Abstract
Circulating tumor cells (CTCs) and circulating tumor DNA (ctDNA) are two cancer-derived blood biomarkers that inform on patient prognosis and treatment efficacy in breast cancer. We prospectively evaluated the clinical validity of quantifying both CTCs (CellSearch) and ctDNA (targeted next-generation sequencing). Their combined value as prognostic and early monitoring markers was assessed in 198 HER2-negative metastatic breast cancer patients. All patients were included in the prospective multicenter UCBG study COMET (NCT01745757) and treated by first-line chemotherapy with weekly paclitaxel and bevacizumab. Blood samples were obtained at baseline and before the second cycle of chemotherapy. At baseline, CTCs and ctDNA were respectively detected in 72 and 74% of patients and were moderately correlated (Kendall’s τ = 0.3). Only 26 (13%) patients had neither detectable ctDNA nor CTCs. Variants were most frequently observed in TP53 and PIK3CA genes. KMT2C/MLL3 variants detected in ctDNA were significantly associated with a lower CTC count, while the opposite trend was seen with GATA3 alterations. Both CTC and ctDNA levels at baseline and after four weeks of treatment were correlated with survival. For progression-free and overall survival, the best multivariate prognostic model included tumor subtype (triple negative vs other), grade (grade 3 vs other), ctDNA variant allele frequency (VAF) at baseline (per 10% increase), and CTC count at four weeks (≥5CTC/7.5 mL). Overall, this study demonstrates that CTCs and ctDNA have nonoverlapping detection profiles and complementary prognostic values in metastatic breast cancer patients. A comprehensive liquid-biopsy approach may involve simultaneous detection of ctDNA and CTCs.
Similar content being viewed by others
Introduction
Liquid biopsy, a minimally invasive approach, has shown in the last decades prominent validity for diagnosis, prognosis, and for monitoring treatment efficacy in different cancer types. Throughout the years, several studies have explored the validity of different circulating analytes, including circulating tumor cells (CTCs), circulating tumor DNA (ctDNA), cell-free RNAs (mRNAs, long noncoding RNAs, and microRNAs), extracellular vesicles, and proteins markers. However, CTCs and ctDNA remain the most widely studied circulating biomarkers so far.
CTCs, released from the primary tumor or metastatic lesions, can be detected by immunocytological techniques1 and have been reported to play a key role in cancer metastasis through intravasation into the bloodstream and dissemination at distant sites2. In early-stage breast cancer (BC) patients, CTC enumeration has been used as an independent and powerful biomarker for survival and relapse prognostication3,4. This was also demonstrated in patients with metastatic breast cancer (mBC) where a decrease in CTC counts 3–5 weeks after the start of a new line of therapy correlates with treatment efficacy5,6,7.
ctDNA, a fraction of total cell-free DNA (cfDNA), is released into the circulation by tumor cells undergoing apoptosis/necrosis or by active secretion and its biological role is currently not known8. Using next-generation sequencing (NGS) or droplet-based digital PCR (ddPCR) technologies, multiple studies have demonstrated the utility of ctDNA detection as a prognostic biomarker for patients’ survival in early-stage and metastatic BC patients9,10,11. Unlike tumor tissue biopsies, ctDNA analysis is spatially unbiased, which allows systematic investigation of tumor molecular alterations and improves the detection of potential actionable mutations. ctDNA profiling has been successfully used in breast cancer patients to explore intratumor clonal heterogeneity and early disease evolution12,13,14 or as a diagnostic tool for treatment selection15,16. The use of CTCs and ctDNA biomarkers, therefore, provides real-time assessment of tumor dynamics and represents a major tool for selecting the best therapy and for monitoring treatment efficacy17.
Until now, the clinical significance of CTCs and ctDNA biomarkers has mostly been assessed independently and only a limited number of studies have quantified both biomarkers in parallel using the same blood draws from the same patients. In the present study, we have investigated the clinical value of concomitant analysis of CTCs and ctDNA. As part of a large prospective biomarker study (COMET) conducted in HER2-negative mBC patients treated by first-line chemotherapy, CTCs and ctDNA were analyzed at two time points: prior to treatment and after four weeks of first-line chemotherapy. The detection profiles and prognostic values of these two circulating biomarkers were then compared to understand their overlap and complementary contribution in mBC patients’ management.
Results
CTC enumeration and ctDNA detection at baseline and their associations with patients’ characteristics
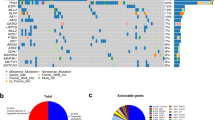
Out of the 510 patients enrolled in the whole COMET study, analyses were performed on the 198 consecutive patients for whom CTC counts and ctDNA measurements were available at baseline (study flow chart in Supplementary Fig. 1). Clinical and pathological characteristics of this subset of 198 patients are displayed in Table 1. At baseline, 142 (72%) patients had one or more CTCs and 98 (49%) patients had five or more CTCs detected in 7.5 mL of blood (median: 4 CTC, range: 0–30,000). Variants were detected in 147 patients (Supplementary Table 1). A total of 251 single-nucleotide variants and 78 small indels were identified among the 54 genes targeted by the NGS approach (Supplementary Data 1). The median number of mutated genes per baseline sample was 1 (range 0–9) in the whole population and two in the 147 patients with detected ctDNA. Fifty-one (26%), 53 (27%), and 94 (47%) patients displayed zero, one, or more than one mutated gene in their baseline ctDNA sample, respectively. Variants were most frequently observed in TP53 and PIK3CA genes (Fig. 1).
Associations between the two blood tumor markers examined as dichotomized variables and patient characteristics are shown in Table 1. In the population of 147 patients with detectable ctDNA (i.e., with at least one variant detected), TP53 variants were more often found in younger, premenopausal patients with triple-negative or grade-3 breast cancers, with a higher number of metastatic sites or synchronous metastasis. PIK3CA variants were more frequent but not restricted to HR-positive cancers and were mostly observed in premenopausal patients. Besides one variant with no known pathogenic function (P-123), ESR1 variants associated with hormone-therapy resistance were restricted to the HR-positive cancers and more frequent in patients with liver metastases.
Moreover, considering blood biomarkers as continuous variables, we detected the associations between high fractions of ctDNA and high tumor grade, triple-negative status, altered performance status, and high number of metastatic sites. CTCs were higher in patients with lobular subtypes, altered performance status, and in the presence of liver or bone metastases (Supplementary Table 2).
Correlation between CTC enumeration and ctDNA detection
In patients with detectable ctDNA (n = 147), a reporter variant was selected to analyze ctDNA-detection profiles and prognostic values in comparison with CTC measurements. The median allelic frequency of reporter variants at baseline was 7.6% (range 0.6–84.8%). CTC counts and ctDNA levels in each patient were moderately correlated (Fig. 2A; Kendall’s τ = 0.3, p < 0.001). At baseline, 82 (41%) patients had detectable ctDNA and five or more CTCs detected, 35 (18%) patients had no detectable ctDNA and less than 5 CTCs detected, and only 26 (13%) had neither ctDNA nor CTCs detected.
The number of CTCs detected in patients with certain mutated genes was then compared with the rest of the population with detectable ctDNA but no variant in the corresponding gene (Supplementary Fig. 2). We observed that the 16 patients with a KMT2C/MLL3 variant had a lower CTC count (median 1.5 CTC, IQR 0–5) than the 131 patients with no KMT2C/MLL3 variant detected (median 10 CTC, IQR 1–59.5, p = 0.007). Twenty patients had a GATA3 variant and demonstrated an opposite trend, with an overall higher CTC count (median: 27 CTC, IQR: 4–74) than the 120 patients with no GATA3 variant detected (median 6 CTCs, IQR 1–40; p = 0.07).
Prognostic value of CTCs and ctDNA at baseline
The prognostic impact of CTCs or ctDNA levels was first assessed at baseline independently. Both biomarkers were considered as dichotomized and continuous variables in univariate analysis. Shorter progression-free survival (PFS) and overall survival (OS) were observed in patients with higher CTC counts (Supplementary Fig. 3A–B; Supplementary Table 5) or higher ctDNA VAFs (Supplementary Fig. 3C–D; Supplementary Table 5). OS and PFS for combined CTC and ctDNA are shown in Figs. 3A and 3C (see also Table S5 for the corresponding CTC counts and ctDNA VAFs).
When considered as continuous variables, baseline ctDNA and CTC levels demonstrated a linear and log-linear impact, respectively, on the patient PFS and OS (Supplementary Fig. 4). There was no significant interaction between VAF impact on survival and individual gene (Supplementary Fig. 5).
Alterations in specific genes were also investigated to evaluate whether they have a prognostic impact on patients’ survival. This analysis was restricted to genes with variants detected in at least 15 patients (TP53, PIK3CA, GATA3, ESR1, KMT2C, and CDH1). We observed that patients with TP53 variants displayed a worse overall survival (Supplementary Fig. 6). This was also significant in the subgroup of patients with HR-positive mBC. On the other hand, the detection of ESR1 variants had no significant prognostic impact in this cohort, mostly composed of endocrine-resistant or HR-negative mBC (data not shown).
Finally, the added prognostic value of both CTCs and ctDNA to a prognostic model based on patient clinical and pathological characteristics was evaluated by different combinations. For PFS, the model included HR status, number of metastatic sites, tumor grade, and metastasis-free interval (CP model #1). For OS, the model included HR status, tumor grade, and metastatic site (visceral vs other, CP model #2). At baseline, the added value of both CTCs and ctDNA was higher when considered as continuous variables (linear or log-linear coding). At four weeks, biomarker levels gave a best fit when considered as dichotomized variables (<5 vs ≥5 for CTC and detectable vs not detectable for ctDNA). For PFS, we found that the increase in the prognostication accuracy of the model was not significant for CTC count at baseline (p = 0.2), while adding ctDNA VAF to the CP model #1 significantly increased the performance of the model (p = 0.004) (Table 2). Interestingly, for OS, both ctDNA and CTC levels at baseline significantly increased the prognostication accuracy of CP model #2 (p < 0.0001 and p = 0.008, respectively).
CTC enumeration and ctDNA detection at four weeks
CTC enumeration and ctDNA levels in patients were obtained after four weeks of treatment (196 patients had CTCs quantified, 191 had ctDNA quantified, and 189 both, Supplementary Fig. 1). Out of the 196 patients with CTCs quantified at four weeks, 73 (37%) had one or more CTCs and 43 (22%) had five or more CTCs per 7.5 mL of blood (median: 0 CTC, range: 0–27,710). Regarding ctDNA, among the 191 patients assessed at four weeks, 124 (65%), 31 (16%), and 36 (19%) patients displayed zero, one, or more than one mutated gene, respectively. In patients with at least one variant detected at baseline, the median allelic frequency of reporter variants at four weeks was 0% (range 0–55%). Among the 189 patients with both CTC and ctDNA quantified at four weeks, 25 (13%) patients had detectable ctDNA and 5 or more CTCs and 94 (50%) had neither ctDNA nor CTCs detected. Overall, the detection rates at four weeks were significantly lower than at baseline for both tumor markers (both p < 0.0001) and were again moderately correlated (Fig. 2B; Kendall’s τ = 0.4, p < 0.001). Supplementary Table 3 depicts the different trajectories in terms of level changes for both biomarkers during the first four weeks of treatment. While CTC count and ctDNA levels had similar trajectories for 82 (43%) patients (both increasing or decreasing, highlighted in blue), an opposite trajectory (one marker becoming lower or undetectable while the other increased, highlighted in yellow) was observed in 28 (15%) patients.
Prognostic value of CTCs and ctDNA after four weeks of treatment
In univariate analysis, CTCs and ctDNA levels at four weeks had a significant prognostic impact on both PFS and OS, when considered separately (Supplementary Fig. 7). OS and PFS for combined parameters are shown in Figs. 3B and 3D. We then evaluated the added prognostic value of CTCs or ctDNA level at four weeks to the optimal baseline prognostic model (based on CP model #1 and baseline ctDNA level for PFS and CP model #2, baseline ctDNA level, and baseline CTC count for OS, as displayed in Table 2). Both significantly increased the prognostication accuracy of the model in the same proportion. Finally, the best prognostic model for PFS included tumor subtype (triple negative vs other), grade (grade 3 vs other), ctDNA VAF at baseline (per 10% increase), and CTC count at four weeks (≥5CTC/7.5 mL) (Table 3).
Bootstrapping revealed moderate optimism of the model, with a decrease of the c-statistic from 0.68 to 0.67 and a calibration slope equal to 0.91. The variables retained in the best prognostic model for OS were tumor subtype, grade, ctDNA VAF at baseline (per 10% increase), and CTC count at four weeks (≥5CTC/7.5 mL). The bootstrapped c-statistic of the OS model was 0.73 (equal to the apparent c-statistic) and the calibration slope was 0.92. Overall, the time-dependent area under the curve (AUC) for 12-month PFS and OS status increased from 0.70 and 0.75 (models with baseline clinical variables only) to 0.76 and 0.85 (full PFS and OS models), respectively.
A normalization of the delta of ctDNA values and CTC counts between baseline and four weeks was also performed with the use of a Min–Max scaling (Supplementary Fig. 8). The HR associated with scaled ctDNA variation (dichotomized at the median) was 1.46–95%, C.I. 1.008–2.112, p = 0.045 for OS and 1.36–95% C.I. 0.97–1.89, p = 0.07 for PFS. The HR associated with scaled CTC count variation (dichotomized at the median) was 1.75–95%, C.I. 1.21–2.51, p = 0.003 for OS and 1.44–95%, C.I. 1.05–1.99, p = 0.03 for PFS.
Discussion
As previously described5,6, CTCs were here detected in the peripheral blood of up to 70–80% of mBC patients. They were also associated with poor PFS and OS in multivariate analysis18,19. As the molecular analysis of CTCs is challenging, many studies concentrated on the enumeration of CTCs20. In contrast, ctDNA can be comprehensively analyzed with a variety of methods on archived plasma samples11,21. The proof-of-concept analysis showing that ctDNA is an informative, inherently specific, and highly sensitive biomarker for mBC was first described by Dawson et al. in 201322. Among the analytes tested, ctDNA provided the earliest marker of treatment response compared with CA15-3 and CTC count in a series of 30 patients. The prognostic value of ctDNA detection was confirmed by a few larger studies23,24. Various techniques for ctDNA detection and quantification have since been reported25,26,27. COMET is the largest prospective study assessing the respective prognostic values of CTCs and ctDNA and their early changes during treatment in 189 mBC patients homogeneously treated with first-line chemotherapy. Prior retrospective studies on cohorts of mBC reported a prognostic impact for either one or both blood biomarkers, but the low patient numbers did not allow concluding on their correlation and clinical validity22,28,29,30. In a larger study, monitoring during treatment was limited31. More recently, this complementarity has been reported in the post-neoadjuvant setting at one time point after surgery32.
In this study, we observed a moderate correlation between CTCs and ctDNA detection at baseline and at four weeks. Interestingly, only 26 patients (13%) had neither ctDNA nor CTCs detected at baseline. This suggests that, similar to serum markers and CTCs18, combining ctDNA and CTCs could increase the number of patients assessable for marker detection and monitoring. The mutational landscape retrieved by ctDNA analysis was consistent with prior reports in HER2-negative BC, PIK3CA and TP53 being the two most frequent variants. As expected, TP53 variants were more often found in younger, premenopausal patients with triple-negative or grade-3 BCs, and with a higher number of metastatic sites. PIK3CA variants were more frequent but not restricted to HR-positive cancers. Activating ESR1 mutations, which are selected as a mechanism of resistance to endocrine therapy33, were restricted to HR-positive cancers, and more frequently observed in patients with liver metastases. The large size of the COMET study allowed investigating associations between the tumor mutational landscape (retrieved by ctDNA analysis) and CTC-detection rates. We found that mBC patients with variants in the lysine (K)-specific methyltransferase 2 C gene (KMT2C, formerly known as mixed-lineage leukemia 3, MLL3, a histone methyltransferase involved in transcriptional coactivation) had a lower CTC count at baseline. KMT2C is frequently mutated in HR-positive breast cancer and associated with shorter PFS under anti-estrogen therapy34. In a gastric model, KMT2C inactivation promoted epithelial-to-mesenchymal transition and acquisition of stem cell-like phenotypes35. This may account for the significantly lower number of CTCs detected by the CellSearch system, which detects EpCAM and cytokeratin-positive epithelial CTCs. In addition, we observed a trend toward a higher CTC count in patients with GATA3-mutated BC (p = 0.07). Such association was previously reported in a smaller study36, and might be related to the putative role of GATA3 in BC cell migration and dissemination37. Finally, in our series, somatic variants in CDH1 were seen in both infiltrating lobular and ductal carcinoma patients, although variants in this gene have been shown to be associated with lobular histology38. Higher CTC counts as a continuous variable were observed in patients with lobular cancers (p = 0.05). However, we did not confirm the correlation between CDH1 variants and CTC count that was previously reported36.
We largely confirmed the validity of CTCs and ctDNA levels as prognostic markers before and during therapy. In addition to standard multivariate analyses, we measured whether one or both blood biomarkers significantly improve the accuracy of multivariate clinicopathological models. This well-established statistical method was already used to demonstrate the superiority of CTC count over serum markers6. Both approaches showed that more accurate prognostic information could be obtained when both blood biomarkers are considered. In detail, the best prognostic models for PFS and OS included ctDNA VAF at baseline (as a continuous variable) and CTC count at four weeks (as a dichotomized variable), highlighting that both blood biomarkers have nonoverlapping and complementary clinical validities. These results raise the question of using both markers for different purposes: (1) detection rates of both CTCs and ctDNA at baseline would help patient-survival prognostication and first-line strategy choice and (2) quantification at four weeks would help monitoring treatment efficacy. For this matter, CTC counts seem to outperform ctDNA. On the other side, ctDNA analysis at inclusion allows to better apprehend the tumor mutational landscape. While baseline CTC count demonstrated its utility in mBC patients as a decision tool before initiating a first-line treatment with chemotherapy or single-agent endocrine therapy39, CTC count monitoring during treatment has not yet demonstrated its clinical utility. In the SWOG 500 trial, patients whose mBC was resistant to first-line chemotherapy (as indicated by persistently high CTC count during therapy) did not benefit from the earlier introduction of a second-line chemotherapy40. Clinical utility of early ctDNA variation monitoring for mBC treatment remains also to be demonstrated10. Consensus recommendations by the American Society of Clinical Oncology have concluded that some ctDNA assays have clinical validity and utility as a noninvasive tool to assess the mutational landscape of a tumor15. However, none of these assays has been fully validated as an early monitoring tool during therapy.
One limitation of our study is that the CellSearch method, as explained above, may not detect CTCs that have lost epithelial features, but no other method has demonstrated its validity on a large scale at this time. Other limitations could be related to the small size of the NGS panel used to analyze ctDNA, potential inclusion of variants arising from clonal hematopoiesis, and limited sensitivity of the assay to detect variants with allelic frequency below 0.5%. However, higher sensitivity would not have changed the limited correlation observed between the CTCs and ctDNA as quantitative variables.
In that context, our study demonstrates that CTCs and ctDNA, which stem from very different biological processes, should be considered as yielding complementary and nonoverlapping clinical information.
Methods
Patients, treatment, and blood sampling
Patients used in this study were included in the multicentric prospective biomarker study “COMET” (NCT01745757) and were treated with weekly paclitaxel and bevacizumab as first-line chemotherapy. As per the 2012 European Medicine Agency label for bevacizumab, patients with triple-negative or hormone-receptor (HR)-positive, HER2-negative mBC were enrolled in this study. Other main inclusion criteria were written informed consent, age >18 years old, performance status (PS) of 0–2, life expectancy ≥3 months, and no prior chemotherapy for mBC. All patients received intravenous paclitaxel 90 mg/m2 on days 1, 8, and 15 with bevacizumab 10 mg/kg on days 1 and 15. Treatment was repeated every four weeks, until disease progression or unacceptable toxicity. The study was approved by an ethics committee (Comité de Protection des Personnes “Ile de France VII”) in June 2012 and registered under the following number NCT01745757. The primary endpoint of the study was the change of circulating endothelial cells (reported elsewhere41) and CTCs during therapy. Patients gave informed consent to use leftover plasma and tumor samples for further research, which allowed ctDNA analysis in this study although not mentioned in the initial research plan. Blood was drawn at baseline (prior to the start of chemotherapy) and at four weeks (prior to the start of the second cycle of treatment). Both markers were quantified in a central laboratory (The Circulating Tumor Biomarkers Laboratory, Institut Curie, France).
CTC enumeration
CTC counts were determined using the standard CellSearch® assay. Blood collected into CellSave® tubes was maintained at room temperature and processed within 96 h by experienced operators. Technical details of the CellSearch® technique (Menarini Silicon Biosystems, Raritan, NJ) have been described elsewhere1.
ctDNA analysis
For plasma isolation and cfDNA extraction, blood collected in EDTA tubes was processed on-site within 2 h of blood draw. Tubes were first centrifuged at 820 g for 10 min at RT. The supernatant was then centrifuged at 16000 g for 10 min at 4 °C and stored at −80 °C. cfDNA was extracted from 1 to 2 mL of plasma using the QIAamp circulating nucleic acid kit (Qiagen), following the manufacturer’s instructions. cfDNA was stored at −20 °C and quantified using a LINE1 real-time PCR assay as previously described (LINE1 forward primer: 5′-TCACTCAAAGCCGCTCAACTAC-3′; LINE1 reverse primer: 5′-TCTGCCTTCATTTCGTTATGTACC-3′)42.
For NGS library construction and sequencing, ctDNA detection was performed by capture-based targeted resequencing using a custom panel of 54 genes (Supplementary Table 1)43,44,45. Libraries were prepared using the Kapa HyperPrep kit (Roche) following the manufacturer’s instructions using xGen Dual Index UMI adapters from IDT, which contain one unique molecular identifier (UMI). Hybridization steps were performed on batches of 12 samples with the SeqCap EZ Accessory Kit V2 (Roche) and a custom panel of SeqCap EZ probes (Roche) covering 220 kb of the human genome. Sequencing was performed on an Illumina HiSeq 2500 instrument in batches of 84 samples using the Rapid Run mode PE 100. Matched archived tumor samples (available for 38 patients) were sequenced on an Illumina MiSeq instrument using a v2 cartridge PE 150.
For sequencing data analysis, raw reads were demultiplexed following IDT UMI recommendations and mapped to the hg19 human genome using bwa v0.7.15 (mem algorithm, default parameters46). UMI consensus sequences were generated and filtered using fgbio v0.5.0a-0 (com.fulcrumgenomics) with the following parameters: minimum mapping quality of 20 to keep an alignment, one error allowed per UMI. All UMIs supported by at least one read were selected and mapped to the reference genome. Only alignments intersecting the targeted sequence were kept using BedTools (v2.21.047). Sequencing metrics for all libraries are described in Supplementary Data 2. Average coverage and consensus depths for cfDNA libraries were 2588X (range 121–9919) and 437X (range 6–1697), respectively. Average coverage for FFPE samples was 286X (range 58–616). Varscan2 (v2.4.148) was then used to call both SNVs and indels in the alignment files considering or without considering UMI consensus sequences. For FFPE samples, UMIs were not considered. Only variants with an allelic frequency higher than 0.1% at a locus covered by at least eight reads were reported (mapping quality at 20 and base quality at 13). For FFPE samples, a minimum threshold of 5% VAF was used to filter out the variants. Variants were then annotated using Annovar (July 2017 version49) and the following annotations: refGene (October 2015 version), dbsnp147 and 1000 G (08/2015 version, all), Exome Sequencing Project (ESP6500, all), ExAc project, Cosmic (v70), ICGC (v21), and prediction algorithms such as SIFT, PolyPhen2, LRT, MutationTaster, PhyloP, and CADD (version ljb26). No technical filtering (based on metrics linked to the variants) was applied to cfDNA samples under the assumption that the confidence in variants called through the consensus of UMIs is high. The following filters were applied: (i) keep only exonic or splicing variant, (ii) discard variants frequent in the population (in at least one database with a VAF higher than 1%), and (iii) discard synonymous variants. A final curated list of nonsynonymous variants was obtained by manual filtering after visual inspection using IGV50. To increase the sensitivity of our approach (see below), whenever a variant was identified in FFPE tumor tissue, the same variant was systematically searched at baseline and week-four plasma samples with and without considering the UMIs. The same approach was applied to week four samples for variants identified in baseline plasma samples considering the UMIs. For patients with multiple variants detected in their cfDNA, a reporter variant was selected for monitoring. Prioritization was given in this order to variants (1) with positivity at baseline, (2) with positivity at four weeks, (3) with a COSMIC ID, (4) which were SNVs, and (5) presenting the highest VAF at baseline. All evaluations were carried out by qualified personnel with no knowledge of the patient’s clinical status.
Three quality controls of the ctDNA-detection assay were used to assess the sensitivity and the specificity of the custom-targeted resequencing approach.
(1) First, Seraseq™ ctDNA Mutation Mix v2 reference material, including 40 mutations in 28 genes, was tested, among which 12 (7 SNVs, three deletions, and two insertions) were covered by the panel (PIK3CA, PTEN, KRAS, TP53, and GNAS). Thirty-six samples comprising three replicates of Seraseq™ ctDNA Mutation Mix v2 at 0% (WT), 0.5, 1, and 2% VAF and at 3 different input quantities (8, 16, and 24 ng) were sequenced on an Illumina HiSeq 2500 instrument as described above. Average coverage was 3655X (3247–4226X) and consensus depth for UMIs was 817X (range 630–1036X). Average sensitivity was 97% at 2% VAF, 55% at 1% VAF, and 9% at 0.5% VAF for the UMI approach. When UMIs were not considered, average sensitivity was increased to 100% at 2% VAF, 94% at 1% VAF, and 77% at 0.5% VAF. Specificity was then assessed by quantifying detection rates for the same 12 variants in the WT reference material. We observed 100% and 97% specificity for the approach considering and not considering UMIs, respectively.
(2) Second, the detection rates of variants reported in Supplementary Data 1 were assessed in cfDNA obtained from 13 healthy individuals purchased from EFS (Etablissement francais du sang, France). Only 10 of the 329 variants found in patients were also detected in healthy controls. However, the higher VAFs observed in patient samples (average = 8.16%, median = 4.29% in patients vs average = 0.77%, median = 0.5% in healthy controls), together with other supporting data such as detection at the second time point or in tumor tissue, provided enough confidence for considering those variants as true-positive calls in patients samples.
(3) Finally, archived tumor material of patients, who gave written informed consent and treated at Institut Curie (available for 38 patients), was retrospectively sequenced to evaluate if variants identified in plasma samples were also observed in the tumor. These formalin-fixed paraffin-embedded (FFPE) tissues were macrodissected to increase tumor cell content prior to DNA extraction. DNA was extracted with the DNA FFPE Tissue Kit (Qiagen) and sequenced with the same targeted NGS method. Out of the 25 samples for which we obtained good-quality data, 24 (96%) displayed all or at least half of the variants observed in plasma DNA (Supplementary Data 1; P-94 and P-52 had 1/2 variant detected, P-129 and P-142 had 3/4 variants detected). The only patient (P-52) with no match between plasma and tissue had a single ESR1 (p.D538G) variant detected in ctDNA, which was most likely absent from the primary tumor as these alterations are frequently acquired in patients under hormone therapy. The one variant not detected in P-142 was also in ESR1 (p.D538G).
Statistics
Associations between patient characteristics and blood biomarker levels when considered as dichotomized variables were studied using chi-square or Fischer’s exact test for categorical variables and Student’s t-test for age. When considered as continuous variables, Mann–Whitney, Kruskall–Wallis, or Student’s t-test were used for continuous data and Pearson correlation test for age. Associations between patient characteristics and TP53, PIK3CA, or ESR1 gene variant status were studied using chi-square or Fisher’s exact test for categorical variables and Student’s t-test for age.
The correlation between CTC and ctDNA levels was assessed with the Kendall’s tau rank correlation. The comparison of the distribution of CTC levels according to the mutated gene status was performed with the Mann–Whitney test. We studied the genes for which variants were present in more than 10 patients (eight genes). No adjustments for multiple comparisons were made. Differences between baseline and week-four biomarker levels were studied with the use of the Wilcoxon signed-rank test.
PFS was defined as the time to disease progression or death of any cause. Overall survival was defined as the time to death, whatever the cause. Both PFS and OS were measured from the inclusion or the date of second sampling (four weeks) for the analyses based on biomarker levels at four weeks (landmark analyses). PFS and OS curves based on separate or combined CTC and ctDNA values were estimated by the Kaplan–Meier method.
The added value of biomarkers was assessed by comparing log likelihoods of models with and without biomarkers. We first fitted a model with baseline clinical and pathological characteristics (CP model) using a multivariable Cox regression analysis. We followed a backward stepwise-selection procedure. The stopping rule for exclusion of predictors was based on the Akaike information criterion (AIC). The variables tested in the model were age (<50 vs ≥50 years), performance status (0 vs ≥1), tumor subtype, tumor grade (≤2 vs 3), number of metastatic sites (≤2 vs 3), metachronous relapses, and presence of visceral metastases. The CP model was then compared with models in which biomarker levels were added (Table 2) by means of the Log-Likelihood ratio test. For this comparison, the added value of biomarkers was studied in two ways: after dichotomization (<5 vs ≥5 for CTC and detectable vs not detectable for ctDNA) and using restricted cubic splines (RCS) with four knots. For CTC, a log transformation was performed to deal with the skewed distribution. When RCS was used, we tested for nonlinearity in the biomarker effect. If no improvement in fit was found, the linear coding was used. To determine which form of the biomarker (continuous or dichotomized) should be kept in the model, models were compared with AIC. Optimal prognostic models were evaluated by conducting a bootstrap validation of model performance. Discrimination was assessed with the c statistic and calibration with the calibration slope. All tests were two-sided. A P value of less than 0.05 was considered to indicate statistical significance. All statistical analyses were done with R software, version 3.4.2 (R Foundation for Statistical Computing).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The ctDNA sequencing data are available in the NCBI Sequence Read Archive (SRA) under accession number PRJNA745047. PFS, OS data, CTC counts, and ctDNA values for individual participant after de-identification are publicly available in Supplementary Data 3 and Rdata. The sequence metrics used for ctDNA analysis are available in Supplementary Data 2.
Code availability
The source R codes for the figures are available for nonprofit research use as Supplementary software.
References
Allard, W. J. et al. Tumor cells circulate in the peripheral blood of all major carcinomas but not in healthy subjects or patients with nonmalignant diseases. Clin. Cancer Res. 10, 6897–6904 (2004).
Pantel, K. & Alix-Panabieres, C. Liquid biopsy and minimal residual disease - latest advances and implications for cure. Nat. Rev. Clin. Oncol. 16, 409–424 (2019).
Bidard, F. C. et al. Circulating tumor cells in breast cancer patients treated by neoadjuvant chemotherapy: a meta-analysis. J. Natl Cancer Inst. 110, 560–567 (2018).
Thery, L. et al. Circulating tumor cells in early breast cancer. JNCI Cancer Spectr. 3, pkz026 (2019).
Cristofanilli, M. et al. The clinical use of circulating tumor cells (CTCs) enumeration for staging of metastatic breast cancer (MBC): international expert consensus paper. Crit. Rev. Oncol. Hematol. 134, 39–45 (2019).
Bidard, F. C. et al. Clinical validity of circulating tumour cells in patients with metastatic breast cancer: a pooled analysis of individual patient data. Lancet Oncol. 15, 406–414 (2014).
Magbanua, M. J. M. et al. Serial analysis of circulating tumor cells in metastatic breast cancer receiving first-line chemotherapy. J. Natl Cancer Inst. 113, 443–452 (2021).
Cabel, L. et al. Clinical potential of circulating tumour DNA in patients receiving anticancer immunotherapy. Nat. Rev. Clin. Oncol. 15, 639–650 (2018).
Hrebien, S. et al. Early ctDNA dynamics as a surrogate for progression-free survival in advanced breast cancer in the BEECH trial. Ann. Oncol. 30, 945–952 (2019).
O’Leary, B. et al. Early circulating tumor DNA dynamics and clonal selection with palbociclib and fulvestrant for breast cancer. Nat. Commun. 9, 896 (2018).
Heitzer, E., Haque, I. S., Roberts, C. E. S. & Speicher, M. R. Current and future perspectives of liquid biopsies in genomics-driven oncology. Nat. Rev. Genet. 20, 71–88 (2019).
Garcia-Murillas, I. et al. Mutation tracking in circulating tumor DNA predicts relapse in early breast cancer. Sci. Transl. Med. 7, 302ra133 (2015).
De Mattos-Arruda, L. et al. Capturing intra-tumor genetic heterogeneity by de novo mutation profiling of circulating cell-free tumor DNA: a proof-of-principle. Ann. Oncol. 29, 2268 (2018).
Garcia-Murillas, I. et al. Assessment of molecular relapse detection in early-stage breast cancer. JAMA Oncol. 5, 1473–1478 (2019).
Merker, J. D. et al. Circulating tumor DNA analysis in patients with cancer: American Society of Clinical Oncology and College of American Pathologists Joint Review. J. Clin. Oncol. 36, 1631–1641 (2018).
Corcoran, R. B. & Chabner, B. A. Application of cell-free DNA analysis to cancer treatment. N. Engl. J. Med. 379, 1754–1765 (2018).
Mattox, A. K. et al. Applications of liquid biopsies for cancer. Sci. Transl. Med. 11, eaay1984 (2019).
Bidard, F. C. et al. Assessment of circulating tumor cells and serum markers for progression-free survival prediction in metastatic breast cancer: a prospective observational study. Breast Cancer Res. 14, R29 (2012).
Pierga, J. Y. et al. High independent prognostic and predictive value of circulating tumor cells compared with serum tumor markers in a large prospective trial in first-line chemotherapy for metastatic breast cancer patients. Ann. Oncol. 23, 618–624 (2012).
Schochter, F. et al. Are circulating tumor cells (CTCs) ready for clinical use in breast cancer? An overview of completed and ongoing trials using CTCs for clinical treatment decisions. Cells 8, 1412 (2019).
Bidard, F. C., Pierga, J. Y., Soria, J. C. & Thiery, J. P. Translating metastasis-related biomarkers to the clinic-progress and pitfalls. Nat. Rev. Clin. Oncol. 10, 169–179 (2013).
Dawson, S. J. et al. Analysis of circulating tumor DNA to monitor metastatic breast cancer. N. Engl. J. Med. 368, 1199–1209 (2013).
Lee, J. H., Jeong, H., Choi, J. W., Oh, H. E. & Kim, Y. S. Liquid biopsy prediction of axillary lymph node metastasis, cancer recurrence, and patient survival in breast cancer: a meta-analysis. Medicine 97, e12862 (2018).
Yang, J. et al. Predictive value of circulating cell-free DNA in the survival of breast cancer patients: a systemic review and meta-analysis. Medicine 97, e11417 (2018).
Stover, D. G. et al. Association of cell-free DNA tumor fraction and somatic copy number alterations with survival in metastatic triple-negative breast cancer. J. Clin. Oncol. 36, 543–553 (2018).
Pairawan, S. et al. Cell-free circulating tumor DNA variant allele frequency associates with survival in metastatic cancer. Clin. Cancer Res. 26, 1924–1931 (2020).
Kruger, D. T. et al. High ctDNA molecule numbers relate with poor outcome in advanced ER+, HER2- postmenopausal breast cancer patients treated with everolimus and exemestane. Mol. Oncol. 14, 490–503 (2020).
Madic, J. et al. Circulating tumor DNA and circulating tumor cells in metastatic triple negative breast cancer patients. Int J. Cancer 136, 2158–2165 (2015).
Suppan, C. et al. Untargeted assessment of tumor fractions in plasma for monitoring and prognostication from metastatic breast cancer patients undergoing systemic treatment. Cancers 11, 1171 (2019).
Rossi, G. et al. Cell-free DNA and circulating tumor cells: comprehensive liquid biopsy analysis in advanced breast cancer. Clin. Cancer Res. 24, 560–568 (2018).
Ye, Z. et al. Association of clinical outcomes in metastatic breast cancer patients with circulating tumour cell and circulating cell-free DNA. Eur. J. Cancer 106, 133–143 (2019).
Radovich, M. et al. Association of circulating tumor DNA and circulating tumor cells after neoadjuvant chemotherapy with disease recurrence in patients with triple-negative breast cancer: preplanned secondary analysis of the BRE12-158 randomized clinical trial. JAMA Oncol. 6, 1410–1415 (2020).
Carausu, M. et al. ESR1 mutations: a new biomarker in breast cancer. Expert Rev. Mol. Diagn. 19, 599–611 (2019).
Gala, K. et al. KMT2C mediates the estrogen dependence of breast cancer through regulation of ERalpha enhancer function. Oncogene 37, 4692–4710 (2018).
Cho, S. J. et al. KMT2C mutations in diffuse-type gastric adenocarcinoma promote epithelial-to-mesenchymal transition. Clin. Cancer Res. 24, 6556–6569 (2018).
Davis, A. A. et al. Association of a novel circulating tumor DNA next-generating sequencing platform with circulating tumor cells (CTCs) and CTC clusters in metastatic breast cancer. Breast Cancer Res. 21, 137 (2019).
Kouros-Mehr, H. et al. GATA-3 links tumor differentiation and dissemination in a luminal breast cancer model. Cancer Cell 13, 141–152 (2008).
Desmedt, C. et al. Genomic characterization of primary invasive lobular breast cancer. J. Clin. Oncol. 34, 1872–1881 (2016).
Bidard, F. C. et al. Efficacy of circulating tumor cell count-driven vs clinician-driven first-line therapy choice in hormone receptor-positive, ERBB2-negative metastatic breast cancer: the STIC CTC randomized clinical trial. JAMA Oncol. 7, 34–41 (2021).
Smerage, J. B. et al. Circulating tumor cells and response to chemotherapy in metastatic breast cancer: SWOG S0500. J. Clin. Oncol. 32, 3483–3489 (2014).
Vasseur, A. et al. Prognostic value of CEC count in HER2-negative metastatic breast cancer patients treated with bevacizumab and chemotherapy: a prospective validation study (UCBG COMET). Angiogenesis 23, 193–202 (2020).
Rago, C. et al. Serial assessment of human tumor burdens in mice by the analysis of circulating DNA. Cancer Res. 67, 9364–9370 (2007).
Curtis, C. et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486, 346–352 (2012).
Rheinbay, E. et al. Recurrent and functional regulatory mutations in breast cancer. Nature 547, 55–60 (2017).
Ciriello, G. et al. Comprehensive molecular portraits of invasive lobular breast cancer. Cell 163, 506–519 (2015).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Koboldt, D. C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Robinson, J. T., Thorvaldsdottir, H., Wenger, A. M., Zehir, A. & Mesirov, J. P. Variant review with the integrative genomics viewer. Cancer Res. 77, e31–e34 (2017).
Acknowledgements
We thank Sybille Everhardt and Sandrine Marques for supervising and managing this research project. We thank Mathieu Chicard for his precious help during NGS library optimization. The COMET study was funded by Roche France and promoted by Unicancer. High-throughput sequencing has been performed by the ICGex NGS platform of the Institut Curie supported by the grants ANR-10-EQPX-03 (Equipex) and ANR-10-INBS-09–08 (France Génomique Consortium) from the Agence Nationale de la Recherche (“Investissements d’Avenir” program) and by the Canceropole Ile-de-France. The circulating tumor biomarkers laboratory is supported by the SiRIC 2 Curie program (SiRIC Grant «INCa-DGOS-Inserm_12554»).
Author information
Authors and Affiliations
Contributions
A.B.S., F.-C.B., C.P. and J.-Y.P. designed the study, interpreted the results, and wrote the paper. V.L., O.T., C.B., W.J., A.G., M.D., C.L., J.-M.F., C.J., T.M., M.-A.M-R. and F.D. participated in the COMET study enrollment and analysis. J.L. supervised the COMET study and biomarker collection. A.B.S., B.A. and S.B. contributed to NGS experiments. E.G. performed the bioinformatics analyses. C.H. and A.R. assisted with the CTC analyses. M.-L.T. and F.B. curated the clinical database and performed the statistical analyses. All authors contributed to paper revision and review.
Corresponding author
Ethics declarations
Competing interests
F.-C.B. and J.-Y.P. received research and travel grants from Menarini Silicon Biosystems and Roche and received lecture fees by Roche. The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bortolini Silveira, A., Bidard, FC., Tanguy, ML. et al. Multimodal liquid biopsy for early monitoring and outcome prediction of chemotherapy in metastatic breast cancer. npj Breast Cancer 7, 115 (2021). https://doi.org/10.1038/s41523-021-00319-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41523-021-00319-4
This article is cited by
-
Benchmarking UMI-aware and standard variant callers for low frequency ctDNA variant detection
BMC Genomics (2024)
-
c-MET-positive circulating tumor cells and cell-free DNA as independent prognostic factors in hormone receptor-positive/HER2-negative metastatic breast cancer
Breast Cancer Research (2024)
-
Liquid biopsy for breast cancer and other solid tumors: a review of recent advances
Breast Cancer (2024)
-
The double agents in liquid biopsy: promoter and informant biomarkers of early metastases in breast cancer
Molecular Cancer (2022)
-
Circulating tumor cells: biology and clinical significance
Signal Transduction and Targeted Therapy (2021)