Abstract
The molecular mechanisms involved in myogenic differentiation are relatively well-known. Myogenic differentiation is regulated by the sequential activation of the basic helix-loop-helix myogenic regulatory transcription factors (MRFs), and biomechanical signals play an important role in the regulation of myogenesis. In this study, we sought to determine whether simulated microgravity culture using Gravite® may affect myoblast differentiation and expression of MRF genes. Although rat myoblasts, L6 cells were differentiated to myotubes in an incubation period-dependent manner, myogenesis of L6 cells was significantly attenuated under simulated microgravity (10-3G) conditions. Real-time Reverse transcription polymerase chain reaction (RT-PCR) showed that expressions of Myog, Myf6, Mef2c, Des, and Ckm under 1 G conditions increase in an incubation period-dependent manner, and that Myod1 expression was specifically observed to increase transiently in the early phase. However, expressions of Myod1 and Myog were significantly inhibited under simulated microgravity conditions. To clarify the molecular mechanisms, L6 cells were treated with 5-AzaC, and further incubated with differentiation medium under 1 G or 10−3 G conditions. The results showed differences in expression levels of Myod1, Myog, and, as well as those of myotube thickness between 1 G and 10−3 G conditions, completely disappeared in this experimental condition. Modified HpaII tiny fragment enrichment by ligation-mediated PCR (HELP)-assay showed that kinetic changes of DNA methylation status were attenuated in simulated microgravity conditions. These results indicate that microgravity regulates myogenesis and Myod1 expression by controlling DNA methylation.
Similar content being viewed by others
Introduction
Sarcopenia is defined as an age-related loss of skeletal muscle mass and strength. Beginning with the 4th decade of life, and symptoms progress with age.1 If muscle mass accounts for up to 60% of body weight, pathological changes in skeletal muscle can cause serious effects on older adults. Nevertheless, the therapeutic outcome of age-related skeletal muscle atrophy and weakness remains unknown.2,3,4 The understanding of molecular mechanisms of myogenic differentiation process will result in better treatment outcomes, since impaired regulation of myogenic differentiation is closely associated with age-related skeletal muscle dysfunction.5,6,7,8 It is well known that myogenic differentiation is regulated by the sequential activation of the basic helix-loop-helix myogenic regulatory transcription factors (MRFs): MyoD, Myf5, myogenin, and MRF4 (Myf6).9,10,11 MyoD in particular, is involved in the commitment of cells to the myogenic lineage.12,13,14 It has been reported that exposure of myoblast or muscle satellite cells to mechanical uniaxial stretch and stretching by a magnetic field, or electrical stimulation, upregulate MRFs expression, resulting in activation of myogenesis.15,16,17 Moreover, differentiation of myoblasts was enhanced on highly aligned fullerene whiskers scaffolds culture.18 In contrast, differentiation of myoblast was suppressed in the microgravity culture for in vitro study and spaceflight experiments, suggesting that biomechanical signals play an important role in the regulation of myogenesis.19,20 It is significant that the muscle mass of astronauts after 2 weeks space flight was diminished by up to 20%.21
DNA methylation is also known to be involved in regulation of myogenic gene expression and myogenesis: The DNA-demethylating agent 5-azacytidine induced myogenesis via upregulated MyoD expression.22,23,24,25 Barrès et al. interestingly reported that acute exercise tends to reduce global methylation on the entire genome, resulting in activation of regulated genes in human skeletal muscle.26 Singh et al. demonstrated that simulated microgravity induced epigenetic changes of genome DNA in human lymphocytes through altered expression of DNMT1, DNMT3a, and DNMT3b.27 However, the effects of space flight or simulated microgravity on epigenetic changes of MRF genes and resulting myogenesis remain unclear. Therefore, this study aims to determine whether simulated microgravity affects myoblast differentiation, expression of MRF genes, and the status of DNA methylation.
Results
Morphological evaluation of myogenesis under microgravity conditions
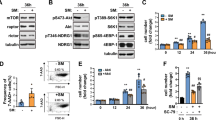
To evaluate myogenesis under simulated microgravity conditions, L6 cells were cultured with differentiation medium under normal 1 G or microgravity (10-3G) conditions for 1, 3, 5, or 7 days (Fig. 1a). Cells were then observed by capturing images, and the maximum transverse diameters of myotube cells (myotube thickness) were measured as indicated in Fig. 1b. As expected, L6 cells were differentiated to myotubes, resulting in an increase in the thickness, under 1 G conditions in an incubation period-dependent manner (Fig. 1c). Interestingly, myogenesis was significantly attenuated under microgravity conditions on day 3, 5, and 7, although myotubes gradually increased (Fig. 1c).
Morphological evaluation of myogenesis under microgravity conditions. a Schematic representation of experimental time course is shown. L6 cells were cultured with growth medium for 2 days, and then replaced with differentiation medium. Cells were then cultured under normal 1 G or 10-3G conditions for 0, 12 h, 1, 3, 5, or 7 days, and observed by taking pictures, and the maximum transverse diameters of myotube cells (myotube thickness) were measured. b Representative microscopic images during myogenesis are shown. White arrow heads: myotube cell; white H-shaped bars: examples of the maximum transverse diameter of myotube cells; white scale bar: 300 μm. c The maximum transverse diameters of myotube cells were measured by image processing software ImageJ, and the mean value of 10 fields was calculated. Columns show the mean of three independent experiments; bars, SD. P values calculated with Student’s t test. (**P < 0.01) (n = 3)
Gene expression analyses of myogenesis-relating genes under microgravity conditions
In order to clarify the molecular mechanisms of attenuation of myogenesis under microgravity conditions, expression levels of myogenesis-related genes were evaluated using real-time RT-PCR (Fig. 2): Expression of Myog, Myf6, Mef2c, Des, and Ckm under 1 G conditions increased in an incubation period-dependent manner. Although transient increases of Myod1 expression under 1 G conditions were observed from 12 h to Day1, those increases were significantly inhibited in microgravity conditions. Myog expressions were also significantly inhibited in microgravity conditions at the same points of time. Expression levels of Pax3, Pax7, and Myf5 did not change under our study conditions.
Gene expression analyses of myogenesis-related genes under microgravity conditions. For expression analysis, L6 cells were cultured under normal 1 G or 10-3G conditions for 0, 12 h, 1, 3, 5, or 7 days. Total RNA was extracted from cell pellets, and expression levels of myogenesis-relating genes were evaluated by real-time RT-PCR. Three independent measurements were averaged, and relative gene expression levels were calculated as a ratio against Actb expression for each experiment. Columns show the mean of three independent experiments; bars, SD. P values calculated with Student’s t test. (*P < 0.05, **P < 0.01) (n = 3)
Effects of 5-Azacytidine treatment on myogenesis under microgravity conditions
Expression of Myod1 and Myog had previously been reported to be regulated by DNA methylation.23,25,28 In order to clarify the molecular mechanisms of the decreased expression of Myod1 and Myog, and the significance in myogenesis under microgravity conditions, L6 cells were then treated with DNA methylation inhibitor, 5-Azacytidine (5-AzaC), before incubation with differentiation medium (Fig. 3a). To optimize concentration and treatment periods of 5-AzaC, L6 cells were first treated with 1, 2, 5, or 10 μΜ of 5-AzaC for 4, 8, or 12 days. As a result, Myod1 expression was observed to increase in a concentration- or treatment period-dependent manner (Fig. 3b). Of these conditions, treatment with 5 μΜ of 5-AzaC for 12 days was confirmed as the effective condition for the following experiments. After treatment with 5 μΜ of 5-AzaC for 12 days, L6 cells were incubated with growth medium for 2 days, and also with differentiation medium for the indicated periods under 1 G or microgravity conditions. Real-time RT-PCR showed that the expression levels of Myod1 and Myog seemed to increase gradually in a treatment period-dependent manner, but the difference between 1 G and 10−3 G conditions observed in Fig. 2, and disappeared in Fig. 3c. In this experimental condition, morphological evaluations were also performed as in Fig. 1. Results showed that differences in myotube thickness between 1 G and microgravity conditions also completely disappeared, although L6 cells were gradually differentiated to myotubes under both conditions (Fig. 3d,e).
Effects of 5-Azacytidine treatment on myogenesis under microgravity conditions. a Schematic representation of experimental time course is shown. L6 cells were cultured with 5 μM of 5-AzaC in growth medium for 12 days, with passage every 4 days. Cells were then seeded on culture flasks and cultured in growth medium for 2 days. After medium replacement with differentiation medium, cells were cultured in normal 1 G or 10−3 G conditions for 0, 12 h, 1, 3, 5, and 7 days. b Optimization of 5-azaC treatment was performed. L6 cells were treated with 1, 2, 5, or 10 μM of 5-AzaC for 4, 8, or 12 days, and expression levels of Myod1 gene were evaluated by real-time RT-PCR. Three independent measurements were averaged and relative gene expression levels were calculated as a ratio against Actb expression for each experiment. Columns show the mean of three independent experiments; bars, SD (n = 3). c Expression levels of Myod1 and Myog genes were evaluated by real-time RT-PCR. Three independent measurements were averaged and relative gene expression levels were calculated as a ratio against Actb expression for each experiment. Columns show the mean of three independent experiments; bars, SD (n = 3). d Representative microscopic images during myogenesis are also shown. White arrow heads: myotube cell; white H-shaped bars: examples of the maximum transverse diameter of myotube cells; white scale bar: 300 μm. e The maximum transverse diameters of myotube cells were measured using image processing software ImageJ, and the mean value of 10 fields was calculated. Columns show the mean of three independent experiments; bars, SD. (n = 3)
DNA methylation in Myod1 gene under microgravity conditions
To determine whether DNA methylation actually contributes to regulation of Myod1 expression, we use modified HELP-assay to detect DNA methylation in the promoter region of Myod1 gene.29 UCSC Genome Browser (http://genome.ucsc.edu) showed that MYOD1 gene virtually overlapped with CpG island, indicating very dense CpG sites, and the proximal promoter region contained actually methylated as shown in the results of reduced representation bisulfite sequencing and methyl 450 K bead arrays (Fig. 4a). A primer set was designed for just upstream from the transcription start site of Myod1, which contained 12 CpG or 3 HpaII sites (Fig. 4b). Since DNA methylation sensitive restriction enzyme HpaII cannot digest methylated DNA, but/although can digest unmethylated DNA, the primer set amplified methylated DNA, but not unmethylated or digested DNA after incubation with HpaII. Therefore, methylated DNA can be quantified by real-time PCR, resulting in evaluation of DNA methylation status. Genomic DNAs were then isolated from L6 cell samples that had been cultured as in Fig. 1, and incubated with HpaII for 12 h. Real-time PCR demonstrated that relative amounts of PCR products amplified from 1 G samples decreased gradually until Day3, and then increased until Day7 (Fig. 4c). It is striking that PCR products from microgravity samples also slightly decreased but to a smaller degree than those of 1 G, suggesting a retaining DNA methylation on Myod1 promoter (Fig. 4c). In order to clarify the underlying mechanisms, real-time RT-PCR analyses of methylation-related genes were performed using the same samples of cDNA as in Fig. 2. Results showed that expressions of DNA methyltransferase, Dnmt1 and Dnmt3a, under 1 G conditions were decreased during the myogenic process (Fig. 4d), while Dnmt3b was not detected (data not shown). Interestingly, expressions of Dnmt1 and Dnmt3a under microgravity conditions also decreased during the myogenic process, but remained significantly higher than those under 1 G conditions, suggesting they retained functions of methyltransferases (Fig. 4d). Expression levels of Tdg also decreased during the myogenic process, but differences between 1 G and microgravity conditions were smaller. Expression levels of Mbd2 varied and there was no trend observed.
DNA methylation in Myod1 gene under microgravity conditions. a Schematic structure of MYOD1 gene in UCSC Genome Browser showed that MYOD1 gene virtually overlapped with CpG island (upper rectangle drawn in red dotted line), indicating very dense CpG sites, and the proximal promoter region contained actually methylated sites (lower rectangle drawn in red dotted line). b The nucleic acid sequence of proximal promoter region in Myod1 gene is shown. Blue circle and arrow indicate transcription start site of Myod1 gene, red rectangles, CpG sites, blue rectangles, HpaII sites, and red arrows, primer sets for modified HELP-assay. c Methylated proximal promoter regions in Myod1 gene are quantified by real-time PCR. Three independent measurements were averaged and relative amounts of PCR products were calculated as the ratio against that of Set0 for each experiment. (n = 3). d Expression levels of methylation related genes were evaluated by real-time RT-PCR. Three independent measurements were averaged and relative gene expression levels were calculated as a ratio against Actb expression for each experiment. Columns show the mean of three independent experiments; bars, SD. P values calculated with Student’s t test. (*P < 0.05, **P < 0.01) (n = 3)
Other regulation of Myod1 and Myog genes under microgravity conditions
In order to clarify regulation mechanisms of expression of Myod1 and Myog genes during the myogenic process under 1 G and microgravity conditions, further analyses of promoter activities and RNA stabilities were performed. First, the 1.47 kb DNA fragment of the Myod1 gene promoter region, or the 1.55 kb DNA fragment of the Myog gene promoter region, was amplified and subcloned into the luciferase reporter plasmid pGL4.16 (designated pGL4.16-Myod1 Pro1474 or pGL4.16-Myog Pro1546, Supplementary Fig 1B). L6 cells were then transfected with these promoter reporters and incubated for 1 day. Next, growth medium was replaced with differentiation medium and incubated for one more day under 1 G or microgravity conditions (Supplementary Fig 1A). Dual-luciferase assays demonstrated that cloned regions had strong promoter activities compared to empty vector, but any differences in those activities under 1 G or microgravity conditions were not observed (Supplementary Fig 1C). L6 cells were next cultured in growth medium for 2 days, and then cultured in differentiation medium for further a day. After adding 5 μg/ml of actinomycin D, cells were cultured under 1 G or microgravity conditions (Supplementary Fig 2A). Real-time RT-PCR showed that gene expression level of Myod1 had decreased gradually, while those of Myog and Actb had not (Supplementary Fig 2B). There were no differences in those levels under 1 G or microgravity conditions (Supplementary Fig 2B).
Discussion
As a consequence of an aging population, the number of elderly who exhibit skeletal muscle atrophy and weakness are increasing, becoming a concern of society.30,31,32 In order to achieve better treatment outcomes, the understanding of the molecular mechanisms of myogenesis is essential, since impaired myogenesis is closely associated with age-related skeletal muscle loss and weakness.5,6,7,8 Previous reports have suggested that biomechanical signals play an important role in the regulation of myogenesis. Specifically, exposure of myoblast and muscle satellite cells to mechanical stretching, electrical stimulation, or microgravity have been found to modify expression levels of MRFs, resulting in alteration of myogenesis.15,16,17,19
In this study, we used our new generation 3D-clinostat “Gravite®” to obtain simulated microgravity conditions (10-3G) for cell culture: We observed that myogenesis of rat myoblast cells (L6 cells) cultured in Gravite® were significantly attenuated, indicating successful creation of microgravity conditions (Fig. 1). L6 cells cultured in differentiation medium under 1 G conditions normally differentiated to myotubes in an incubation period-dependent manner. However, myogenesis of L6 cells was significantly attenuated under microgravity conditions from Day3 to Day7, although myotubes gradually increased, indicating a retardation of myogenesis under microgravity conditions (Fig. 1). Since myogenesis is regulated by the sequential activation of MRFs, expression levels of myogenesis-related genes were evaluated by real-time RT-PCR. Results showed expressions of Myod1, Myog, Myf6, Mef2c, Des, and Ckm under 1 G conditions were increased in an incubation period-dependent manner (Fig. 2). Among them, Myod1 expression under 1 G conditions showed a unique transient increase in the early phase (from 12 h to Day1), suggesting initiation of the myogenic process. And expressions of Myod1, as well as Myog were strongly inhibited under microgravity conditions, suggesting that those inhibitions might have directly affected initiation of myogenesis as in Fig. 1. These significant results suggest that our experimental model would be useful in analysing biological and molecular mechanisms under microgravity conditions.
To clarify the molecular mechanisms of decreased expression of Myod1 and Myog, and their significance in myogenesis under microgravity conditions, L6 cells were first treated with various doses of DNA methylation inhibitor (5-AzaC) for various incubation periods to optimize experimental conditions. As expected, Myod1 expression increased in a concentration- or a treatment period-dependent manner, although treatment with 10 μΜ of 5-AzaC showed slightly toxic effects. We thus decided to try treatment with 5 μΜ of 5-AzaC for 12 days as an effective condition for the following experiments. Real-time RT-PCR showed that expression levels of Myod1 and Myog seemed to increase gradually in a treatment period-dependent manner, but the transient increase of Myod1 in the early phase, seen in Fig. 2, disappeared. And differences in expression levels of Myod1 and Myog between 1 G and 10−3 G conditions also disappeared. Furthermore, differences in myotube thickness between 1 G and microgravity conditions completely disappeared in this experimental condition, suggesting that modifying DNA methylation status is essential to regulate myogenesis and expression of Myod1 and Myog under microgravity conditions (Fig. 3d,e). These results suggest that DNA methylation is a key factor in gravity-regulating myogenesis, and possibly a molecular target for treatment of skeletal muscle dysfunctions and Sarcopenia.
UCSC Genome Browser indicated that MYOD1 gene virtually overlapped with CpG island, indicating very dense CpG sites, and the proximal promoter region actually contained methylated sites (Fig. 4a). In fact, our modified HELP-assay showed that methylated Myod1 promoter had gradually decreased (PCR product decreased) until Day3, and then increased until Day7. These kinetic changes in DNA methylation status apparently contributed to increased expression of Myod1 in the initiation process of myogenesis. Furthermore, these kinetic changes in DNA methylation status were attenuated under microgravity conditions, indicating that microgravity regulated both myogenesis and Myod1 expression by controlling DNA methylation status. The mRNA expressions of Dnmt1 and Dnmt3a gave further evidence that expression levels in DNA methyltransferases might contribute to differences in DNA methylation status under microgravity conditions.
Singh et al. reported that simulated microgravity increased expression levels of DNMT1, DNMT3a, and DNMT3b at 72 h, and reduced them at 7 days in human T-lymphocyte cells. Also, the methylation sensitive-random amplified polymorphic DNA (MS-RAPD) analysis revealed that simulated microgravity exposure resulted in DNA hypomethylation and mutational changes.27 In our study, with similar kinetics, increased levels of Dnmt1 and Dnmt3a at 1-5 days and decreased levels at 7 days were observed, but DNA methylation status should have opposite effects: Hypermethylation resulted in decreased expression of Myod1 and retardation of myogenesis. Effects of Tdg kinetics were not consistent with the DNA methylation status. These differences might be due to differences in cell type, culture system, method of methylation detections, or target gene. Although our findings seem consistent, further investigation will be necessary to clarify the results. Other experiments to evaluate promoter activities and mRNA stabilities have also supported the importance of epigenetic regulation of Myod1 in myogenesis and MRF expression under altered gravity conditions.
In conclusion, we demonstrated that simulated microgravity attenuated myogenesis by controlling DNA methylation status of Myod1 (Fig. 5). It is notable that a DNA methylation inhibitor attenuates the inhibitory effects of microgravity on myogenesis, suggesting the inhibitors’ potential as a molecular targeting therapy for muscle atrophy and weakness. Although this study mainly focused on biological viewpoints, further analyses including the influence of external physico-chemical parameters will probably provide new insight into the molecular mechanisms of myogenesis, and also the development of molecular targeting therapy for age-related skeletal muscle dysfunction in the clinical phase.
Hypothetical model of regulation of myogenesis. Differentiation signals regulate DNA methylation machinery, and especially, the Myod1 gene in myoblast cells, resulting in sequential activation of target genes and consecutive myogenic differentiation. Simulated microgravity attenuated this myogenesis by controlling DNA methylation status of Myod1. This signal pathway was also controlled using DNA methylation inhibitor, 5-AzaC, treatment
Materials and methods
Cell culture
Rat myoblast cells, L6 cells, were obtained from Health Science Research Resources Bank (Osaka, Japan) and maintained with Dulbecco’s modified Eagle’s minimal essential medium-High glucose (DMEM-H) (NACALAI TESQUE, Inc., Kyoto, Japan) containing 10% fetal bovine serum (FBS; BioWhittaker, Verviers, Belgium), 100 U/ml penicillin, and 0.1 mg/ml streptomycin (Sigma, St. Louis, MO, USA) as previously described.17 Cells were cultured with growth medium for 2 days, and then replaced with differentiation medium containing DMEM-H, 2% FBS, 100 U/ml penicillin, and 0.1 mg/ml streptomycin after reaching sub-confluence. Cells were then cultured under normal 1 G or 10-3G conditions, using “Gravite®” (Space Bio-Laboratories Co., Ltd., Hiroshima, Japan).
For expression analysis, cells were cultured under normal 1 G (group 1 G) or 10-3 G conditions (group 10−3G) for 0, 12 h, 1, 3, 5, or 7 days. Cells were then harvested and stored at −80˚C until use. L6 cells were also cultured with 5 μM of 5-Azacytidine (5-AzaC, Wako Pure Chemical Industries, Ltd., Osaka, Japan) in growth medium for 12 days, with passage every 4 days. Cells were then seeded on 12.5 cm2 culture flasks and cultured in growth medium for 2 days. After medium replacement with differentiation medium, cells were cultured in normal 1 G or 10−3 G conditions. Morphological observations and harvesting were performed out at 0, 12 h, 1, 3, 5, and 7 days, and stored at −80 °C until use. Methods were performed in accordance with relevant regulations and guideline.
Gravite®
Microgravity conditions can be produced either by space flight or by free fall; to simulate microgravity, we used a newly developed Gravite® (Space Bio-Laboratories Co., Ltd.), as previously patented (undifferentiated pluripotent stem cell proliferation/differentiation regulation method and system, Patent No. 8034616B2 (US), 2515552 (CA), 1577380 (EPC: GB, FR, DE, IT, SE), ZL02830112.9 (CN), and O731940 (KR), and GRAVITY CONTROLLER, Patent No. 623009 (JP), US9494949B2 (US) and granted in EU). This device produces an environment similar to that of outer space (10−3 G) by rotating a sample around two axes, integrating the gravity vector with the temporal axis. This is accomplished by rotation of a chamber at the center of the device, resulting in uniform dispersion of the gravity vector within a spherical volume, with a constant angular velocity. These specific conditions produced a simulated environment of 10−3 G in 8 minutes actually measured by gravity acceleration sensor, and it was defined as simulated microgravity (10−3 G).33,34,35
Analysis of myogenesis
To evaluate myogenesis, cells were observed by taking pictures of 10 fields using inverted phase contrast microscope (Eclipse, Nikon, Japan) after incubation under 1 G or 10−3 G conditions for indicated periods. The maximum transverse diameters of myotube cells were measured by image processing software ImageJ, and the mean value of 10 fields was calculated.
RNA preparation and real-time RT-PCR
Total RNA was extracted from frozen cell pellets using the NucleoSpin® RNA II kit (MACHEREY-NAGEL, Düren, Germany) according to manufacturer instructions. Two micrograms of total RNA extracted from each cell line were reverse-transcribed using the High-Capacity cDNA Archive™ Kit (Applied Biosystems, Foster City, CA, USA). A 1/200 dilution of the cDNA was subjected to real-time RT-PCR using primers (final concentration of 200 nM each) and MGB probe (final concentration of 100 nM, the Universal Probe Library: UPL, Roche Diagnostics, Tokyo, Japan) (shown in S1 Table) sets with FastStart Universal Probe Master (ROX) (Roche Diagnostics) for quantitation of gene expressions with Actb as an internal housekeeping control. PCR reactions were carried out using 7500 Real-Time PCR System (Applied Biosystems) under the following standard conditions: Three independent measurements were averaged and relative gene expression levels were calculated as a ratio against Actb expression for each experiment.
DNA extraction and methylation analysis
Genomic DNA was isolated from frozen cell pellets using the NucleoSpin® Tissue (MACHEREY-NAGEL) according to manufacturer instructions. Fifty ng of genomic DNA in total 50 μl reaction mixture were digested with methylation-sensitive HpaII at 37 °C for 12 h, and 2 μl of reaction mixture were then subjected to real-time PCR using primer set (shown in S2 Table) to amplify a fragment including CpG sites in Myod1 promoter region with iTaq® SYBR™ Green Supermix with ROX (BIO-RAD, Hercules, CA, USA). PCR reactions were carried out using 7500 Real-Time PCR System (Applied Biosystems) under the following standard conditions: Three independent measurements were averaged and relative gene expression levels were calculated as a ratio to sample DNA at Set0.
Plasmid construction and luciferase reporter experiments
The 1.47 kb DNA fragment of the Myod1 gene (−1264 to +210 from the transcriptional start site at +1) or the 1.55 kb DNA fragment of the Myog gene (−1512 to +34 from the transcriptional start site at +1) were amplified by PCR (primer sets are shown in S3 Table) from L6 genomic DNA, and they were subcloned into the luciferase reporter plasmid pGL4.16 (Promega, Madison, WI, USA) (designated pGL4.16-Myod1 Pro1474 or pGL4.16-Myog Pro1546). L6 cells were seeded on culture flasks, and cultured in growth medium for a day. Then reporter constructs were transiently transfected into L6 cells using TransIT®-LT1 Transfection Reagent (TaKaRa Bio, Inc., Shiga, Japan). The renilla luciferase vector (pRL-SV40, Promega) was used as a transfection efficiency control. After incubation for a day, growth medium was replaced with differentiation medium and incubated for one more day under 1 G or 10-3G conditions. Luciferase luminescence was measured using a single-sample luminometer, the Biolumat LB 9505 (BERTHOLD TECHNOLOGIES GmbH & Co. KG, Bad Wildbad, Germany) with the Dual-Luciferase Reporter Assay System (Promega). Promoter activities were calculated as the ratio of firefly to renilla luciferase readings, and the averages of at least three independent experiments were calculated. Methods were approved by a Hiroshima University Gene Recombination Experiment Safety Committee.
Analyses of mRNA stabilities
L6 cells were seeded on culture flasks, and cultured in growth medium. After 2 days, growth medium was replaced with differentiation medium and incubated for another day. After adding 5 μg/ml of actinomycin D (NACALAI TESQUE) to the medium, cells were cultured under 1 G or 10−3 G conditions for the indicated periods (0, 15, 30, 45, 60, or 120 min). Total RNA was prepared from each harvested cell, and gene expressions of Myod1, Myog, and Actb were analysed using real-time RT-PCR method as mentioned above.
Statistical analysis
All statistical tests were performed using the StatView® version 5.0 software (SAS Institute Inc., NC, USA) and Microsoft® Excel® for Mac version 12.3.6. Student’s t-test was used to determine P-values (*P < 0.05, **P < 0.01).
Data availability
All relevant data are available from the corresponding author.
Change history
18 July 2018
The original version of this Article contained an error in Fig. 4. Fig. 4B was partly cropped, which obscured some of the sequence information. This has been corrected in both the PDF and HTML versions of the Article.
References
Metter, E. J., Conwit, R., Tobin, J. & Fozard, J. L. age-associated loss of power and strength in the upper extremities in women and men. J. Gerontol. Biol. Sci. 52, 267–276 (1997).
Iolascon, G. et al. Physical exercise and sarcopenia in older people: position paper of the Italian Society of Orthopaedics and Medicine (OrtoMed). Clin. Cases Miner. Bone Metab. 11, 215–221 (2014).
Law, T. D., Clark, L. A. & Clark, B. C. Resistance exercise to prevent and manage sarcopenia and dynapenia. Annu Rev. Gerontol. Geriatr. 36, 205–228 (2016).
Bosaeus, I. & Rothenberg, E. Nutrition and physical activity for the prevention and treatment of age-related sarcopenia. Proc. Nutr. Soc. 75, 174–180 (2016).
Degens, H. Age-related skeletal muscle dysfunction: causes and mechanisms. J. Musculoskelet. Neuron. Interact. 7, 246–252 (2007).
Pietrangelo, T. et al. Molecular basis of the myogenic profile of aged human skeletal muscle satellite cells during differentiation. Exp. Gerontol. 44, 523–531 (2009).
Lang, T. et al. Sarcopenia: etiology, clinical consequences, intervention, and assessment. Osteoporos. Int. 21, 543–559 (2010).
Sakuma, K., Aoi, W. & Yamaguchi, A. Molecular mechanism of sarcopenia and cachexia: recent research advances. Pflug. Arch. 469, 573–591 (2017).
Molkentin, J. D. & Olson, E. N. Defining the regulatory networks for muscle development. Curr. Opin. Genet Dev. 6, 445–453 (1996).
Sabourin, L. A. & Rudnicki, M. A. Developmental biology: frontiers for clinical genetics the molecular regulation of myogenesis. Clin. Genet 57, 16–25 (2000).
Berkes, C. A. & Tapscott, S. J. MyoD and the transcriptional control of myogenesis. Semin. Cell Dev. Biol. 16, 585–595 (2005).
Davis, R. L., Weintraub, H. & Lassar, A. B. Expression of a single transfected cDNA converts fibmblasts to myoblasts. Cell 51, 987–1000 (1987).
Weintraub, H. et al. Activation of muscle-specific genes in pigment, nerve, fat, liver, and fibroblast cell lines by forced expression of MyoD (muscle regulatory gene/MyoD retrovirus). Dev. Biol. 86, 5434–5438 (1989).
Choi, J. et al. MyoD converts primary dermal fibroblasts, chondroblasts, smooth muscle, and retinal pigmented epithelial cells into striated mononucleated myoblasts and multinucleated myotubes (myogenesis/myoflbrils/desmin/master switch genes). Dev. Biol. 87, 7988–7992 (1990).
Chandran, R., Knobloch, T. J., Anghelina, M. & Agarwal, S. Biomechanical signals upregulate myogenic gene induction in the presence or absence of inflammation. Am. J. Physiol. Cell Physiol. 293, 267–276 (2007).
Yuge, L. & Kataoka, K. Differentiation of myoblasts is accelerated in culture in a magnetic field. Vitr. Cell. Dev. Biol. -Anim. 36, 383–386 (2000).
Kawahara, Y. et al. Novel electrical stimulation sets the cultured myoblast contractile function to ‘on’. Pathobiology 73, 288–294 (2006).
Minami, K. et al. Highly ordered 1D fullerene crystals for concurrent control of macroscopic cellular orientation and differentiation toward large-scale tissue engineering. Adv. Mater. 27, 4020–4026 (2015).
Torgan, C. E., Burge, I. S. S., Collinsworth, I. A. M., Truskey, G. A. & Kraus, W. E. Differentiation of mammalian skeletal muscle cells cultured on microcarrier beads in a rotating cell culture system. Cell. Eng. 38, 583–590 (2000).
Kulesh, D. A. et al. Space shuttle flight (STS-45) of L8 myoblast cells results in the isolation of a nonfusing cell line variant. J. Cell Biochem 55, 530–544 (1994).
Clement, G. Musculo-skeletal system in space. Fundamentals of space medicine. (Kluwer Academic Publisher, Dordrecht, Netherlands, 2003).
Laker, R. C. & Ryall, J. G. DNA methylation in skeletal muscle stem cell specification, proliferation, and differentiation. Stem Cells Int. 2016, 5725927 (2016).
Montesano, A., Luzi, L., Senesi, P. & Terruzzi, I. Modulation of Cell Cycle Progression by 5-Azacytidine Is Associated with Early Myogenesis Induction in Murine Myoblasts. Int J. Biol. Sci. 9, 391–402 (2013).
Zhang, F., Pomerantz, J. H., Sen, G., Palermo, A. T. & Blau, H. M. Active tissue-specific DNA demethylation conferred by somatic cell nuclei in stable heterokaryons. Proc. Natl Acad. Sci. 104, 4395–4400 (2007).
Taylor, S. M. & Jones, P. A. Multiple new phenotypes induced cells treated with SAzacytidine in 10T 1/2 and 3T3 cells treated with 5-Azacytidine. Cell 17, 771–779 (1979).
Barrès, R. et al. Acute exercise remodels promoter methylation in human skeletal muscle. Cell Metab. 15, 405–411 (2012).
Singh, K. P., Kumari, R. & DuMond, J. W. Simulated microgravity-induced epigenetic changes in human lymphocytes. J. Cell. Biochem. 111, 123–129 (2010).
Zhong, X. et al. Ten-Eleven Translocation-2 (Tet2) is involved in myogenic differentiation of skeletal myoblast cells in vitro. Sci. Rep. 8, 43539 (2017).
Khulan, B. et al. Comparative isoschizomer profiling of cytosine methylation: The HELP assay. Genome Res. 16, 1046–1055 (2006).
Greenlund, L. J. S. & Nair, K. S. Sarcopenia–consequences, mechanisms, and potential therapies. Mech. Ageing Dev. 124, 287–299 (2003).
Cruz-Jentoft, A. J. et al. Sarcopenia: European consensus on definition and diagnosis Report of the European Working Group on Sarcopenia in Older People. Age Ageing 39, 412–423 (2010).
Cooper, C. et al. Frailty and sarcopenia: definitions and outcome parameters. Osteoporos. Int. 23, 1839–1848 (2012).
Mitsuhara, T. et al. Simulated microgravity facilitates cell migration and neuroprotection after bone marrow stromal cell transplantation in spinal cord injury. Stem Cell Res. Ther. 4, 35 (2013).
Kawahara, Y., Manabe, T., Matsumoto, M., Kajiume, T. & Yuge, L. LIF-free embryonic stem cell culture in simulated microgravity. PLoS One 4, e6343 (2009).
Yuge, L. et al. Simulated microgravity maintains the undifferentiated state and enhances the neural repair potential of bone marrow stromal cells. Stem Cells Dev. 20, 893–900 (2011).
Acknowledgements
We thank Prof. N. Hirohashi for his encouragement and Ms. C. Oda for her technical support. A part of this work was carried out at the Analysis Center of Life Science at Hiroshima University.
Author information
Authors and Affiliations
Contributions
Conceptualization: K.T., L.Y.; Data curation: T.F., K.T., T.F.; Formal analysis: T.F., K.T., T.F.; Funding acquisition: K.T., Y.K., L.Y.; Investigation: T.F., K.T., T.F.; Methodology: T.F., K.T., T.F., T.I., Y.K.; Project administration: K.T.; Resources: K.T., Y.K., L.Y.; Supervision: K.T. L.Y.; Writing–original draft: T.F., K.T., T.F., T.I.; Writing–review and editing: K.T. and L.Y.
Corresponding authors
Ethics declarations
Competing interests
L.Y. is a director of Space Bio-laboratories Co., Ltd. (SBL) and K.Y. is a president of SBL. They share holding. The interest conflicts of this research have been approved by the Conflict of Interest Management Committee. By regularly reporting of the research progress to the Conflicts of Interest Management Committee, we will maintain fairness regarding the interests of this research. The remaining authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Furukawa, T., Tanimoto, K., Fukazawa, T. et al. Simulated microgravity attenuates myogenic differentiation via epigenetic regulations. npj Microgravity 4, 11 (2018). https://doi.org/10.1038/s41526-018-0045-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41526-018-0045-0
This article is cited by
-
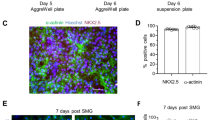
Simulated microgravity attenuates myogenesis and contractile function of 3D engineered skeletal muscle tissues
npj Microgravity (2024)
-
VDR regulates simulated microgravity-induced atrophy in C2C12 myotubes
Scientific Reports (2022)
-
Update on the effects of microgravity on the musculoskeletal system
npj Microgravity (2021)