Abstract
We evaluate microsatellite instability-high (MSI-H) status with cell-free DNA (cfDNA) in metastatic breast cancer (MBC) and the association with clinico-genomic characteristics. Patients with MSI-H in cfDNA (Guardant360®, 74 gene next-generation sequencing (NGS) with MBC are identified. We conduct a retrospective review. The median number of alterations and a median maximum mutant allelic fraction (MAF) in MSI-H and non-MSI-H cohorts are compared with Mann–Whitney U-test. Of 6718 patients with breast cancer with ≥1 plasma NGS alteration, 42 (0.63%) have MSI-H. A median number of genomic alterations per sample is 11 in MSI-H vs. 3 in non-MSI-H (Mann–Whitney U-test p < 0.0001) and the median maximum MAF is 16.8% in MSI-H vs. 2.6% in non-MSI-H (Mann–Whitney U-test p < 0.0001). The co-existing genomic landscape is heterogeneous. The median response duration for seven patients receiving immunotherapy is 92 days (range 29–273 days). CfDNA can identify MSI-H in MBC. Research is needed to validate immunotherapy usage in cfDNA-detected MSI-H MBC.
Similar content being viewed by others
Introduction
Microsatellite instability (MSI) occurs in many types of tumors from defects in mismatch repair genes, resulting in a mismatch repair deficiency1,2,3,4,5. One group identified MSI occurring in 3.8% of cancers assessed, across 39 different cancer types2. Tumors with MSI can be further delineated by the amount of instability, as MSI-high (MSI-H) or MSI-low (MSI-L)6. Somatic or germline mutations in mismatch repair genes can occur, with both types of alterations associated with MSI-H cancers, and germline mutations can also be found in association with Lynch syndrome7,8,9.
Microsatellite instability can be indicative of hyper-mutated tumors arising from mismatch repair deficiency4. Tumors with mismatch repair deficiency have been shown to respond to programmed cell death 1 axis blockade10,11, as have tumors with a high mutational burden12. Immune checkpoint inhibition is now approved for the treatment of advanced solid tumors with MSI-H and/or mismatch repair deficiency13. MSI-H status has typically been determined through the evaluation of tumor tissue4.
Plasma-based genotyping is being explored as a complementary methodology to tumor tissue genotyping as it has the advantage of being less invasive than tumor tissue genotyping and can be repeated at the time of disease progression, which may capture a changing genomic landscape under the pressure of treatment14. Limitations of this technology include reduced sensitivity in low-shedding tumors15. The utility of immunotherapy in MSI-H tumors detected using plasma-based genotyping is currently not known and needs to be understood as plasma-based genotyping assays are being used more commonly in metastatic breast cancer (MBC) to identify targetable mutations14.
The objective of this study is to evaluate MSI-H detection using a plasma-based genotyping assay (Guardant360®, cell-free DNA (cfDNA), up to 74 gene assay) in a cohort of patients with MBC undergoing plasma-based genotyping, and study the association of MSI-H with clinico-genomic characteristics. We demonstrate that MSI-H status can be identified via cfDNA. MSI-H status correlated with a higher median number of genomic alterations and a medium maximum mutant allelic fraction (MAF). We also retrospectively evaluate the response to immunotherapy in seven patients with MSI-H MBC.
Results
Patient characteristics
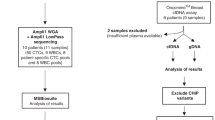
Of 6718 patients with advanced breast cancer with at least one alteration detected in cfDNA, 42 (0.63%) were MSI-H (with 43 samples).
All 42 patients in the MSI-H cohort were female. Their median age was 61 years (range 39–92). As shown in Fig. 1a, the median number of genomic alterations per sample in this MSI-H cohort was 11 (range 2–67) compared to 3 (range 1–206) in the non-MSI-H samples (Mann–Whitney U-test p < 0.0001). Even when limiting to MSI-H and non-MSI-H samples with a maximum variant allele fraction (maxVAF) of ≥5%, MSI-H samples maintained a significantly higher median number of genomic alterations per sample (11 versus 4 alterations, Mann–Whitney U-test p < 0.0001). The median maximum mutant allelic fraction (MAF) for the MSI-H cohort was 16.8% (range 0.9–54.2%) compared to 2.6% (range 0.02–96.5%) for the non-MSI-H samples (Mann–Whitney U-test p < 0.0001) as shown in Fig. 1b.
a Distribution of the number of somatic alterations detected per sample, including synonymous variants and variants of uncertain significance, excluding amplifications, in MSI-H versus non-MSI-H samples. The number of somatic alterations in individual samples is represented by colored dots with median value and interquartile range outlined in black. b Median maximum MAF in MSI-H versus non-MSI-H samples (Mann–Whitney U-test p < 0.0001).
Co-existing genomic landscape
Figure 2A depicts the most common co-existing non-synonymous mutations (including single nucleotide variants (SNVs), indels, and a few amplifications) seen in conjunction with MSI-H in this cohort. As demonstrated in Fig. 2a, the co-existing genomic landscape in patients with MSI-H is heterogeneous, including mutations in TP53, PI3KCA, ESR1, RB1, EGFR, ARID1A, BRAF, NOTCH1, DNA damage repair genes (ATM and BRCA2), APC, PDGFRA, PTEN, MET, KIT, and FGFR2. The frequencies of these mutations are indicated in Fig. 2a.
a Most common non-synonymous variants seen in MSI-H patients with ≥1 cfDNA alteration detected. Absolute number of patients are indicated in (#). b Comparison of 15 most frequent gene mutations (excluding VUS and synonymous variants) in MSI-H samples (red) and non-MSI-H samples (blue). c Differences in the frequency of genes (excluding synonymous variants, but including VUS) in MSI-H (red) vs non-MSI-H (blue) samples (* indicating significant differences, p < 0.05 one-sided Fisher exact test) of the top 20 significantly mutated genes between groups.
Figure 2b compares the most frequent gene mutations (excluding variants of uncertain significance, VUS, and synonymous variants) in MSI-H and non-MSI-H samples, while statistically significant differences in gene frequencies (excluding synonymous variants) in MSI-H vs. non-MSI-H samples in the top 20 mutated genes between these cohorts are shown in Fig. 2c. Figure 2b, c were generated using cBio Cancer Genomics Portal16,17.
Clinical treatment course
For 11/42 MSI-H patients with MBC, clinical data were available, as summarized in Table 1. Both triple-negative breast cancer (TNBC: 36%) and hormone receptor-positive, HER2 negative (HR+/HER2−) MBC (64%) were seen. None of these 11 patients had a known diagnosis of Lynch syndrome. Two of three patients who had undergone MSI testing of a metastatic tumor tissue specimen also had MSI-H positive disease detected via tissue. Patient 2 had next-generation sequencing (NGS) done of their metastatic liver lesion, which also identified MSI-H. Patient 5 had immunohistochemistry (IHC) testing of the mismatch repair (MMR) genes conducted on a metastatic liver lesion, which identified intact expression of the MMR genes. Patient 6 had tissue NGS testing of a supraclavicular lymph node, which also detected MSI-H. Seven patients underwent treatment with various immune checkpoint inhibitors, as described in Table 2. The median duration of treatment was 92 days (range 29–273 days). There did not appear to be an association between the number of prior lines of therapy and the duration of response, albeit the cohort size was limited. Two patients had a sustained benefit to immunotherapy in combination with chemotherapy, including Patient 1, who was treated with first-line atezolizumab/nab-paclitaxel and experienced stable disease for 273 days (Fig. 3), and Patient 7, who was treated with pembrolizumab/capecitabine followed by pembrolizumab/eribulin in the fifth-line setting who remained on immunotherapy treatment for 245 days; both of these patients did not have prior exposure to either the immunotherapy or chemotherapy agents. Of the two patients whose MSI-H call was concordant with tissue testing, Patient 2 received first-line atezolizumab/nab-paclitaxel and remained on treatment for 91 days, while Patient 6 received fourth-line pembrolizumab monotherapy and remained on treatment for 65 days. Patient 5, who had discordant tissue and plasma MSI-H calls, received tenth-line pembrolizumab monotherapy and remained on treatment for 29 days.
Discussion
Plasma-based genotyping assays are being used more commonly in MBC, particularly after approval of a targeted therapy, alpelisib, for PIK3CA mutant HR+/HER2− MBC18. The non-invasive nature of these assays means they can be repeated easily throughout the disease course to capture tumor evolution and may be particularly useful for patients in whom tissue testing is difficult (e.g., for patients with bone-only disease) or not informative (e.g., a “quality not sufficient” result)14, although an acknowledged limitation is a potentially reduced sensitivity in low-shedding tumors. Furthermore, tumor tissue biopsies are generally obtained at the time of diagnosis of MBC, and therefore genotyping done on baseline specimens may not necessarily reflect the current genomic environment after treatment with various therapies, which real-time plasma-based genotyping may capture. As plasma-based genotyping is increasingly being used to detect actionable mutations such as PIK3CA, the ability to identify other genomic alterations such as MSI-H expands the potential clinical application of these assays.
In this study, we evaluated a plasma-based genotyping assay to identify the presence of MSI-H in advanced breast cancer. In our cohort of 6718 patients with at least 1 detectable mutation found in plasma-based genotyping, MSI-H positive tumors accounted for 0.63%. Our findings suggest that while MSI-H is detectable using a plasma-based assay, it is relatively rare in MBC. Previous studies have suggested an MSI-H frequency in breast cancer ranging from 0–1.5%, aligned with the frequency detected here2,3,19,20,21. In one study of 444 breast cancer samples, IHC of the mismatch repair proteins identified mismatch repair deficiency in 17% of patients. However, when these same 75 samples were tested using a PCR assay utilizing the five indicator sites recommended by the National Cancer Institute, 68/75 (91%) were classified as microsatellite stable22. Variations in frequency between all of these studies may reflect differences in how each assay calls MSI-H and differences between the breast cancer cohorts used in each study. Further research is needed to identify the optimal methodology for the detection of MSI-H in breast cancer and validate the clinical utility of plasma-based genotyping in this setting.
In our cohort, MSI-H-positive breast cancers demonstrated a significantly higher number of somatic alterations, even when limited to samples with ≥5% maxVAF. As MSI-H is suggestive of impaired DNA mismatch repair, it seems consistent that MSI-H tumors have significantly more somatic alterations. The prognostic implications of these tumor characteristics in breast cancer need to be further explored. We also observed a wide spectrum of co-existing mutations, including mutations in TP53, PI3KCA, ESR1, RB1, EGFR, ARID1A, BRAF, NOTCH1, DNA damage repair genes (ATM and BRCA2), APC, PDGFRA, PTEN, MET, KIT, and FGFR2, consistent with the known genomic complexity of MBC23. In addition, MBC may harbor the APOBEC signature in keeping with increasing genomic complexity23, and it is not known how MSI-H status correlates with the presence of the APOBEC signature. While there is limited literature on how the APOBEC mutational signature relates to MSI-H, prior studies have observed an increased tumor mutational burden in both MSI-H and APOBEC mutated tumors24,25. Understanding the relationship of the APOBEC signature with MSI-H, and how these biomarkers may correlate with response to immunotherapy, is a direction for future research, as we were unable to assess these associations in our cohort.
Although our clinical response cohort is small, it is intriguing to note that some patients had durable responses to treatment with immunotherapy and chemotherapy, including two patients who remained on immunotherapy and chemotherapy for over 245 days. It is also interesting to note that both patients with concordant plasma and tissue MSI-H calls who received immunotherapy (±chemotherapy) remained on treatment for less than 100 days. The patient with discordant plasma and tissue MSI-H calls received pembrolizumab monotherapy for 29 days, although this was the most heavily pretreated patient in the cohort at their tenth-line of therapy. The small sample size, the combination of immunotherapy ± chemotherapy regimens used, and the mixture of lines of therapy make it difficult to interpret the duration of response seen here. However, the fact that a patient who was MSI-H via both plasma and tissue treated with immunotherapy plus chemotherapy in the first-line had a duration of response of only 91 days suggests that further work is needed to better identify patients with MBC who may benefit from immunotherapy.
MSI-H testing has primarily been concentrated amongst the cancers associated with Lynch syndrome19. With the expanded use of molecular testing across cancer types, the use of MSI-H as a pan-cancer biomarker is still in its infancy. Notably, the FDA approved the use of pembrolizumab across solid tumors for patients with MSI-H metastatic disease and no satisfactory alternative treatments, meaning that patients with MSI-H MBC could qualify for pembrolizumab regardless of their programmed death ligand 1 (PD-L1) status13. However, although the FDA approval applies across solid tumors, many cancer types, including breast, had low or no representation amongst the five KEYNOTE studies used to support this approval26,27. For example, the KEYNOTE-158 trial, which contained the largest cohort of non-colorectal MSI-H cancers, included five patients with breast cancer and did not provide a breast cancer-specific response rate28. While the utility of immune checkpoint inhibitor therapy in some of these cancer types requires further exploration, preliminary data does support the clinical utility of plasma-based NGS testing in identifying MSI-H patients who may benefit from immune checkpoint inhibitor therapy in this setting. In a study of nine patients with metastatic castration-resistant prostate cancer found to be MSI-H via this same liquid biopsy assay who went on to receive pembrolizumab, four patients (44%) achieved a PSA decrease >50% after a median of 4 weeks, three of whom had a PSA decrease >99%27. Among five patients evaluable for response, the overall response rate was 60%, with one complete response and two partial responses27.
There is limited data on how MSI status correlates with PD-L1 expression, the current biomarker used to select immunotherapy for MBC, although pan-cancer studies suggest a relatively low correlation between PD-L1 expression and MSI-H status29. A prior study evaluating tumor tissue biopsies of TNBC did not identify a correlation between MSI status and PD-L1 expression30. Similarly, the overlap between tumor mutational burden (TMB) score and MSI-H has yet to be fully explored in MBC. An analysis done using the same cfDNA assay as in our study compared the frequency of TMB high (defined in a cancer-type-specific manner using the 80th percentile score for mutations/megabase) and MSI-H across multiple cancer types. The frequency of overlap varied significantly, with 7% of breast cancer cases classified as TMB high, also denoted as MSI-H31. Recent studies in breast cancer have demonstrated the preliminary efficacy of immunotherapy in tumors with high mutational burden32,33. Further exploration of the optimal biomarkers for the selection of patients with MBC who may benefit from immune checkpoint inhibitor therapy is needed. The majority of immunotherapy studies in advanced breast cancer have focused on TNBC as it has a higher level of tumor-infiltrating lymphocytes and PD-L1 expression compared to other breast cancer subtypes. The results of these studies have been mixed, with some patients experiencing a sustained duration of response, but overall low objective response rates34. The data available in patients with HR+/HER2− breast cancer is limited, with the KEYNOTE-028 trial of patients with estrogen receptor-positive/HER2- advanced breast cancer and PD-L1 + tumors showing a median progression free survival of 1.8 months and an objective response rate of 12%35. Overall, this data suggests that additional research is needed across breast cancer subtypes to better refine treatment selection, including in patients with HR + /HER2- disease, as demonstrated by the patient with HR+/HER2− advanced breast cancer in our study with a duration of response greater than 245 days on pembrolizumab combination therapy.
The study had a few limitations. First, aside from the 11 patients for whom the ordering provider was willing to share additional clinical data, we have limited clinical data for this cohort. As we were reliant on which providers responded to our inquiry, it is possible the cohort may be biased toward providers more interested in MSI-H and, thus, more likely to give immunotherapy in this setting. Of the 11 patients with clinical data available, only three had tissue testing of a metastatic tumor lesion that included orthogonal confirmation of MSI status, a limitation of the retrospective nature of this study. Given the retrospective nature of this study, we were not able to obtain metastatic tumor tissue for MSI-H testing prospectively to evaluate concordance with the blood-based assay. As this study included patients tested from 2018 to 2020, this lack of tissue MSI-H testing may reflect the historic lack of routine testing for MSI-H conducted in patients with breast cancer. Additionally, the version of the assay these patients were tested with included only partial coverage of MLH1 and did not include PMS2, MSH2, or MSH6. Thus, we cannot comment on the occurrence of co-occurring pathogenic alterations in these genes in this cohort. The rarity of MSI-H in many cancer types, including breast cancer, means that large cohorts for validation of assay performance by cancer type are difficult; there are technically no validated methods for the detection of MMR status in breast cancer22. While the clinical validation cohort used for this assay contained a relatively small number of breast cancer cases, the pan-cancer positive predictive value was 94.7%, and publications examining MSI-H calls made by this assay in other cancer types where MSI-H is rare support the validity of these calls36,37. Further exploration of the optimal methodology for the detection of MSI-H in breast cancer is needed. Second, given the limited sample size and retrospective nature of this study, we were not able to assess the clinical utility of immunotherapy in patients with MSI-H MBC, although a few durable responses were seen in this small cohort of seven patients. A prospective study is needed to determine the clinical utility of immunotherapy treatment in MSI-H MBC detected by plasma-based genotyping.
In summary, our work demonstrates that plasma-based genotyping assays can detect MSI-H in breast cancer, including in patients with TNBC and HR+/HER2− MBC. Additional research is needed to further demonstrate the efficacy of immunotherapy in MSI-H MBC and identify the optimal biomarkers for the selection of patients with MBC who may benefit from immunotherapy combinations.
Methods
Patient population
A retrospective analysis of a de-identified database containing results from consecutive patients with advanced breast cancer treated within the United States who underwent cell-free circulating tumor DNA (cfDNA) analysis with Guardant360® (next-generation sequencing (NGS), up to 74 gene panels) between 9/27/2018 and 3/12/2020 was conducted. As this database is derived from commercial testing, only internal access from select employees of Guardant Health is permitted. From this subset, the cohort with MSI-H in cfDNA analysis with Guardant360® was identified. Patient age, gender, and cancer type were extracted from the test requisition form. This research was approved by the Quorum Institutional Review Board (IRB) for the generation of de-identified datasets for research purposes. The ordering providers for all 42 MSI-H cases were contacted regarding their interest in participating in this study. Providers responded affirmatively for 11 of these patients, and clinical data were provided by the treating physician to evaluate clinical features and response to immunotherapy. All patients in the dataset had consented (written informed consent) to Guardant360® testing by their treating physician.
Cell-free DNA analysis
For this study, cell-free circulating tumor DNA (cfDNA) analysis was performed using the Guardant360® assay (Guardant Health, Redwood City, California, USA). Guardant360® is a CLIA-certified, College of American Pathologists-accredited, New York State Department of Health-approved, cfDNA NGS assay, with analytical and clinical validation previously described38,39. The assay includes analysis of single nucleotide variants (SNVs) in up to 74 genes (the assay evolved from a 73-gene to a 74-gene panel over the course of the study period), as well as small insertions/deletions (indels), copy number amplifications, and gene rearrangements/fusions in a subset of genes39. The genes included in this assay are shown in Supplementary Table 1.
The analytical performance of MSI-H determination via the Guardant360® cfDNA assay has previously been described40. Briefly, NGS reads from cfDNA across 90 microsatellite loci are integrated into the MSI-H caller, and samples with a total number of unstable sites exceeding a pre-determined threshold are denoted as MSI-H. Clinical validation of this assay for MSI-H determination using 949 unique patients across 40 different cancer types demonstrated a sensitivity of 86.6% and a specificity of 99.5% in samples with a maximum variant allele fraction in cfDNA ≥0.2%40. We note that this clinical validation cohort did rely more heavily on cancers where MSI is more frequent (e.g., colorectal cancer) and included 30 breast cancer samples. However, the positive predictive value of the assay when compared to tissue was 94.7% across cancer types, and publications examining MSI-H calls made by this assay in other cancer types where MSI-H is rare support the validity of these calls36,37.
Statistical analysis
A Mann–Whitney U-test was used to compare the median number of alterations per sample and a median maximum mutant allelic fraction (MAF) for the MSI-H cohort and non-MSI-H samples during the study period. A p value less than 0.05 was considered statistically significant. All statistical analyses were done using GraphPad Prism version 9.1.0 for macOS, GraphPad Software, San Diego, California, USA, www.graphpad.com.
Data availability
The dataset generated and/or analyzed for the current study are not publicly available as they are derived from commercial testing and can only be shared under a fully executed data use agreement. Inquiries regarding such data use agreements can be made to: medicalaffairs@guardanthealth.com.
References
Malla, S. B. et al. In-depth clinical and biological exploration of DNA damage immune response as a biomarker for oxaliplatin use in colorectal cancer. Clin. Cancer Res. 27, 288–300 (2021).
Bonneville, R. et al. Landscape of microsatellite instability across 39 cancer types. JCO Precis. Oncol. 2017, PO.17.00073 (2017).
Hause, R. J., Pritchard, C. C., Shendure, J. & Salipante, S. J. Classification and characterization of microsatellite instability across 18 cancer types. Nat. Med. 22, 1342–1350 (2016).
Long, D. R., Waalkes, A., Panicker, V. P., Hause, R. J. & Salipante, S. J. Identifying optimal loci for the molecular diagnosis of microsatellite instability. Clin. Chem. 66, 1310–1318 (2020).
Markowitz, S. D. & Bertagnolli, M. M. Molecular origins of cancer: molecular basis of colorectal cancer. N. Engl. J. Med. 361, 2449–2460 (2009).
Kim, G. P. et al. Prognostic and predictive roles of high-degree microsatellite instability in colon cancer: a National Cancer Institute-National Surgical Adjuvant Breast and Bowel Project Collaborative Study. J. Clin. Oncol. 25, 767–772 (2007).
Halvarsson, B., Anderson, H., Domanska, K., Lindmark, G. & Nilbert, M. Clinicopathologic factors identify sporadic mismatch repair-defective colon cancers. Am. J. Clin. Pathol. 129, 238–244 (2008).
Hampel, H. et al. Feasibility of screening for Lynch syndrome among patients with colorectal cancer. J. Clin. Oncol. 26, 5783–5788 (2008).
Aaltonen, L. A. et al. Incidence of hereditary nonpolyposis colorectal cancer and the feasibility of molecular screening for the disease. N. Engl. J. Med. 338, 1481–1487 (1998).
Le, D. T. et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science 357, 409–413 (2017).
Le, D. T. et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med. 372, 2509–2520 (2015).
Marabelle, A. et al. Association of tumour mutational burden with outcomes in patients with advanced solid tumours treated with pembrolizumab: prospective biomarker analysis of the multicohort, open-label, phase 2 KEYNOTE-158 study. Lancet Oncol. 21, 1353–1365 (2020).
FDA., U. S. FDA grants accelerated approval to pembrolizumab for first tissue/site agnostic indication. https://www.fda.gov/drugs/resources-information-approved-drugs/fda-grants-accelerated-approval-pembrolizumab-first-tissuesite-agnostic-indication (2017).
Vidula, N. et al. Tumor tissue- versus plasma-based genotyping for selection of matched therapy and impact on clinical outcomes in patients with metastatic breast cancer. Clin. Cancer Res. 27, 3404–3413 (2021).
Meador, C. B. et al. High sensitivity of plasma cell-free DNA genotyping in cases with evidence of adequate tumor content. JCO Precis. Oncol. https://doi.org/10.1200/po.20.00420 (2021).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013).
André, F. et al. Alpelisib plus fulvestrant for PIK3CA-mutated, hormone receptor-positive, human epidermal growth factor receptor-2-negative advanced breast cancer: final overall survival results from SOLAR-1. Ann. Oncol. 32, 208–217 (2021).
Latham, A. et al. Microsatellite instability is associated with the presence of Lynch syndrome pan-cancer. J. Clin. Oncol. 37, 286–295 (2019).
Trabucco, S. E. et al. A novel next-generation sequencing approach to detecting microsatellite instability and pan-tumor characterization of 1000 microsatellite instability-high cases in 67,000 patient samples. J. Mol. Diagn. 21, 1053–1066 (2019).
Vanderwalde, A., Spetzler, D., Xiao, N., Gatalica, Z. & Marshall, J. Microsatellite instability status determined by next-generation sequencing and compared with PD-L1 and tumor mutational burden in 11,348 patients. Cancer Med. 7, 746–756 (2018).
Fusco, N. et al. Mismatch repair protein loss as a prognostic and predictive biomarker in breast cancers regardless of microsatellite instability. JNCI Cancer Spectr. 2, pky056 (2018).
Bertucci, F. et al. Genomic characterization of metastatic breast cancers. Nature 569, 560–564 (2019).
Barroso-Sousa, R. et al. Prevalence and mutational determinants of high tumor mutation burden in breast cancer. Ann. Oncol. 31, 387–394 (2020).
Jia, Y. et al. Tumor mutation burden and immune microenvironment analysis of urothelial carcinoma. J. Clin. Oncol. 39, 2021 (2021).
Marcus, L., Lemery, S. J., Keegan, P. & Pazdur, R. FDA approval summary: pembrolizumab for the treatment of microsatellite instability-high solid tumors. Clin. Cancer Res. 25, 3753–3758 (2019).
Broderick, J. M. Pembrolizumab approved by the FDA for microsatellite instability-high and mismatch repair deficient cancers. https://www.targetedonc.com/view/pembrolizumab-approved-by-the-fda-for-microsatellite-instabilityhigh-and-mismatch-repair-deficient-cancers (2017).
Marabelle, A. et al. Efficacy of pembrolizumab in patients with noncolorectal high microsatellite instability/mismatch repair-deficient cancer: results from the phase II KEYNOTE-158 study. J. Clin. Oncol. 38, 1–10 (2020).
Luchini, C. et al. ESMO recommendations on microsatellite instability testing for immunotherapy in cancer, and its relationship with PD-1/PD-L1 expression and tumour mutational burden: a systematic review-based approach. Ann. Oncol. 30, 1232–1243 (2019).
Ren, X. Y. et al. Mismatch repair deficiency and microsatellite instability in triple-negative breast cancer: a retrospective study of 440 patients. Front. Oncol. 11, 570623 (2021).
Drusbosky, L. et al. Blood-based tumor mutational burden from circulating tumor DNA (ctDNA) across advanced solid malignancies using a commercially available liquid biopsy assay. J. Clin. Oncol. 39, 3040–3040 (2021).
Barroso-Sousa, R. et al. Nimbus: A phase 2 trial of nivolumab plus ipilumumab for patients with hypermutated her2-negative metastatic breast cancer (MBC). Cancer Res. 82, GS2–10 (2022).
Winer, E. P. et al. Association of tumor mutational burden (TMB) and clinical outcomes with pembrolizumab (pembro) versus chemotherapy (chemo) in patients with metastatic triple-negative breast cancer (mTNBC) from KEYNOTE-119. J. Clin. Oncol. 38, 1013–1013 (2020).
Howard, F. M., Pearson, A. T. & Nanda, R. Clinical trials of immunotherapy in triple-negative breast cancer. Breast Cancer Res. Treat. 195, 1–15 (2022).
Rugo, H. S. et al. Safety and antitumor activity of pembrolizumab in patients with estrogen receptor-positive/human epidermal growth factor receptor 2-negative advanced breast cancer. Clin. Cancer Res. 24, 2804–2811 (2018).
Barata, P. et al. Clinical activity of pembrolizumab in metastatic prostate cancer with microsatellite instability high (MSI-H) detected by circulating tumor DNA. J. Immunother. Cancer https://doi.org/10.1136/jitc-2020-001065 (2020).
Chakrabarti, S. et al. Detection of microsatellite instability-high (MSI-H) by liquid biopsy predicts robust and durable response to immunotherapy in patients with pancreatic cancer. J. Immunother. Cancer https://doi.org/10.1136/jitc-2021-004485 (2022).
Odegaard, J. I. et al. Validation of a plasma-based comprehensive cancer genotyping assay utilizing orthogonal tissue- and plasma-based methodologies. Clin. Cancer Res. 24, 3539–3549 (2018).
Zill, O. A. et al. The landscape of actionable genomic alterations in cell-free circulating tumor DNA from 21,807 advanced cancer patients. Clin. Cancer Res. 24, 3528–3538 (2018).
Willis, J. et al. Validation of microsatellite instability detection using a comprehensive plasma-based genotyping panel. Clin. Cancer Res. 25, 7035–7045 (2019).
Acknowledgements
No funding was provided for this study, although two of the co-authors (C.W. and K.P.) are employees of Guardant Health, as noted in the competing interests section.
Author information
Authors and Affiliations
Contributions
N.V.: Designed study, wrote the first draft of the manuscript, revised manuscript, data collection, interpretation, analysis, review, and approval of the final manuscript. A.L., S.K., K.H., G.A., A.E., D.J., E.R., C.F., P.M., K.P., and J.S.: Data collection, interpretation, analysis, and review and approval of the final manuscript. C.W.:Data collection, interpretation, analysis, and review and approval of the final manuscript. Writing/revising of the manuscript. A.B.: Designed the study, revised manuscript, data collection, interpretation, analysis, and review and approval of the final manuscript.
Corresponding author
Ethics declarations
Competing interests
Individual disclosures for co-authors are as noted below: A.L.: The author declares no competing financial and non-financial interests. S.K.: The author declares no competing financial and non-financial interests. K.H.: The author declares no competing financial and non-financial interests. A.E.: The author declares no competing financial and non-financial interests. E.R.: The author declares no competing financial and non-financial interests. CF: The author declares no competing financial and non-financial interests. P.M.: The author declares no competing financial and non-financial interests. N.V.: The author declares no competing non-financial interests but the following competing financial interests: Research funding to the institution (Massachusetts General Hospital): Merck, Daehwa, Pfizer, Radius, and Novartis; Advisory board participation (single session): AbbVie, OncoSec, Aadi, Gilead. D.J.: The author declares no competing non-financial interests but the following competing financial interests: Consulting fees from Novartis, Genentech, Syros, Eisai, Vibliome, Mapkure, and Relay Therapeutics. Contracted research with Novartis, Genentech, Syros, Pfizer, Eisai, Takeda, Ribon Therapeutics, Infinity, InventisBio, Cyteir, Blueprint Medicines, and Arvinas. Ownership interest in Relay Therapeutics and PIC Therapeutics. C.W.: The author declares no competing non-financial interests but the following competing financial interests: Employee and stockholder of Guardant Health. G.A.: The author declares no competing non-financial interests but the following competing financial interests: Speaker bureau for Guardant Health and Natera. K.P.: The author declares no competing non-financial interests but the following competing financial interests: Employee and stockholder of Guardant Health JS: The author declares no competing non-financial interests but the following competing financial interests: Honoria for consulting and/or advisory board: AbbVie Inc., Agendia, Amgen Biotechnology, Aptitude Health, AstraZeneca, Bayer, Bristol-Myers Squibb, Carrick Therapeutics, Celgene Corporation, Clovis Oncology, Daiichi Sankyo, Eisai, GI Therapeutics, Genentech, Gilead Sciences, GRAIL, Halozyme Therapeutics, Heron Therapeutics, Immunomedics, Ipsen Biopharmaceuticals, Lilly, Merck, Myriad, Nektar Therapeutics, Novartis, Ontada, Pfizer, Pharmacyclics, Pierre Fabre Pharmaceuticals, Puma Biotechnology, Prime Oncology, Roche, Samsung Bioepis, Sanofi, Seagen, Syndax Pharmaceuticals, Taiho Oncology, Takeda, Synthon. A.B.: The author declares no competing non-financial interests but the following competing financial interests: Consultant/Advisory board: Pfizer, Novartis, Genentech, Merck, Radius Health, Immunomedics/Gilead, Sanofi, Daiichi Pharma/AstraZeneca, Phillips, Eli Lilly, Foundation Medicine. Contracted Research/Grant (to institution): Genentech, Novartis, Pfizer, Merck, Sanofi, Radius Health, Immunomedics/Gilead, Daiichi Pharma/AstraZeneca, Eli Lilly.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vidula, N., Lipman, A., Kato, S. et al. Detection of microsatellite instability high (MSI-H) status by targeted plasma-based genotyping in metastatic breast cancer. npj Breast Cancer 8, 117 (2022). https://doi.org/10.1038/s41523-022-00490-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41523-022-00490-2
This article is cited by
-
Pembrolizumab response in stage IV luminal-type breast cancer with high microsatellite instability: a case report
Journal of Medical Case Reports (2024)