Abstract
Neuro-ichthyotic syndromes are a group of rare genetic diseases mainly associated with perturbations in lipid metabolism, intracellular vesicle trafficking, or glycoprotein synthesis. Here, we report a patient with a neuro-ichthyotic syndrome associated with deleterious mutations in the ALDH1L2 (aldehyde dehydrogenase 1 family member L2) gene encoding for mitochondrial 10-formyltetrahydrofolate dehydrogenase. Using fibroblast culture established from the ALDH1L2-deficient patient, we demonstrated that the enzyme loss impaired mitochondrial function affecting both mitochondrial morphology and the pool of metabolites relevant to β-oxidation of fatty acids. Cells lacking the enzyme had distorted mitochondria, accumulated acylcarnitine derivatives and Krebs cycle intermediates, and had lower ATP and increased ADP/AMP indicative of a low energy index. Re-expression of functional ALDH1L2 enzyme in deficient cells restored the mitochondrial morphology and the metabolic profile of fibroblasts from healthy individuals. Our study underscores the role of ALDH1L2 in the maintenance of mitochondrial integrity and energy balance of the cell, and suggests the loss of the enzyme as the cause of neuro-cutaneous disease.
Similar content being viewed by others
Introduction
Neuro-ichthyotic syndromes are a group of rare genetic diseases mainly associated with perturbations in lipid metabolism, intracellular vesicle trafficking, or glycoprotein synthesis.1 Congenital dry and scaly skin and progressive neurological symptoms are hallmarks of this group of diseases. Sjögren–Larsson syndrome (SLS: MIM#270200) is one of the most recognized neuro-ichthyotic syndromes characterized by congenital ichthyosis, leukoencephalopathy, intellectual disability, and spastic di- or tetraplegia.2,3,4 In 95% of patients it is caused by mutations of the ALDH3A2 gene which encodes for the fatty aldehyde dehydrogenase (FALDH), a microsomal enzyme that oxidizes long-chain aldehydes to fatty acids.2,5,6 SLS patients without mutations in the ALDH3A2 gene have also been identified, leaving the cause of the symptoms unknown.7 Here, we report a patient with a congenital neuro-ichthyotic syndrome but atypical phenotype displaying dysmorphic features, and abnormalities on MRI and MR (1H-MRS) spectroscopy in the absence of ALDH3A2 gene mutations and no spastic paraplegia to suggest classic SLS. The diagnosis of Coffin–Lowry syndrome, made after identification of a deleterious frameshift mutation in the RPS6KA3 gene,8,9 does not explain all features of the patient. We provide evidence that the neuro-ichthyotic syndrome in this case is associated with the loss of expression of the ALDH1L2 gene, which encodes a mitochondrial folate enzyme.
Results
Patient’s developmental history and morphological features
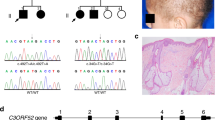
The patient (male) presented at birth with hypotonia, abnormally thick fingers and toes, and ichthyosis. Pruritis and facial dysmorphism were apparent since the age of 11 months (Fig. 1). He had normal statural, ponderal, and head circumference growth. Motor acquisitions during the 1st year were severely delayed with sitting acquired at 11 months and independent walking at 3 years. At 14 years, the patient was able to speak using efficient isolated words and had good communication skills. He was unable to read and presented hyperactivity and attention deficit increasing over time. To date, the patient had normal walking and has not developed neurological signs including spastic paraplegia or ataxia. Morphological studies of skeleton, abdomen, and heart were normal. Electroencephalogram showed diffuse moderate bradyrythmia. Somatosensory evoked potentials revealed prolonged latencies on the four limbs and motor evoked potentials showed the lack of cortical response. MRI at 1.5 T demonstrated early diffuse hypomyelination with coalescing and dilated Virchow–Robin spaces. 1H-MRS reveals two abnormal lipid peaks in the white matter persistent at short and long echo times (other peaks were normal, except a small increase of inositol) (Fig. 1). Positions of these peaks in our patient correspond to positions of the characteristic peaks in classic SLS but the overall profile was different. Specifically, in SLS the 1.3 ppm peak is more prominent while the 0.9 ppm peak is smaller (Fig. 1m). In our patient, the peaks were still evident at 6 years of age, the finding which in association with the leukodystrophy initially suggested the SLS diagnosis. Of note, in SLS patients both peaks are permanent through age while in our patient these peaks decreased over time and were not detected after 8 years of age (Supplementary Fig. 1).
Particularities of the patient phenotype. Patient at the age of 3 years a, b and 14 years c presented with a facial dysmorphism with epicanthus, hypertelorism, broad nasal root, anteverted nares, long philtrum, thin upper lip. The written consent for publication of these photos was obtained from child’s parents. Cerebral MRI shows diffuse hypomyelination at the age of 2 years with white matter appearing respectively in hypersignal on T2-weighted sequences, in hypersignal on FLAIR sequences and in normosignal on T1-weighted sequences d, g, j. Progressive myelination and dilatation and coalescing of Virchow–Robin spaces at the age of 6 e, h, k and 14 years f, i, l. 1H-MRS in the corona radiata for classical SLS patients shows a typical major peak at 1.3 ppm and a smaller peak at 0.9 ppm (m, arrows). 1H-MRS for our patient reveals a similar pattern with two peaks at 1.3 and 0.9 ppm (arrows) at the age of 2 n and 6 years o. However, the peak at 1.3 ppm appears smaller than the peak at 0.9 ppm in our patient. We observed no decrease in N-acetyl-aspartate (NAA)/creatine (Cr) ratio, or in choline (Cho) peak suggesting normal maintenance of neuronal and myelin content but a small increase of inositol peak that may be due to some astrocytic stress. Family pedigrees p and patient’s genotype r, s
Mutations identified by whole exome sequencing
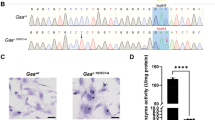
Sequencing of the ALDH3A2 exons, exon/intron junctions, and the full-length cDNA did not reveal any mutations in this gene in our patient. The following mutations were found by the whole exome analysis: (1) a de novo hemizygous mutation (c.263dup, p.Ser89Leufs*4) in the RPS6KA3 gene on the X chromosome,10,11 a deleterious frameshift mutation not carried by the patient’s mother. This finding confirmed that the patient was affected by a Coffin–Lowry syndrome8 and partially explained the patient’s features but did not explain the congenital pruritic ichthyosis, MRI or 1H-MRS features typical of SLS. (2) This patient also has compound heterozygous mutations in the ALDH1L2 gene, which encodes for a mitochondrial 10-formyltetrahydrofolate dehydrogenase.12 Discovered mutations, one intronic near splice site (c.2517-5C>T) and one frame shift (c.827del/p.Val276Glyfs*33; rs770401066, dbSNP NCBI) were not present in the homozygous state in ExAC or gnomAD. The patient’s asymptomatic parents and brother were heterozygous for one of these mutations. Segregation analysis revealed that the father harbors the Val276Glyfs*33 frameshift mutation, while the mother harbors the c.2517-5C>T intronic mutation. The presence of a mutant mRNA resulting from the frame shift was confirmed in patient’s fibroblasts by the direct sequencing. The mutated sequence predicts a truncated protein of 307 aa, including the 22 aa mitochondrial leader sequence, 253 aa of the N-terminal folate-binding domain/hydrolase catalytic center,13 and the 32 aa random peptide with no identity to known proteins resulted from the frameshift (Supplementary Fig. 2). Such truncated proteins are usually not folded properly,13 and apparently are rapidly degraded. Indeed, the truncated protein was not detected in the patient’s fibroblasts. Deficiency of the ALDH1L2 gene has not been reported and the overall consequences of the enzyme loss for the cell are not clear. We examined fibroblasts from this patient, his parents and healthy unrelated individual, and present evidence that the loss of ALDH1L2 impairs the mitochondrial function and is the likely cause of a new neuro-ichthyotic syndrome.
Characterization of patient’s fibroblasts
Compared to fibroblasts from a healthy individual (control, C cells), patient’s fibroblasts (R cells) have barely detectable ALDH1L2 protein (Fig. 2a–c and Supplementary Fig. 3). Levels of ALDH1L2 mRNA were also significantly lower in patient’s cells (Fig. 2d and Supplementary Fig. 4). Since one of the alleles of the ALDH1L2 gene in the patient has mutation near the splice site, we attributed the decrease in the mRNA level to the impaired transcription caused by the mutation. Indeed, splice site mutations are known to cause loss of gene expression.14 Levels of the ALDH3A2 protein were not different between the two cell lines (Fig. 2a and Supplementary Fig. 3), an indication that FALDH deficiency was not the primary cause of the patient’s symptoms. The ALDH1L2 enzyme catalyzes the conversion of 10-formyl-THF to THF and CO2 simultaneously producing NADPH from NADP+ (Fig. 2e).12,15,16 Therefore, the ALDH1L2 activity is likely to affect folate metabolism but the extent of the enzyme contribution to the maintenance of reduced folate pools is not clear. The total folate levels were not significantly different between patient’s (R) and control (C) fibroblasts (Fig. 2f), only 10-formyl-THF was noticeably and significantly different between two fibroblast cultures (Fig. 2f). ALDH1L1, the cytosolic homolog of ALDH1L2 and a major user of 10-formyl-THF17,18 was not present in either fibroblast culture (Supplementary Fig. 5). Therefore, the three-fold increase of this folate upon the ALDH1L2 loss (Fig. 2f) indicates that the enzyme is a major user of 10-formyl-THF. The ratio of NADPH/NADP+, metabolites also involved in ALDH1L2 catalysis, was more than four-fold lower in patient versus control fibroblasts (Fig. 2g and Supplementary Fig. 6), supporting the role of ALDH1L2 as the main source of NADPH generation.19 Furthermore, patient’s fibroblasts have much lower ATP levels in mitochondria as well as in whole cells (Fig. 2h and Supplementary Figs. 7 and 8) with the ATP/ADP ratio indicating a very low energy status in patient’s fibroblasts (Fig. 3b, d). Another characteristic feature of ALDH1L2-deficient fibroblasts is a decreased proliferation rate (Fig. 2i), which was not responsive to the increase of folate in media (10 μM leucovorin or 20 μM folic acid). In fact, the metabolomics analysis demonstrated differences between the patient and control fibroblasts beyond folate metabolism (Fig. 2j) with statistically significant (p < 0.05) differences for 250 out of 475 assigned metabolites.
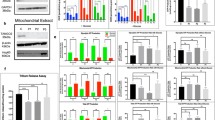
Difference between fibroblasts from the patient (R cells) and fibroblasts from healthy individual (C cells). a–c R cells have much lower levels of ALDH1L2 protein as evaluated by Western blot assay (a, ratios of averaged band intensities are indicated; statistics is shown in Supplementary Fig. 3) and confocal microscopy b in cells or by Western blot assay in isolated mitochondria (c). In panel a, samples were from different plates (biological replicates) with 20 μg of the total protein loaded per well. d Levels of ALDH1L2 mRNA are lower in R cells (mean ± SE of three biological replicates). e Cytosolic and mitochondrial folate pathways. f Levels of folate coenzymes (FA, folic acid; THF, tetrahydrofolate; 5-MTHF, 5-methyl-THF; 10-CHO-THF, 10-formyl-THF) in C and R cells (only 10-CHO-THF was noticeably and significantly different between the two cell lines). For each cell type mean ± SE of three independent experiments (each done in quadruplicate) is shown (3 biological replicates each includes 4 technical replicates). g Ratio of NADPH/NADP+ in C and R cells (mean ± SE of four biological replicates). h Levels of ATP in C and R cells (mean ± SE of four biological replicates). For panels d, g, h, p values were below 0.001 (Student’s t-test) for the comparison of R and C cells (detailed statistical analysis for these panels is shown in Supplementary Figs. 4 and 6–8). i Proliferation rate of C and R cells measured in real-time (xCelligence); samples with three densities of cells were monitored for each cell type. In each case, experiments were done in duplicate with automated averaging of data points. j PCA for metabolites (475 total) measured in C and R cells (n = 4; samples are biological replicates)
Patient fibroblasts have lower ATP and energy index, and altered mitochondrial morphology in comparison with fibroblasts from parents or SLS patient. a Western blots assays of ALDH1L2 and ALDH3A2 in fibroblasts isolated from the patient (R), both parents (mother, M; father, F) and from an SLS patient (SLS). LR denotes patient’s fibroblasts transduced for ALDH1L2 expression. Numbers on the bottom of each panel indicate band intensity (arbitrary units) for ALDH1L2 and ALDH3A2 relative to the intensity of corresponding actin band. b Doubling time and energy index of different fibroblast cultures (cell labeling as in panel a). c ATP levels measured by a colorimetric assay in different fibroblast cultures. Three different samples (biological replicates) were used in this experiment; for each sample, 4 measurements (technical replicates) were performed and the average of these measurements were used to calculate mean ± SE. d TMRM (tetramethylrhodamine) to MitoTracker Green ratio in different fibroblasts. Six samples (biological replicates) were analyzed for each cell type. e Levels of ROS evaluated by confocal microscopy after DCF (2′,7′-dichlorodihydrofluorescein diacetate) staining in patient (R cells) and control (C cells) fibroblasts (values were calculated from the analysis of 10 cells for each cell type; laser power was kept uniform for all measurements). f Confocal images (108×) of different fibroblast cultures (as in panel a). Live cells were stained with Hoechst (nucleus staining, light-blue), MitoTracker Green (mitochondrial staining, green), or TMRM (mitochondrial staining, red); scale bars, 10 µM. For panels c, d, *p < 0.05; **p < 0.01; ***p < 0.001
Comparison of our patient’s fibroblasts with fibroblasts derived from parents
Fibroblast cultures generated from both parents (mother, M, heterozygous for the intronic mutation; father, F, heterozygous for the mutation causing a premature stop codon) showed ALDH1L2 protein expression though its levels were noticeably higher in father’s cells (Fig. 3a). Both cell lines have similar levels of ALDH3A2 proteins comparable with those of patient’s fibroblasts (Fig. 3a), and both demonstrated a much faster proliferation rate than R cells as indicated by the doubling time (Fig. 3b). ATP levels in the father’s cells were remarkably higher than in R cells (Fig. 3c). Furthermore, mitochondria from R cells have lower membrane potential (Fig. 3d) and showed increased levels of reactive oxygen species (Fig. 3e). Correspondingly, metabolomic analysis has shown increased levels (fold-change > 2; p < 0.05) of several oxidative stress biomarkers such as methionine sulfoxide, 5-oxoproline, and ophthalmate, in patient’s cells (Supplementary Data File 1). Confocal microscopy has shown differences in mitochondrial morphology in patient’s fibroblasts with the appearance of rounded isolated mitochondria, which were not seen in healthy control or father’s fibroblasts (Fig. 3f). The morphology of mitochondria in mother’s fibroblasts was similar to the morphology of the control and father’s fibroblasts though mother’s cells were smaller in size compared to fibroblasts from other individuals (Fig. 3f). These cells as well have normal doubling time (Fig. 3b) and mitochondrial membrane potential (Fig. 3d).
Electron microscopy further confirmed altered mitochondrial morphology in R cells compared to father’s cells (Supplementary Fig. 9). In contrast to mitochondria of father’s cells, which are filamentous as commonly seen in cultured fibroblasts,20 mitochondria of R cells appear to be shorter and distorted. Another noticeable feature of R cells was the presence of large vesicles not seen in F cells (Supplementary Fig. 9). We suggest that these alterations are linked to the metabolic effects caused by the ALDH1L2 deficiency. Thus, significant differences in metabolic profile in R cells were associated with amino acid, nucleotide, and lipid pathways (Fig. 4 and Supplementary Data File 1). Of note, accumulation of all common amino acids is indicative of decreased protein biosynthesis,21 which is in line with decreased proliferation and low energy status in R cells. Strong changes in the lipid profiles were seen in R cells with the most dramatic increase of acylcarnitine metabolites (Fig. 4) and the reduction of mono- and diglycerides as well as all classes of phospholipids (Supplementary Data File 1).
Metabolomic analysis of R, F, and LR fibroblasts. a Lentivirus-based expression of ALDH1L2 in LR fibroblasts restores levels of the enzyme seen in control or parent’s cells (Western blot assay of isolated mitochondria and confocal image of fibroblasts stained with ALDH1L2-specific antibody, St indicates lane with molecular weight standards, VDAC is shown as mitochondrial marker; green fluorescence indicates ALDH1L2; nuclei were co-stained with DAPI). b PCA (principal component analysis, performed with SIMCA Version 15.0.2, Sartorius Stedim Data Analytics AB, Umeå, Sweden) of metabolomic data (total of 516 metabolites) for R, F, and LR fibroblasts (n = 5 biological replicates in each case). c Heat map representation of the metabolite comparison between R, F, and LR cells (performed with Qlucore Omics Explorer v.3.4 software, Qlucore, Lund, Sweden; data were filtered by p value ≤ 0.05). d Schematic depicting the TCA cycle and its connection to carnitine pathway. e, f Levels of Krebs cycle metabolites and carnitine and most acylcarnitine derivatives are similar in F and LR cells but compared to R cells are much lower in both cell lines. Statistically significant differences (n = 5) are highlighted in green (p < 0.05, decreased metabolites), red (p < 0.05, increased metabolites), or light red (p < 0.1, increased metabolites). g Proposed mechanism for the effect of the ALDH1L2 loss
Restoration of biochemical properties by expression of wild-type ALDH1L2
Re-expression of ALDH1L2 (via viral transduction) in patient fibroblasts (LR cells, Fig. 4a) restored the morphological features of father’s cells, including re-appearance of filamentous mitochondria (Fig. 3f) and disappearance of large vesicles (Supplementary Fig. 9); decreased the doubling time (Fig. 3b); increased levels of ATP (Fig. 3c) and mitochondrial membrane potential (Fig. 3d) thus improving the energy status of these cells and making them more similar to father’s fibroblasts. Overall, the metabotype of patient cells with restored ALDH1L2 expression was shifted towards the metabotype of father cells (Fig. 4b, c) indicating that the metabotype of patient cells is associated with the loss of ALDH1L2. Specifically, Krebs cycle intermediates and acyl carnitines were similar in LR and F fibroblasts (Fig. 4d–f). These data indicate that the ALDH1L2 loss affects fatty acid metabolism. In fact, several reports implicated the enzyme in β-oxidation and fatty acid metabolism22,23,24 though underlying mechanisms are not clear. Alternatively, alterations in fatty acid metabolism could be a cellular response to oxidative stress associated with the ALDH1L2 loss.25 The phenotype rescue by transduction of ALDH1L2 indicates that the metabolic changes and mitochondrial dysfunction were not caused by Coffin–Lowry syndrome.
Comparison of our patient’s fibroblast with fibroblasts derived from an SLS patient
We have also compared our patient’s fibroblasts with the fibroblast culture from a patient with classical SLS caused by the loss of ALDH3A2 enzyme due to a homozygous mutation (c.471+1delG in intron 3) associated with the splicing abnormality of the ALDH3A2 gene. Levels of ALDH1L2 appeared normal in the SLS patient and were similar to the protein level in father’s fibroblasts (Fig. 3a). These cells have a standard doubling time and ATP levels similar to father’s cells characterized in this study (Fig. 3b, d). Confocal microscopy has shown that SLS fibroblasts have a typical mitochondrial morphology (Supplementary Fig. 10a) distinct from that observed in R cells (Fig. 3f). The metabotype of SLS fibroblasts was different from the metabotype of fibroblasts from our patient as well as metabotypes of healthy control or asymptomatic parents of our patient (Supplementary Figs. 10b, 10c, and 11; and Supplementary Data File 1). These differences suggest that our patient has a distinct biochemical basis for SLS-like symptoms.
Discussion
Patient reported here displays characteristics leading to the diagnosis of SLS-like neuro-ichthyotic syndrome. Since this patient does not have mutations in the ALDH3A2 gene, the cause of the disease remained unclear until exome sequencing revealed inherited mutations in the ALDH1L2 gene. The patient also has a deleterious frameshift mutation in the RPS6KA3 gene, which encodes for ribosomal S6 kinase (RSK2), a growth factor-regulated serine/threonine kinase and a member of the RAS-MAPK signaling pathway.10,11 Mutations in this gene are associated with Coffin–Lowry syndrome, a rare X-linked genetic disorder characterized by intellectual disability, tapering fingers, cranio-facial and skeletal abnormalities.8 Affected males were also reported to have cardiac problems, obesity, short stature, microcephaly, dental abnormalities, deafness, visual problems, hypotonia, hyperlaxity, behavioral troubles, epilepsy, sleep apneas, or drop attacks.26,27 Brain MRI can show cerebral atrophy, hypoplasia of the corpus callosum or the cerebellar vermis, ventricular dilatation or asymmetry, dilatation of the Virchow–Robin spaces leading to periventricular white matter cystic lesions, and constricted foramen magnum.28,29,30,31
Recognizing Coffin–Lowry syndrome in very young children is often difficult since physical characteristics are mild and not specific, and screening for ribosomal S6 kinase mutations is essential in most cases to confirm the diagnosis.32 In our case as well the diagnosis of Coffin–Lowry syndrome was considered only after discovering the deleterious mutation in RPS6KA3. This was a de novo somatic mutation not carried by the patient’s mother, a very common case in Coffin–Lowry disease.27,33,34 The mutation may explain some specific features of our patient (facial dysmorphism, toe and hand abnormalities, moderate cognitive and attention disabilities, and dilatation of Virchow–Robin spaces on MRI) but it is unlikely to contribute to pruritic ichthyosis, severe diffuse hypomyelination seen on MRI, and abnormal lipid peaks on 1H-MRS, features usually observed in SLS. Therefore, we attribute these features to the loss of ALDH1L2 protein. Such loss is defined by the nature of the compound mutations. Thus, the premature stop codon was predicted to produce a truncated protein (307 amino acid residues compared to 923 amino acid residues in normal protein), which includes 32 random amino acids at the C-terminus resulted from the frameshift. Such proteins are usually non-functional and are likely to undergo rapid degradation.35 In agreement with this conclusion, we did not detect the truncated protein in our patient’s fibroblasts. Of note, our previous studies indicated that the truncation of the enzyme beyond 290 aa (this does not include the mitochondrial leader sequence) produces non-functional protein.13 In the case of the frameshift mutation it would be 285 amino acids including the random peptide at the C-terminus. The splice site mutation in the second allele apparently compromised splicing, which would explain the drop in the mRNA level observed in patient’s fibroblasts. In fact, disrupted constitutive splicing most often results in loss of gene expression due to aberrant splicing.14
Our experiments with cultured patient’s fibroblasts provided a strong support for ALDH1L2 as a causative factor of patient’s conditions. Thus, cultured fibroblasts displayed abnormal mitochondrial morphology and very slow proliferation capacities, phenomena not reported for fibroblasts from patients with either Sjogren–Larsson or Coffin–Lowry syndrome. In fact, mitochondria in keratinocytes of SLS patients appear normal36,37 and our study demonstrated normal proliferation of fibroblasts from an SLS patient. Fibroblast cultures established from patients with Coffin–Lowry syndrome were characterized for signaling pathways downstream of RSK29,38,39 but effects of the protein on proliferation or mitochondria function/morphology in such fibroblasts were not reported. Interestingly though, Rsk2 deficiency in mice was associated with enhanced proliferative capacity of fibroblast-like synoviocytes,40 the effect opposite to that observed for ALDH1L2-deficient fibroblasts in our experiments. Perhaps the strongest indication for ALDH1L2 mutations as the underlying cause of the neuro-ichthyotic syndrome in our patient was the restoration of the normal mitochondrial morphology and most of the metabotype, seen in father’s cells, after re-introduction of the wild-type enzyme to the patient’s fibroblasts.
ALDH1L2 encodes a 923 amino acid residues (including 22 amino acids of the mitochondrial leader sequence in the N-terminus) protein, which resides in the mitochondrial matrix.12 So far deficiency of the ALDH1L2 gene has not been reported, and the overall consequences of the enzyme loss for the cell are not yet clear. Nonetheless, the importance of mitochondrial folate pathways for the cell is well established,41,42,43,44 and recent reports further underscored the role of folate-bound mitochondrial serine metabolism for mitochondrial integrity and oxidative phosphorylation.45,46,47 Our study provides strong evidence that ALDH1L2 is a key player in these processes and the loss of the enzyme due to deleterious gene mutations leads to neuro-ichthyotic disease. What could be underlying mechanisms of such effect? ALDH1L2 enzyme converts 10-formyl-THF to THF and CO2 in an NADP+-dependent reaction thus producing NADPH (Fig. 2e).15 Though the biological role of this reaction is not fully understood, the enzyme could be a major source of NADPH in mitochondria.19 In turn, mitochondrial NADPH is crucial for the maintenance of reduced glutathione, the major antioxidant, and the loss of NADPH is associated with increased oxidative stress. In this regard, growing body of evidence indicates the link between oxidative stress and mitochondrial dysfunction.48,49,50 For example, increased oxidative stress in skin fibroblasts of patients with multiple acyl-CoA dehydrogenation deficiency led to fragmented mitochondria,25 the morphology similar to that observed in fibroblasts of our patient. In fact, the role of ALDH1L2 protein in preventing oxidative stress has been suggested by the study of melanoma cell metastasis.51 Thus, we propose that the ALDH1L2 loss induces mitochondrial dysfunction due to reduced NADPH and increased oxidative stress (Fig. 4g).
Methods
Legal authorization and ethics approval
The parents have given their permission (written consent form) for publication of the child’s photos. Written informed consents were obtained from parents for the genetic analyses. This report is in accordance with the French “Reference methodology” (MR-001) modified on 5th January 2006 and signed by the CHU of Clermont-Ferrand on 15th March 2007 for standard patient care. This research obtained authorization of the local ethics committee (CHU of Clermont-Ferrand).
Reagents
All reagents were purchased from Sigma-Aldrich (St. Louis, MO, USA) unless otherwise specified.
Gene sequencing
ALDH3A2 exons, exon/intron junctions, and the full-length cDNA were sequenced essentially as we previously described.52 Whole exome sequencing was performed by IntegraGen SA (Evry, France) using the SureSelect V4 capture kit (Agilent, Massy, France) and the HighSeq2000 sequencer (Illumina, San Diego, CA)53 after written informed consent obtained from patient’s parents. Peripheral blood samples were drawn from the antecubital vein into 4 ml EDTA-containing tubes. Genomic DNA extraction was performed automatically from 2 ml of whole blood on a QIAsymphony SP instrument, by using the QIAsymphony DSP DNA Midi kit (QIAGEN), following the manufacturer’s protocol. There were 37,950 variations in the patient. Of these variants, 35,310 were classified as single nucleotide variations and 2640 were indels. Further analysis was focused on genes encoding proteins related to aldehyde dehydrogenases. This approach selected a single mutant gene. Interpretation was based on Human Genome Build 37 (NCBI/hg19).
Generation of fibroblasts culture
Written consent has been obtained from parents prior to biopsies according to the institutional document for tissue biopsies and referring to the French laws for ethics and protection of subjects participating in medical research. Skin biopsies were performed at the anterior forearm after anesthesia with lidocaine/prilocaine cream applied topically for 1 h. A cylindrical skin plug including epidermis and dermis was removed using a sterile 3 mm skin punch and placed in 0.9% NaCl solution. Skin samples were washed and transferred to culture dishes containing DMEM/F12, 10% of FBS, penicillin/streptomycin/amphotericin (final concentration of 200 U/ml, 0.2 and 0.5 μg/ml, respectively) and maintained at 37 °C and under humidified air containing 5% CO2. After 1 week, the medium of the explant cultures was changed every 2–3 days. When confluent, cells were expanded. All fibroblast cell lines were maintained in DMEM (Gibco) supplemented with 10% FBS (Atlanta Biologicals, Flowery Branch, GA, USA) and 5% penicillin/streptomycin/neomycin cocktail (Gibco).
Doubling time assay
Cell number was determined by hemocytometer or Countess II cell counter (Thermo, Waltham, MA, USA). In folate supplementation experiments, folinic acid or folic acid were added at 10 and 20 µM, respectively. Doubling time assays were performed by seeding a confluent 10-cm dishe of each cell line onto 6-cm dishes in triplicate. The seed time was considered T0 and cells were left to proliferate for 24 h. Cell numbers at 0 and 24 h were determined by trypan blue exclusion assay using hemocytometer.
RNA extraction and cDNA synthesis
Total RNA was isolated from 2 × 106 cells using an RNeasy mini kit (Qiagen). One microgram of total RNA was used in a reverse-transcription reaction to generate cDNA using high capacity cDNA reverse transcription kit (Applied Biosystems).
Real-time PCR
Quantification of mRNAs was carried out by real-time PCR Realplex4 Mastercycler (Eppendorf, Hauppauge, NY, USA) using RT2 SYBR Green PCR master mix (Applied Biosystem) in final 20 μl PCR mixture containing 10 μl SYBR Premix EX Taq, 2 μl cDNA (100 ng), 0.4 μl (10 μM) forward and 0.4 μl (10 μM) reverse primers and 6.8 μl ddH2O. The PCR protocol was as follows: initial 95 °C melting for 5 min, then 40 cycles of denaturation at 95 °C for 30 s, annealing at 60 °C for 30 s, and elongation at 72 °C for 20 s. Levels of ALDH1L1 and ALDH1L2 mRNA were normalized by the levels of actin as housekeeping gene. The fold change in mRNA expression was calculated using 2−ΔΔCt.
Cell lysate preparation and mitochondria isolation
Mitochondrial fractions were obtained using a mitochondrial isolation kit (MACS; Miltenyi Biotech, Auburn, CA, USA) following the manufacturer’s protocol.
Real-time cell analysis
Experiments were carried out using an xCELLigence RTCA DP instrument (ACEA Biosciences, San Diego, CA, USA) placed in a humidified incubator at 37 °C and 5% CO2 according to the manufacturer’s manual. Cell proliferation was monitored using E-plate 16 (ACEA Biosciences). The background impedance reading for each well was set up using cell-free medium (100 µl per well) after pre-incubation at room temperature for 30 min. Cells were seeded in each well in 100 µl cell suspensions across a concentration range of 5 × 103 to 8 × 104 cells/well and allowed to attach for 30 min at room temperature. Plates were locked in the instrument, and impedance readings of each well were automatically recorded every 15 min for the duration of the experiment.
Metabolomic analysis
Cells were cultured in 15 cm dishes, grown to 70–80% confluency, harvested, and subsequently flash frozen. Sample preparation for analysis was carried out at Metabolon Inc., as described.54 Briefly, individual samples were subjected to methanol extraction then split into aliquots for analysis by ultrahigh performance liquid chromatography/mass spectrometry (UHPLC/MS). The global biochemical profiling analysis comprised of four unique arms consisting of reverse phase chromatography positive ionization methods optimized for hydrophilic compounds (LC/MS Pos Polar) and hydrophobic compounds (LC/MS Pos Lipid), reverse phase chromatography with negative ionization conditions (LC/MS Neg), as well as a HILIC chromatography method coupled to negative (LC/MS Polar).55 All of the methods alternated between full scan MS and data dependent MSn scans. The scan range varied slightly between methods but generally covered 70–1000 m/z. Metabolites were identified by automated comparison of the ion features in the experimental samples to a reference library of chemical standard entries that included retention time, molecular weight (m/z), preferred adducts, and in-source fragments as well as associated MS spectra and curated by visual inspection for quality control using software developed at Metabolon. Identification of known chemical entities was based on comparison to metabolomic library entries of purified standards.56
Statistical analysis
Two types of statistical analyses were performed: (1) significance tests and (2) classification analysis. Standard statistical analyses were performed in ArrayStudio on log‐transformed data. For analyses not standard in ArrayStudio, the R program (http://cran.r-project.org/) was used. Following log transformation and imputation of missing values, if any, with the minimum observed value for each compound, Welch’s two sample t-test was used as significance test to identify biochemicals that differed significantly (p < 0.05) between experimental groups. An estimate of the false discovery rate (q‐value) was calculated to take into account the multiple comparisons that normally occur in metabolomic‐based studies. Classification analyses used included principal components analysis (PCA), hierarchical clustering, and OPLS-DA. For the scaled intensity graphics, each biochemical in original scale (raw area count) was rescaled to set the median across all samples equal to 1.
ATP and NADPH/NADP+ assays
Cells were cultured in 15 cm dishes, harvested at 70–80% confluency, and flash frozen. ATP and NADPH/NADP+ were measured in whole cell lysate or isolated mitochondria using colorimetric ATP and fluorescence NADPH/NADP+ kits (Abcam), respectively according to the manufacturer’s protocols. Fifty microliters of the prepared sample were used in the assay. Experiments were carried out four times in triplicate.
Metabolite extraction and HPLC assays
Metabolite extraction and HPLC analysis were performed according to published procedures.57,58 Cells cultured in 10-cm plates were washed two times with ice-cold PBS immediately prior to the addition of extraction buffer (1.5 ml of 9:1 methanol/chloroform mixture). Following the addition of extraction buffer, plates were scraped, and the extracts were kept at −20 °C overnight. Samples were spun down at 4 °C for 20 min at 21,000 × g, supernatants were placed to new Eppendorf tubes, and dried out using Centrivap (Labconco). Metabolites were dissolved in 1 mM Tris pH 9.0, passed through a 0.22 μm filter (Thermo), and resolved on a Symmetry C18 guard column (3.9 × 5 mm/4.6 × 250 mm, 5 μm particle size) using a Waters HPLC system (Milford, MA, USA). The elution solvents were (A) 0.1 M KH2PO4 pH 6.0 and (B) 0.1 M KH2PO4, 50% methanol pH 6.0. Elution conditions were solvent A (11 min) and a linear gradient of 0–20% solvent B. Peaks were detected by absorbance at 260 nm. The retention time was 8, 9, and 14 min for ATP, ADP, and AMP, respectively.
Western blot analysis
Cells were lysed using RIPA buffer containing protease and phosphatase inhibitor cocktails. The Bradford protein assay (BioRad) was used to quantify protein concentrations. Equal amounts of protein (20 μg) were separated on a 7.5–12% Tris–glycine SDS polyacrylamide gel and were transferred to nitrocellulose membranes (GE). Membranes were blocked with 5% BSA in TBS supplemented with 0.1% Tween-20 for 1 h at room temperature and were incubated with primary antibody overnight at 4 °C. After incubating with horseradish peroxidase-conjugated secondary antibodies (anti-rabbit, NXA931; anti-mouse, NA934V; both from GE; 1:10,000), membranes were developed using supersignal chemiluminescence reagents (Thermo). Blots were striped with a buffer containing 1.5% glycine, 0.1% SDS, and 1% Tween-20 at pH 2.2. ALDH1L2 was detected using in-house ALDH1L2-specific polyclonal antibody (1:2000).12 ALDH3A2 (ab113111), VDAC (ab154856), and GAPDH (ab8245) antibodies were from Abcam. Actin antibody (sc-47778) was purchased from Santa Cruz. All commercial antibodies were used at the dilution of 1:1000. All blots derived from the same experiment were processed in parallel. Band intensities were quantified using ImageJ software, NIH (https://imagej.nih.gov/ij/). Uncropped blot images are shown in Supplementary Fig. 12.
Immunocytochemistry
Cells were plated onto glass bottom microwell dishes (MatTek Corp., Ashland, MA, USA), left to attach overnight and then were stained with 250 nM TMRM, 100 nM Mitotracker, and 0.5 μg/ml Hoechst (all from Molecular Probes). In a separate experiment, cells were also stained with 40 μM H2DCFDA for whole cell ROS visualization. Cells were stained independently and imaged immediately following dye incubation to ensure changes in ROS were not a product of imaging time differences. For fixed cell immunocytochemistry, cells were cultured on glass coverslips and subsequently fixed in 4% paraformaldehyde, quenched with ammonia sulfate, and washed with PHEM buffer. Cells were permeabilized with 0.1% Triton-100X and blocked with 5% goat serum for 1 h at room temperature. Cells were then stained with anti-ALDH1L2 antibody (1:250) for 1 h, rinsed with PHEM, and incubated with secondary anti-rabbit IGG goat antibody (Molecular Probes, A21441, 1:500). Coverslips were mounted to glass slides with mountant containing DAPI (Thermo) and left to dry overnight at 4 °C and were imaged the following day.
Analysis of mitochondrial mass and membrane potential
Cells (1.5 × 105) were seeded onto 6 cm plates and left to attach overnight. Cells were harvested, resuspended in 5 ml of fresh medium and stained with 250 nM TMRM and then with 100 nM MitoTracker Green (both from Molecular Probes) to determine mitochondrial polarization and mitochondrial mass respectively. Intensity of each dye was determined using CytoFLEX flow cytometer (Beckman Coulter, Indianapolis, IN, USA). Quantitative analysis was completed using CytoFLEX software.
ALDH1L2 lentiviral transduction
The construct of human ALDH1L2 cDNA cloned to a pLenti-GIII-CMV lentiviral vector was purchased from Applied Biological Materials Inc. (Richmond, BC, Canada). Recombinant lentivirus for the ALDH1L2 expression was produced using ViraPower lentiviral expression system (Thermo) according to the manufacturer’s protocol. Patient’s fibroblasts grown in a 6-well plate (1 × 106 cells per well) were transduced with 2 ml of the mixture of RPMI and the generated viral stock (1:1) for 24 h. The efficiency of lentiviral transduction was confirmed by the killing curve selection assay with puromycin and individual clones were cultured.
Assays of reduced folate pools
Approximately 5 × 106 cells were collected and rapidly washed three times with ice-cold PBS. The cell pellet was resuspended in 50 mM Tris–HCl buffer, pH 7.4, containing 50 mM sodium ascorbate. Cells were lysed by heating for 3 min in a boiling water bath. Cell lysates were chilled on ice and centrifuged for 5 min at 17,000 × g at 4 °C. Folate pools were measured in cell lysates by the ternary complex assay method as described.59,60 Folate levels were calculated per mg of cellular protein measured by Bradford assay.
Transmission electron microscopy
Cells were plated on Nunc Permanox slide chambers (3000 cells per well) in DMEM and left to attach overnight. Cells were fixed for 1 h at room temperature with 2.5% formaldehyde/glutaraldehyde (1:1) in 0.1 M sodium cacodylate buffer pH 7.4. Slides were stored at 4 °C until further processing. Following three rinses with 0.1 M sodium cacodylate buffer, pH 7.4, cells were post-fixed with 1% osmium tetroxide/1.25% potassium ferrocyanide/0.1 M sodium cacodylate buffer for 1 h at room temperature. After washes in deionized water, cells were dehydrated using increasing concentrations of ethanol (30%, 50%, 75%, and 100% twice, 10 min each) and embedded in Polybed 812 epoxy resin (Polysciences, Inc., Warrington, PA). Cells were sectioned en face to the substrate at 70 nm using a diamond knife and Leica Ultracut UCT ultramicrotome (Leica Microsystems, Inc., Buffalo Grove, IL). Ultrathin sections were collected on 200 mesh copper grids and stained with 4% aqueous uranyl acetate for 12 min, followed by Reynolds’ lead citrate for 8 min. Samples were observed with a JEOL JEM-1230 transmission electron microscope operating at 80 kV (JEOL USA, Peabody, MA) and digital images acquired using a Gatan Orius SC1000 CCD camera and Gatan Microscopy Suite 3.0 software (Gatan, Inc., Pleasanton, CA). Magnification for each image set includes 5000×, 10,000×, and 20,000×.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
All data generated or analyzed during this study are included in this published article and Supplementary Information files or are available from the author upon reasonable request.
References
Rizzo, W. B., Jenkens, S. M. & Boucher, P. Recognition and diagnosis of neuro-ichthyotic syndromes. Semin. Neurol. 32, 75–84 (2012).
Rizzo, W. B. Sjogren–Larsson syndrome: molecular genetics and biochemical pathogenesis of fatty aldehyde dehydrogenase deficiency. Mol. Genet. Metab. 90, 1–9 (2007).
Willemsen, M. A. et al. Clinical, biochemical and molecular genetic characteristics of 19 patients with the Sjogren–Larsson syndrome. Brain 124, 1426–1437 (2001).
Weustenfeld, M. et al. Genotype and phenotype variability in Sjogren–Larsson syndrome. Hum. Mutat. 40, 177–186 (2019).
Cho, K. H., Shim, S. H. & Kim, M. Clinical, biochemical, and genetic aspects of Sjogren–Larsson syndrome. Clin. Genet. 93, 721–730 (2018).
Rizzo, W. B. Genetics and prospective therapeutic targets for Sjogren–Larsson syndrome. Expert Opin. Orphan Drugs 4, 395–406 (2016).
Scalais, E., Verloes, A., Sacre, J. P., Pierard, G. E. & Rizzo, W. B. Sjogren–Larsson-like syndrome with bone dysplasia and normal fatty alcohol NAD+ oxidoreductase activity. Pediatr. Neurol. 8, 459–465 (1992).
Trivier, E. et al. Mutations in the kinase Rsk-2 associated with Coffin–Lowry syndrome. Nature 384, 567–570 (1996).
Yang, X. et al. ATF4 is a substrate of RSK2 and an essential regulator of osteoblast biology; implication for Coffin–Lowry syndrome. Cell 117, 387–398 (2004).
Hauge, C. & Frodin, M. RSK and MSK in MAP kinase signalling. J. Cell Sci. 119, 3021–3023 (2006).
Carriere, A., Ray, H., Blenis, J. & Roux, P. P. The RSK factors of activating the Ras/MAPK signaling cascade. Front. Biosci. 13, 4258–4275 (2008).
Krupenko, N. I. et al. ALDH1L2 is the mitochondrial homolog of 10-formyltetrahydrofolate dehydrogenase. J. Biol. Chem. 285, 23056–23063 (2010).
Reuland, S. N., Vlasov, A. P. & Krupenko, S. A. Modular organization of FDH: exploring the basis of hydrolase catalysis. Protein Sci. 15, 1076–1084 (2006).
Faustino, N. A. & Cooper, T. A. Pre-mRNA splicing and human disease. Genes Dev. 17, 419–437 (2003).
Strickland, K. C. et al. Enzymatic properties of ALDH1L2, a mitochondrial 10-formyltetrahydrofolate dehydrogenase. Chem. Biol. Interact. 191, 129–136 (2011).
Krupenko, S. A. & Krupenko, N. I. ALDH1L1 and ALDH1L2 folate regulatory enzymes in cancer. Adv. Exp. Med. Biol. 1032, 127–143 (2018).
Tibbetts, A. S. & Appling, D. R. Compartmentalization of Mammalian folate-mediated one-carbon metabolism. Annu. Rev. Nutr. 30, 57–81 (2010).
Krupenko, S. A. FDH: an aldehyde dehydrogenase fusion enzyme in folate metabolism. Chem. Biol. Interact. 178, 84–93 (2009).
Fan, J. et al. Quantitative flux analysis reveals folate-dependent NADPH production. Nature 510, 298–302 (2014).
Brantova, O. et al. Ultrastructural changes of mitochondria in the cultivated skin fibroblasts of patients with point mutations in mitochondrial DNA. Ultrastruct. Pathol. 30, 239–245 (2006).
Ashkavand, Z. et al. Metabolic reprogramming by folate restriction leads to a less aggressive cancer phenotype. Mol. Cancer Res. 15, 189–200 (2017).
Sun, R. et al. Benzene exposure alters expression of enzymes involved in fatty acid beta-oxidation in male C3H/He mice. Int. J. Environ. Res. Public Health 13, https://doi.org/10.3390/ijerph13111068 (2016).
Edhager, A. V. et al. Proteomic investigation of cultivated fibroblasts from patients with mitochondrial short-chain acyl-CoA dehydrogenase deficiency. Mol. Genet. Metab. 111, 360–368 (2014).
Ibeagha-Awemu, E. M. et al. Transcriptome adaptation of the bovine mammary gland to diets rich in unsaturated fatty acids shows greater impact of linseed oil over safflower oil on gene expression and metabolic pathways. BMC Genomics 17, https://doi.org/10.1186/s12864-016-2423-x (2016).
Cornelius, N., Corydon, T. J., Gregersen, N. & Olsen, R. K. Cellular consequences of oxidative stress in riboflavin responsive multiple acyl-CoA dehydrogenation deficiency patient fibroblasts. Hum. Mol. Genet. 23, 4285–4301 (2014).
Nishimoto, H. K. et al. The historical Coffin–Lowry syndrome family revisited: identification of two novel mutations of RPS6KA3 in three male patients. Am. J. Med. Genet. A 164A, 2172–2179 (2014).
Pereira, P. M., Schneider, A., Pannetier, S., Heron, D. & Hanauer, A. Coffin–Lowry syndrome. Eur. J. Hum. Genet. 18, 627–633 (2010).
Miyata, Y. et al. Periventricular small cystic lesions in a patient with Coffin–Lowry syndrome who exhibited a novel mutation in the RPS6KA3 gene. Brain Dev. 40, 566–569 (2018).
Tos, T., Alp, M. Y., Aksoy, A., Ceylaner, S. & Hanauer, A. A familial case of Coffin–Lowry syndrome caused by RPS6KA3 C.898C>T mutation associated with multiple abnormal brain imaging findings. Genet. Couns. 26, 47–52 (2015).
Wang, Y. et al. A novel RSK2 (RPS6KA3) gene mutation associated with abnormal brain MRI findings in a family with Coffin–Lowry syndrome. Am. J. Med. Genet. A 140, 1274–1279 (2006).
Kesler, S. R. et al. Altered neurodevelopment associated with mutations of RSK2: a morphometric MRI study of Coffin–Lowry syndrome. Neurogenetics 8, 143–147 (2007).
Touraine, R. L., Zeniou, M. & Hanauer, A. A syndromic form of X-linked mental retardation: the Coffin–Lowry syndrome. Eur. J. Pediatr. 161, 179–187 (2002).
Jacquot, S. et al. Mutation analysis of the RSK2 gene in Coffin–Lowry patients: extensive allelic heterogeneity and a high rate of de novo mutations. Am. J. Hum. Genet. 63, 1631–1640 (1998).
Hanauer, A. & Young, I. D. Coffin–Lowry syndrome: clinical and molecular features. J. Med. Genet. 39, 705–713 (2002).
Oren, Y. S., Pranke, I. M., Kerem, B. & Sermet-Gaudelus, I. The suppression of premature termination codons and the repair of splicing mutations in CFTR. Curr. Opin. Pharmacol. 34, 125–131 (2017).
Matsuoka, L. Y., Kousseff, B. G. & Hashimoto, K. Studies of the skin in Sjogren–Larsson syndrome by electron microscopy. Am. J. Dermatopathol. 4, 295–301 (1982).
Ito, M., Oguro, K. & Sato, Y. Ultrastructural study of the skin in Sjogren–Larsson syndrome. Arch. Dermatol. Res. 283, 141–148 (1991).
Sassone-Corsi, P. et al. Requirement of Rsk-2 for epidermal growth factor-activated phosphorylation of histone H3. Science 285, 886–891 (1999).
Xing, J., Ginty, D. D. & Greenberg, M. E. Coupling of the RAS-MAPK pathway to gene activation by RSK2, a growth factor-regulated CREB kinase. Science 273, 959–963 (1996).
Derer, A. et al. Rsk2 controls synovial fibroblast hyperplasia and the course of arthritis. Ann. Rheum. Dis. 75, 413–421 (2016).
Momb, J. et al. Deletion of Mthfd1l causes embryonic lethality and neural tube and craniofacial defects in mice. Proc. Natl. Acad. Sci. USA 110, 549–554 (2013).
Tani, H. et al. Mice deficient in the Shmt2 gene have mitochondrial respiration defects and are embryonic lethal. Sci. Rep. 8, https://doi.org/10.1038/s41598-017-18828-3 (2018).
Pai, Y. J. et al. Glycine decarboxylase deficiency causes neural tube defects and features of non-ketotic hyperglycinemia in mice. Nat. Commun. 6, https://doi.org/10.1038/ncomms7388 (2015).
Ben-Sahra, I., Hoxhaj, G., Ricoult, S. J., Asara, J. M. & Manning, B. D. mTORC1 induces purine synthesis through control of the mitochondrial tetrahydrofolate cycle. Science 351, 728–733 (2016).
Minton, D. R. et al. Serine catabolism by SHMT2 is required for proper mitochondrial translation initiation and maintenance of formylmethionyl-tRNAs. Mol. Cell 69, 610–621 (2018).
Morscher, R. J. et al. Mitochondrial translation requires folate-dependent tRNA methylation. Nature 554, 128–132 (2018).
Gao, X. et al. Serine availability influences mitochondrial dynamics and function through lipid metabolism. Cell Rep. 22, 3507–3520 (2018).
Ying, W. NAD+/NADH and NADP+/NADPH in cellular functions and cell death: regulation and biological consequences. Antioxid. Redox Signal. 10, 179–206 (2008).
Dai, D. F., Chiao, Y. A., Marcinek, D. J., Szeto, H. H. & Rabinovitch, P. S. Mitochondrial oxidative stress in aging and healthspan. Longev. Healthspan 3, https://doi.org/10.1186/2046-2395-3-6 (2014).
Bradshaw, P. C. Cytoplasmic and mitochondrial NADPH-coupled redox systems in the regulation of aging. Nutrients 11, https://doi.org/10.3390/nu11030504 (2019).
Piskounova, E. et al. Oxidative stress inhibits distant metastasis by human melanoma cells. Nature 527, 186–191 (2015).
Sarret, C. et al. Sjogren–Larsson syndrome: novel mutations in the ALDH3A2 gene in a French cohort. J. Neurol. Sci. 312, 123–126 (2012).
Barbier, M. et al. MFAP5 loss-of-function mutations underscore the involvement of matrix alteration in the pathogenesis of familial thoracic aortic aneurysms and dissections. Am. J. Hum. Genet. 95, 736–743 (2014).
Hatano, T., Saiki, S., Okuzumi, A., Mohney, R. P. & Hattori, N. Identification of novel biomarkers for Parkinson’s disease by metabolomic technologies. J. Neurol. Neurosurg. Psychiatry 87, 295–301 (2016).
Evans, A. M. et al High resolution mass spectrometry improves data quantity and quality as compared to unit mass resolution mass spectrometry in high‐throughput profiling metabolomics. Metabolomics 4, https://doi.org/10.4172/2153-0769.1000132 (2014).
Dehaven, C. D., Evans, A. M., Dai, H. & Lawton, K. A. Organization of GC/MS and LC/MS metabolomics data into chemical libraries. J. Cheminform. 2, https://doi.org/10.1186/1758-2946-2-9 (2010).
Lorenz, M. A., Burant, C. F. & Kennedy, R. T. Reducing time and increasing sensitivity in sample preparation for adherent mammalian cell metabolomics. Anal. Chem. 83, 3406–3414 (2011).
Formato, M., Masala, B. & De Luca, G. The levels of adenine nucleotides and pyridine coenzymes in red blood cells from the newborn, determined simultaneously by HPLC. Clin. Chim. Acta 189, 131–137 (1990).
Oleinik, N. V., Krupenko, N. I., Reuland, S. N. & Krupenko, S. A. Leucovorin-induced resistance against FDH growth suppressor effects occurs through DHFR up-regulation. Biochem. Pharmacol. 72, 256–266 (2006).
Hoeferlin, L. A., Oleinik, N. V., Krupenko, N. I. & Krupenko, S. A. Activation of p21-dependent G1/G2 arrest in the absence of DNA damage as an antiapoptotic response to metabolic stress. Genes Cancer 2, 889–899 (2011).
Acknowledgements
The authors thank Mélanie Rigal, Catherine Vaurs-Barrière, Patricia Combes, and Geneviève Giraud (Clermont-Ferrand); Dr. Ting Li and Dr. David Horita (Nutrition Research Institute, UNC Chapel Hill). S.A.K. is supported by the National Institutes of Health grant R01 DK117854. The Microscopy Services Laboratory, Department of Pathology and Laboratory Medicine, UNC, is supported in part by P30 CA016086 Cancer Center Core Support Grant to the UNC Lineberger Comprehensive Cancer Center.
Author information
Authors and Affiliations
Contributions
The overall study design: C.S. and S.A.K. Clinical follow-up: C.S., C.F. and O.B.-T. ALDH3A2 gene sequencing: C.S. and E.E.-P. Whole exome sequencing: I.D. and O.B.-T. Generation of fibroblast culture: E.E.-P. Folate assays and HPLC nucleotide assays: P.P. and N.I.K. Metabolomic data analysis and interpretation: S.S., N.I.K. and S.A.K. Electron microscopy: E.P. and P.P. Fibroblast characterization, transfection, sample preparation, and analysis: Z.A. and E.P. All authors contributed to the discussion of results and manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sarret, C., Ashkavand, Z., Paules, E. et al. Deleterious mutations in ALDH1L2 suggest a novel cause for neuro-ichthyotic syndrome. npj Genom. Med. 4, 17 (2019). https://doi.org/10.1038/s41525-019-0092-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41525-019-0092-9
This article is cited by
-
Network-based inference of master regulators in epithelial membrane protein 2-treated human RPE cells
BMC Genomic Data (2022)
-
Therapeutic targeting of the mitochondrial one-carbon pathway: perspectives, pitfalls, and potential
Oncogene (2021)
-
Aldh1l2 knockout mouse metabolomics links the loss of the mitochondrial folate enzyme to deregulation of a lipid metabolism observed in rare human disorder
Human Genomics (2020)
-
Cytosolic 10-formyltetrahydrofolate dehydrogenase regulates glycine metabolism in mouse liver
Scientific Reports (2019)