data processing, filtering, and modeling is available (Additional Files 6, 7, and 8).
Statistical analysis
Differential testing for individual CpG sites
The generalized additive models for location, scale, and shape (gamlss; version 5.4-1) R package was used to test for differentially methylated CpGs (DMCs)95. This package provides functions to perform multivariate regression modeling while maintaining flexibility regarding term structure and distribution. For DMC testing, we assumed a normal distribution (standard linear regression model) and included both the mu (mean) and sigma (scale parameter) in models to test for differences in the mean by disease state while accounting for potential differences in variability. Linear regression was used in lieu of negative-binomial or beta regression modeling due to recent work showing that linear regression does not increase the likelihood of false positives in EPIC array-based epigenome-wide association studies48. All models were stratified by sex, and age and estimated glial cell proportion were included as covariates. Sex stratification was performed in response to a skewed, non-uniform p-value histogram that was produced by an initial model that simply included a covariate to adjust for sex (Supplementary Figure 2). The need to stratify by sex was not unexpected and this possibility was included in our preregistration plan. After stratifying for sex, p-value histograms were largely uniform, with peaks near zero (Supplementary Figure in Dryad data repository), as would be expected from well-calibrated hypothesis tests96.
An unadjusted p-value cutoff of <9 × 10−8 was used to assess the significance of DMCs, as recommended in a recent report on human epigenome-wide association studies48. All statistical models were run using R statistical software (version 4.1.0). Annotation of detected differential probes was performed using the Illumina EPIC array manifests. Code for data processing, filtering, and modeling is available as supplementary files.
Differential testing for genomic regions
The DMRcate R package (version 2.8.5) was used to test for differentially methylated regions97. In this package, the model matrix was established using the following code: design = model.matrix(~factor(disease) + glial_cell_proportion + age). For DMR analysis, Lamba was set to 1000 and the minimum number of CpGs was set to 2. All models were stratified by sex, and both age and estimated glial cell proportion were once again included as covariates in the modeling. The Benjamini–Hochberg False Discovery Rate (FDR) method was used to generate minimum smoothed q-values that account for multiple testing98. For the regional analysis, the significance cutoff was set at a minimum smoothed FDR q-value < 0.05.
Gene ontology pathway enrichment
The clusterProfiler R package (version 4.2.2) was used to determine whether differentially methylated genes were enriched for specific gene ontology biological process (GOBP) pathways99. Specifically, the enrichGO function was used to run hypergeometric over-representation tests on lists of differentially methylated gene IDs. Gene ID lists were split into hypermethylated DMCs or DMRs and hypomethylated DMCs or DMRs prior to analysis. The list of genes with annotated probes in the EPIC array manifest was used as the universe of total tested genes. Separate, sex-stratified analyses were performed for genes annotated to DMCs and DMRs. GOBP pathway enrichment analysis was not performed on male DMCs due to the low number of significant CpGs (n = 3). For enrichment analyses in clusterProfiler, redundant pathways were consolidated using the simplify function with default parameters, and significance cutoff was set at FDR q-value < 0.05.
Epigenetic clock analysis
Epigenetic age was estimated for all parietal cortex samples using a recently published algorithm designed specifically for the human cortex84. Estimates of epigenetic age (years) and known age at death (years) were then compared using standard linear regression with an age*disease interaction term. The interaction term was used to determine whether categorical PD status (control vs. PD) altered trajectories of epigenetic age compared to known chronological age. Epigenetic age regression modeling was performed on all data combined, as well as sex-stratified data. For these regression analyses, significance was set at p-value < 0.05.
Data availability
This study was preregistered with Open Science Framework: https://osf.io/z4vbw100.
All data acquired and analyzed for this study are available in the associated Dryad data repository: https://doi.org/10.5061/dryad.7d7wm37w0.
References
Fahn, S. & Stanley, F. Description of Parkinson’s disease as a clinical syndrome. Ann. N. Y. Acad. Sci. 991, 1–14 (2003).
Farrer, M. J. Genetics of Parkinson disease: paradigm shifts and future prospects. Nat. Rev. Genet. 7, 306–318 (2006).
Ferreira, M. & Massano, J. An updated review of Parkinson’s disease genetics and clinicopathological correlations. Acta Neurol. Scand. 135, 273–284 (2017).
Allis, C. D. & Jenuwein, T. The molecular hallmarks of epigenetic control. Nat. Rev. Genet. 17, 487–500 (2016).
Faulk, C. & Dolinoy, D. C. Timing is everything: the when and how of environmentally induced changes in the epigenome of animals. Epigenetics 6, 791–797 (2011).
Jakovcevski, M. & Akbarian, S. Epigenetic mechanisms in neurological disease. Nat. Med. 18, 1194–1204 (2012).
Marques, S. & Outeiro, T. F. Epigenetics in Parkinson’s and Alzheimer’s diseases. Sub-Cell. Biochem. 61, 507–525 (2013). in.
Miranda-Morales, E. et al. Implications of DNA methylation in Parkinson’s disease. Front. Mol. Neurosci. 10, 225 (2017).
Cheng, Y., Bernstein, A. I., Chen, D. & Jin, P. 5-Hydroxymethylcytosine: a new player in brain disorders? Exp. Neurol. 268, 3–9 (2014).
Lardenoije, R., Pishva, E., Lunnon, K. & van den Hove, D. L. Neuroepigenetics of aging and age-related neurodegenerative disorders. Prog. Mol. Biol. Transl. Sci. 158, 49–82 (2018).
Labbé, C., Oswaldo, L.-B. & Ross, O. A. Epigenetic regulation in Parkinson’s disease. Acta Neuropathol. 132, 515–530 (2016).
Wullner, U., Kaut, O., DeBoni, L., Piston, D. & Schmitt, I. DNA methylation in Parkinson’s disease. J. Neurochem. 139, 108–120 (2016).
(IPDGC), I. P. D. G. C. & (WTCCC2), W. T. C. C. C. 2. A two-stage meta-analysis identifies several new loci for Parkinson’s disease. PLoS Genet. 7, e1002142 (2011).
Kaut, O., Schmitt, I. & Wüllner, U. Genome-scale methylation analysis of Parkinson’s disease patients’ brains reveals DNA hypomethylation and increased mRNA expression of cytochrome P450 2E1. Neurogenetics 13, 87–91 (2012).
Coupland, K. G. et al. DNA methylation of the MAPT gene in Parkinson’s disease cohorts and modulation by vitamin E. Vitr. Mov. Disord. 29, 1606–1614 (2014).
Jowaed, A., Schmitt, I., Kaut, O., Wüllner, U. & Wu, U. Methylation regulates alpha-synuclein expression and is decreased in Parkinson’s disease patients’ brains. J. Neurosci. 30, 6355–6359 (2010).
Matsumoto, L. et al. CpG demethylation enhances alpha-synuclein expression and affects the pathogenesis of Parkinson’s disease. PLoS ONE 5, 1–9 (2010).
Desplats, P. et al. α-synuclein sequesters Dnmt1 from the nucleus. J. Biol. Chem. 286, 9031–9037 (2011).
Ai, S. X. et al. Hypomethylation of SNCA in blood of patients with sporadic Parkinson’s disease. J. Neurol. Sci. 337, 123–128 (2014).
Tan, Y. Y. et al. Methylation of alpha-synuclein and leucine-rich repeat kinase 2 in leukocyte DNA of Parkinson’s disease patients. Park. Relat. Disord. 20, 308–313 (2014).
Song, Y. et al. Pyrosequencing analysis of SNCA methylation levels in leukocytes from Parkinson’s disease patients. Neurosci. Lett. 569, 85–88 (2014).
Pihlstrøm, L., Berge, V., Rengmark, A. & Toft, M. Parkinson’s disease correlates with promoter methylation in the α-synuclein gene. Mov. Disord. 30, 577–580 (2015).
Schmitt, I. et al. L-dopa increases α-synuclein DNA methylation in Parkinson’s disease patients in vivo and in vitro. Mov. Disord. 30, 1794–1801 (2015).
Masliah, E., Dumaop, W., Galasko, D. & Desplats, P. Distinctive patterns of DNA methylation associated with Parkinson disease. Epigenetics 8, 1030–1038 (2013).
Moore, K., McKnight, A. J., Craig, D. & O’Neill, F. Epigenome-wide association study for Parkinson’s disease. NeuroMolecular Med. 16, 845–855 (2014).
Young, J. I. et al. Genome-wide brain DNA methylation analysis suggests epigenetic reprogramming in Parkinson disease. Neurol. Genet. 5, e342 (2019).
Marshall, L. L. et al. Epigenomic analysis of Parkinson’s disease neurons identifies Tet2 loss as neuroprotective. Nat. Neurosci. 23, 1203–1214 (2020).
Pezzi, J. C. et al. Association between DNA methyltransferase gene polymorphism and Parkinson’s disease. Neurosci. Lett. 639, 146–150 (2017).
Cerri, S., Mus, L. & Blandini, F. Parkinson’s disease in women and men: what’s the difference? J. Parkinsons. Dis. 9, 501 (2019).
Georgiev, D., Hamberg, K., Hariz, M., Forsgren, L. & Hariz, G.-M. Gender differences in Parkinson’s disease: a clinical perspective. Acta Neurol. Scand. 136, 570–584 (2017).
Gillies, G. E., Pienaar, I. S., Vohra, S. & Qamhawi, Z. Sex differences in Parkinson’s disease. Front. Neuroendocrinol. 35, 370 (2014).
Henderson-Smith, A. et al. DNA methylation changes associated with Parkinson’s disease progression: outcomes from the first longitudinal genome-wide methylation analysis in blood. Epigenetics 14, 365–382 (2019).
Henderson, A. R. et al. DNA methylation and expression profiles of whole blood in Parkinson’s disease. Front. Genet. 0, 509 (2021).
Jin, S.-G., Kadam, S. & Pfeifer, G. P. Examination of the specificity of DNA methylation profiling techniques towards 5-methylcytosine and 5-hydroxymethylcytosine. Nucleic Acids Res. 38, e125 (2010).
Bernstein, A. I. & Jin, P. High-throughput sequencing-based mapping of cytosine modifications. in epigenetic technological applications. https://doi.org/10.1016/B978-0-12-801080-8.00003-X (2015).
Driver, J. A., Logroscino, G., Gaziano, J. M. & Kurth, T. Incidence and remaining lifetime risk of Parkinson disease in advanced age. Neurology 72, 432–438 (2009).
Collier, T. J., Kanaan, N. M. & Kordower, J. H. Ageing as a primary risk factor for Parkinson’s disease: evidence from studies of non-human primates. Nat. Rev. Neurosci. 12, 359–366 (2011).
Reeve, A., Simcox, E. & Turnbull, D. Ageing and Parkinson’s disease: why is advancing age the biggest risk factor? Ageing Res. Rev. 14, 19–30 (2014).
Rodriguez, M., Rodriguez-Sabate, C., Morales, I., Sanchez, A. & Sabate, M. Parkinson’s disease as a result of aging. Aging Cell 14, 293–308 (2015).
Sulzer, D. Multiple hit hypotheses for dopamine neuron loss in Parkinson’s disease. Trends Neurosci. 30, 244–250 (2007).
Horvath, S. DNA methylation age of human tissues and cell types. Genome Biol. 14, R115 (2013).
Jones, M. J., Goodman, S. J. & Kobor, M. S. DNA methylation and healthy human aging. Aging Cell 14, 924–932 (2015).
Jung, M. & Pfeifer, G. P. Aging and DNA methylation. BMC Biol. 13, 7 (2015).
Kochmanski, J., Montrose, L., Goodrich, J. M. & Dolinoy, D. C. Environmental deflection: the impact of toxicant exposures on the aging epigenome. Toxicol. Sci. 156, kfx005 (2017).
Horvath, S. & Ritz, B. R. Increased epigenetic age and granulocyte counts in the blood of Parkinson’s disease patients. Aging (Albany NY) 7, 1130–1142 (2015).
Beach, T. G. et al. Arizona Study of Aging and Neurodegenerative Disorders and Brain and Body Donation Program. Neuropathology 35, 354–389 (2015).
Goedert, M., Spillantini, M. G., Del Tredici, K. & Braak, H. 100 years of Lewy pathology. Nat. Rev. Neurol. 9, 13–24 (2013).
Mansell, G. et al. Guidance for DNA methylation studies: Statistical insights from the Illumina EPIC array. BMC Genomics 20, 366 (2019).
Li, D., Hsu, S., Purushotham, D., Sears, R. L. & Wang, T. WashU Epigenome Browser update 2019. Nucleic Acids Res. 47, W158–W165 (2019).
Ernst, J. & Kellis, M. ChromHMM: automating chromatin-state discovery and characterization. Nat. Methods 9, 215–216 (2012).
Kochmanski, J., VanOeveren, S. E., Patterson, J. R. & Bernstein, A. I. Developmental dieldrin exposure alters DNA methylation at genes related to dopaminergic neuron development and Parkinson’s disease in mouse midbrain. Toxicol. Sci. 169, 593–607 (2019).
Zhang, L. et al. Sex-specific DNA methylation differences in Alzheimer’s disease pathology. Acta Neuropathol. Commun. 9, 77 (2021).
Bonifati, V. et al. Mutations in the DJ-1 gene associated with autosomal recessive early-onset parkinsonism. Science 299, 256–259 (2003).
Repici, M. & Giorgini, F. DJ-1 in Parkinson’s disease: clinical insights and therapeutic perspectives. J. Clin. Med. 8, 1377 (2019).
Ariga, H. et al. Neuroprotective function of dj-1 in Parkinson’s disease. Oxid. Med. Cell. Longev. https://doi.org/10.1155/2013/683920 (2013).
Hijioka, M., Inden, M., Yanagisawa, D. & Kitamura, Y. DJ-1/PARK7: a new therapeutic target for neurodegenerative disorders. Biol. Pharm. Bull. 40, 548–552 (2017).
Taira, T. et al. DJ-1 has a role in antioxidative stress to prevent cell death. EMBO Rep. 5, 213–218 (2004).
Kahle, P. J., Waak, J. & Gasser, T. DJ-1 and prevention of oxidative stress in Parkinson’s disease and other age-related disorders. Free Radic. Biol. Med. 47, 1354–1361 (2009).
Tan, Y. et al. Methylation status of DJ-1 in leukocyte DNA of Parkinson’s disease patients. Transl. Neurodegener. 5, 5 (2016).
Navarro-Sánchez, L., Águeda-Gómez, B., Aparicio, S. & Pérez-Tur, J. Epigenetic study in Parkinson’s disease: a pilot analysis of DNA methylation in candidate genes in brain. Cells 7, 150 (2018).
Lu, H.-C. et al. Disruption of the ATXN1–CIC complex causes a spectrum of neurobehavioral phenotypes in mice and humans. Nat. Genet. 49, 527–536 (2017).
Crespo-Barreto, J., Fryer, J. D., Shaw, C. A., Orr, H. T. & Zoghbi, H. Y. Partial loss of ataxin-1 function contributes to transcriptional dysregulation in spinocerebellar ataxia type 1 pathogenesis. PLOS Genet 6, e1001021 (2010).
Zoghbi, H. Y. & Orr, H. T. Pathogenic mechanisms of a polyglutamine-mediated neurodegenerative disease, spinocerebellar ataxia type 1. J. Biol. Chem. 284, 7425–7429 (2009).
Luo, Y. The function and mechanisms of nurr1 action in midbrain dopaminergic neurons, from development and maintenance to survival. Int. Rev. Neurobiol. 102, 1–22 (2012).
Dong, J., Li, S., Mo, J. L., Cai, H. B. & Le, W. D. Nurr1-based therapies for Parkinson’s disease. CNS Neurosci. Ther. 22, 351–359 (2016).
Decressac, M., Volakakis, N., Björklund, A. & Perlmann, T. NURR1 in Parkinson disease—from pathogenesis to therapeutic potential. Nat. Rev. Neurol. 9, 629–636 (2013).
Kashani, A., Betancur, C., Giros, B., Hirsch, E. & El Mestikawy, S. Altered expression of vesicular glutamate transporters VGLUT1 and VGLUT2 in Parkinson disease. Neurobiol. Aging 28, 568 (2007).
Kouwenhoven, W. M. et al. VGluT2 expression in dopamine neurons contributes to postlesional striatal reinnervation. J. Neurosci. 40, 8262–8275 (2020).
Buck, S. A. et al. VGLUT2 is a determinant of dopamine neuron resilience in a rotenone model of dopamine neurodegeneration. J. Neurosci. 41, 4937–4947 (2021).
Shen, H. et al. Genetic deletion of vesicular glutamate transporter in dopamine neurons increases vulnerability to MPTP-induced neurotoxicity in mice. Proc. Natl Acad. Sci. 115, E11532–E11541 (2018).
Steinkellner, T. et al. Role for VGLUT2 in selective vulnerability of midbrain dopamine neurons. J. Clin. Invest. 128, 774 (2018).
Eskenazi, D. et al. Dopamine neurons that cotransmit glutamate, from synapses to circuits to behavior. Front. Neural Circuits 15, 665386 (2021).
Chuang, Y.-H. et al. Longitudinal epigenome-wide methylation study of cognitive decline and motor progression in Parkinson’s disease. J. Parkinsons. Dis. 9, 389 (2019).
Nishimura, T., Kubosaki, A., Ito, Y. & Notkins, A. L. Disturbances in the secretion of neurotransmitters in IA-2/IA-2β null mice: changes in behavior, learning and life span. Neuroscience 159, 427 (2009).
Grünblatt, E. et al. Gene expression profiling of parkinsonian substantia nigra pars compacta; alterations in ubiquitin-proteasome, heat shock protein, iron and oxidative stress regulated proteins, cell adhesion/cellular matrix and vesicle trafficking genes. J. Neural Transm. 111, 1543–1573 (2004).
Sandor, C. et al. Transcriptomic profiling of purified patient-derived dopamine neurons identifies convergent perturbations and therapeutics for Parkinson’s disease. Hum. Mol. Genet. 26, 552–566 (2017).
Gezer, A. O. et al. Developmental exposure to the organochlorine pesticide dieldrin causes male-specific exacerbation of α-synuclein-preformed fibril-induced toxicity and motor deficits. Neurobiol. Dis. 141, 104947 (2020).
van der Plaat, D. A. et al. Occupational exposure to pesticides is associated with differential DNA methylation. Occup. Environ. Med. 75, 427–435 (2018).
Alto, L. T. & Terman, J. R. Semaphorins and their signaling mechanisms. Methods Mol. Biol. 1493, 1 (2017).
Yazdani, U. & Terman, J. R. The semaphorins. Genome Biol. 2006 73 7, 1–14 (2006).
Horvath, S. & Raj, K. DNA methylation-based biomarkers and the epigenetic clock theory of ageing. Nat. Rev. Genet. 19, 371–384 (2018).
Levine, M. E. et al. An epigenetic biomarker of aging for lifespan and healthspan. Aging (Albany NY) 10, 573 (2018).
Bergsma, T. & Rogaeva, E. DNA methylation clocks and their predictive capacity for agingphenotypes and healthspan. Neurosci. Insights 15, 2633105520942221 (2020).
Shireby, G. L. et al. Recalibrating the epigenetic clock: implications for assessing biological age in the human cortex. Brain 143, 3763–3775 (2020).
Nevalainen, T. et al. Obesity accelerates epigenetic aging in middle-aged but not in elderly individuals. Clin. Epigenetics 9, 20 (2017).
Chen, Y., Damayanti, N. P., Irudayaraj, J., Dunn, K. & Zhou, F. C. Diversity of two forms of DNA methylation in the brain. Front. Genet. 5, 46 (2014).
Kochmanski, J. & Bernstein, A. I. The impact of environmental factors on 5-hydroxymethylcytosine in the brain. Curr. Environ. Heal. Reports 1–12 (2020) https://doi.org/10.1007/s40572-020-00268-3 (2020).
Al-mahdawi, S., Virmouni, S. A., Pook, M. A. & Evans-Galea, M. The emerging role of 5-hydroxymethylcytosine in neurodegenerative diseases. Front. Neurosci. 8, 397 (2014).
Chen, Y. et al. Discovery of cross-reactive probes and polymorphic CpGs in the Illumina Infinium HumanMethylation450 microarray. Epigenetics 8, 203–209 (2013).
Zhou, W., Laird, P. W. & Shen, H. Comprehensive characterization, annotation and innovative use of Infinium DNA methylation BeadChip probes. Nucleic Acids Res. 45, e22 (2017).
Fortin, J.-P., Triche, T. J., Hansen, K. D. & Hansen, K. D. Preprocessing, normalization and integration of the Illumina HumanMethylationEPIC array with minfi. Bioinformatics 33, 558–560 (2017).
Guintivano, J., Aryee, M. J. & Kaminsky, Z. A. A cell epigenotype specific model for the correction of brain cellular heterogeneity bias and its application to age, brain region and major depression. Epigenetics 8, 290–302 (2013).
Tian, Y. et al. ChAMP: updated methylation analysis pipeline for Illumina BeadChips. Bioinformatics 33, 3982–3984 (2017).
Jiao, C. et al. Positional effects revealed in Illumina methylation array and the impact on analysis. Epigenomics 10, 643–659 (2018).
Wahl, S. et al. On the potential of models for location and scale for genome-wide DNA methylation data. BMC Bioinform. 15, 232 (2014).
Breheny, P., Stromberg, A. & Lambert, J. p-value histograms: inference and diagnostics. High-Throughput 7, 23 (2018).
Peters, T. J. et al. De novo identification of differentially methylated regions in the human genome. Epigenetics Chromatin 8, 6 (2015).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B (Methodol.) 57, 289–300 (1995).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284 (2012).
Bernstein, A. I. & Kochmanski, J. DNA modifications in human PD. Open Science Framework https://osf.io/z4vbw (2021).
Acknowledgements
This work was supported by the National Institute of Environmental Health Sciences of the National Institutes of Health under award R00 ES024570 (A.I.B.). The Banner Sun Health Research Institute Brain and Body Donation Program has been supported by the National Institute of Neurological Disorders and Stroke (U24 NS072026 National Brain and Tissue Resource for Parkinson’s Disease and Related Disorders), the National Institute on Aging (P30 AG19610 Arizona Alzheimer’s Disease Core Center), the Arizona Department of Health Services (contract 211002, Arizona Alzheimer’s Research Center), the Arizona Biomedical Research Commission (contracts 4001, 0011, 05-901 and 1001 to the Arizona Parkinson’s Disease Consortium) and the Michael J. Fox Foundation for Parkinson’s Research. The authors thank the Van Andel Genomics Core, especially Marie Adams, for providing consultation, library preparation, and next-generation sequencing facilities and services (Illumina EPIC array). We also thank the Van Andel Institute Flow Cytometry Core, especially Rachael Sheridan and Joshua L. Schipper, for their assistance with flow cytometry. Lastly, we are grateful to the Banner Sun Health Research Institute Brain and Body Donation Program of Sun City, Arizona for the provision of human brain tissue.
Author information
Authors and Affiliations
Contributions
A.I.B.: conceptualization, funding acquisition, methodology, project administration, supervision, writing—review and editing. J.K.: conceptualization, data curation, formal analysis, investigation, methodology, visualization, writing—original draft, writing—review and editing. N.K.: conceptualization, investigation, methodology. All authors read and approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kochmanski, J., Kuhn, N.C. & Bernstein, A.I. Parkinson’s disease-associated, sex-specific changes in DNA methylation at PARK7 (DJ-1), SLC17A6 (VGLUT2), PTPRN2 (IA-2β), and NR4A2 (NURR1) in cortical neurons. npj Parkinsons Dis. 8, 120 (2022). https://doi.org/10.1038/s41531-022-00355-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41531-022-00355-2
This article is cited by
-
Genetic variation and pesticide exposure influence blood DNA methylation signatures in females with early-stage Parkinson’s disease
npj Parkinson's Disease (2024)
-
Linking environmental risk factors with epigenetic mechanisms in Parkinson’s disease
npj Parkinson's Disease (2023)
-
Epigenome-wide association study of peripheral immune cell populations in Parkinson’s disease
npj Parkinson's Disease (2023)