Abstract
Metastasis is the major cause of death in breast cancer patients. Although previous large-scale analyses have identified frequently altered genes specific to metastatic breast cancer (MBC) compared with those in primary breast cancer (PBC), metastatic site-specific altered genes in MBC remain largely uncharacterized. Moreover, large-scale analyses are required owing to the low expected frequency of such alterations, likely caused by tumor heterogeneity and late dissemination of breast cancer. To clarify MBC-specific genetic alterations, we integrated publicly available clinical and mutation data of 261 genes, including MBC drivers, from 4268 MBC and 5217 PBC patients from eight different cohorts. We performed meta-analyses and logistic regression analyses to identify MBC-enriched genetic alterations relative to those in PBC across 15 different metastatic site sets. We identified 11 genes that were more frequently altered in MBC samples from pan-metastatic sites, including four genes (SMARCA4, TSC2, ATRX, and AURKA) which were not identified previously. ARID2 mutations were enriched in treatment-naïve de novo and post-treatment MBC samples, compared with that in treatment-naïve PBC samples. In metastatic site-specific analyses, associations of ESR1 with liver metastasis and RICTOR with bone metastasis were significant, regardless of intrinsic subtypes. Among the 15 metastatic site sets, ESR1 mutations were enriched in the liver and depleted in the lymph nodes, whereas TP53 mutations showed an opposite trend. Seven potential MBC driver mutations showed similar preferential enrichment in specific metastatic sites. This large-scale study identified new MBC genetic alterations according to various metastatic sites and highlights their potential role in breast cancer organotropism.
Similar content being viewed by others
Introduction
Breast cancer is the most common type of cancer in women and the second-most common cause of death due to its metastasis or progression to advanced disease1. Metastasis is a complex process consisting of cancer cell dissemination, intravasation, circulation in the bloodstream, extravasation, and colonization, with various cell types involved in the tumor and affected organ microenvironment. Numerous studies have identified driver mutations in primary cancer, revealing that the accumulation of driver mutations in cancer cells may contribute to cancer cell proliferation and survival2. Several large-scale genomic studies have suggested candidate driver genes of metastatic breast cancer (MBC) using similar methods implemented in studies of primary breast cancer (PBC)3. As a result, 31 candidate driver genes of MBC were identified through analyses of various datasets, including those based on whole-genome sequencing, whole-exome sequencing, or targeted gene sequencing with appropriate statistical methods4,5,6. Most of these studies attempted to identify the driver genes enriched in MBC by comparing the frequency of driver genes between unpaired MBC and PBC samples. Among the 31 candidate driver genes of MBC, several (e.g., ESR1, TP53, and NF1) were revealed to be enriched in MBC compared with those in PBC. Moreover, Razavi et al.7 identified more than 20 additional MBC-enriched altered genes that were not identified in other studies by comparing the frequency of 468 cancer-related genes between MBC and PBC unpaired samples. The identified MBC-enriched altered genes may have functional impacts on MBC clones in the metastatic sites of breast cancer, which should be elucidated in detail.
Despite efforts to identify the driver genes of MBC and MBC-enriched altered genes compared with those in PBC, further large-scale genomic studies of MBC are needed for several reasons. First, power analyses showed that ~1000 PBC samples were needed to achieve 90% statistical power to identify PBC drivers of low frequency (0.02) at one background mutation per mega base pairs, suggesting that more MBC samples would be required to uncover MBC driver genes with similar power2,8. Second, the metastasis of breast cancer may follow a late dissemination model, in which the tumor cells disseminate from a primary tumor site in the late phase of tumorigenesis; therefore, most genetic alterations are likely to be shared between primary and metastatic tumors7,9,10. This similarity may make it difficult to identify driver genes that are specifically altered in MBC but rarely detected in PBC, unless matched MBC and PBC samples are compared. Third, intra-genetic heterogeneity of metastatic tumors results in a low allele frequency of candidate driver mutations among the analyzed samples. Compared with gene-level analysis for identifying driver genes, which uses the collective frequencies of multiple variants in each gene, identifying driver mutations by single variant analysis is more challenging because of the expected rarity of each variant and the requirement of a sufficient sample size to ensure appropriate statistical power. Although several driver mutations enriched in MBC were identified by comparing the genomes of patients with MBC and PBC, the number of identified mutations remains low, partly because of the relatively limited scale of previous studies10.
Moreover, few studies have investigated the genetic alterations in MBC samples according to their metastatic sites. MBC cells were shown to preferentially metastasize to specific organ sites, in a process referred to as organotropism, including the bone (occurrence rate of 47–60%), liver (19–20%), lung (16–34%), and brain (10–16%), suggesting that different altered genes may contribute to the survival of cancer cells at various metastatic sites11,12,13. Although some molecular characteristics of MBC cells in such metastatic sites have been elucidated, large-scale genomic analyses are required to uncover significantly altered genes in specific metastatic sites.
Therefore, to comprehensively characterize genetic alterations in MBC across various metastatic sites, in this study, we integrated sequencing data of 4268 MBC and 5217 PBC samples from eight different cohorts [Dana-Farber Cancer Institute (DFCI), MD Anderson Cancer Center (MDA), Memorial Sloan Kettering Cancer Center (MSK), Vanderbilt-Ingram Cancer Center (VICC), Foundation Medicine Adult Cancer Clinical Dataset (FMAD), Wellcome Trust Sanger Institute (WTSI), INSERM, and The Cancer Genome Atlas (TCGA)], and analyzed 261 cancer-related genes, including most of the previously identified MBC driver genes, to identify genetic alterations in MBC according to their metastatic sites. We also investigated the possible driver mutations of MBC across different metastatic sites.
Results
Pan-metastasis analysis
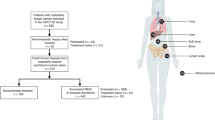
To investigate MBC-specific genetic alterations related to organotropism, we performed three analyses using different datasets and methods (Fig. 1). After performing quality control (QC) and integrating data (Supplementary Tables 1–7 and Supplementary Figs. 1–9), the final number of MBC and PBC samples analyzed was 4268 and 5217, respectively, across seven datasets from the eight cohorts (Table 1). Most of the samples were unpaired between metastasis and primary tumors, except for 17 paired samples from the WTSI cohort. To identify the genes enriched in MBC, we first selected the 261 genes that were concurrently targeted by at least 7 of 11 targeted sequencing panels used in our cohorts (Supplementary Tables 3 and 6). These genes included most of the 31 candidate MBC driver genes identified in three previous studies (Fig. 2a, dark green)10,14,15. In particular, the selected genes included 19 of 20 genes identified by at least two studies, suggesting that these 261 genes were adequate for further analyses to identify MBC-specific genetic alterations. Most of the mutated genes showed a long-tail distribution with a low frequency; over 98% of the 261 genes showed a cumulative mutational frequency of <10% for both MBC and PBC samples (Fig. 2b). Ten of the most frequently altered genes in MBC samples exhibited no difference in the frequency of genetic alterations, compared to that in PBC samples, except for ESR1, ARID1A, and NF1 which were more frequently altered in MBC samples (Supplementary Fig. 10). This indicates a heterogeneous driver landscape and the importance of aggregated data to identify frequently mutated genes in MBC compared with those in PBC9,10.
This study aims to investigate the six research questions using three analyses including pan-metastasis analysis, metastatic site-specific analysis, and driver mutation analysis. For each question, different datasets and data formats were used. Illustrations are created with Biorender.com and maftools47.
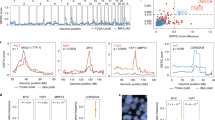
a Candidate MBC driver genes and MBC-enriched altered genes compared with those in PBC are shown. Thirty-one genes identified as MBC driver genes in three previous studies are shown in dark green. These studies used MutSigCV or dNdScv for different datasets to identify MBC driver genes. To identify MBC-enriched altered genes, these studies compared the mutational frequency of the detected driver genes between unpaired MBC and PBC, whereas we and Razavi et al.7 compared cancer-related targeted genes between unpaired MBC and PBC. The targeted genes in the study by Razavi et al.7 and our study are shown in light blue, and the rest of the targeted genes are listed in Supplementary Table 6 and are described in Razavi et al.7, respectively. Asterisks indicate that the tested genes were significantly enriched in MBC for each study (significantly enriched genes in PBC are colored in orange and denoted by an asterisk). To compare the mutational frequency per gene between MBC and PBC, we used meta-analysis and logistic regression analysis, whereas the other studies used Fisher’s exact test. b Top 30 frequently altered genes are presented. The upper panel shows MBC samples, and the lower panel shows PBC samples. c Forest plots present four significantly altered genes (SMARCA4, TSC2, ATRX, and AURKA) that were identified by our meta-analyses and logistic regression analyses and were not shown as MBC-enriched genes in previous studies7,10,14,15. Results of fixed-effects meta-analyses and heterogeneity tests between cohorts are shown. P values were adjusted by the false discovery rate (FDR). d Frequency of samples with mutations of the identified 11 genes according to treatment history and subtypes. Statistically significant difference of the mutational frequency in PBC with de novo MBC or post-treatment MBC are indicated by an asterisk (FDR < 0.05 from logistic regression analysis).
Next, to identify frequently altered genes in MBC, we conducted meta-analyses to compare the frequency of specific mutation types in each gene between MBC and PBC samples and validated the results by multiple logistic regression analysis adjusted for the cohort. Single-nucleotide variants (SNVs)/insertion and deletions (Indels) in 11 genes were frequently identified in MBC (ESR1, SMARCA4, NF1, FGFR4, ARID2, PARP1, TSC2, ATRX, ARID1A, AURKA, and STAG2 at false discovery rate [FDR] < 5%) compared with those in PBC, and ESR1 amplifications were frequently identified in MBC samples (Fig. 2a–b and Supplementary Table 8). Among these significantly altered genes, mutation in estrogen receptor 1 (ESR1), a known cause of primary hormone therapy resistance, showed the highest odds ratio (OR) (7.39; 95% confidence interval [CI] = 5.90–9.26; FDR = 1.64 × 10−85; heterogeneity FDR = 1), consistent with the findings of previous studies (Supplementary Fig. 11)7,10,14,15,16,17,18,19. We identified four MBC-enriched genes, SMARCA4, TSC2, ATRX, and AURKA which were not identified in previous large-scale studies for MBC patients (Fig. 2c)7,10,14,15.
We compared the mutational frequency of the 11 genes between treatment-naïve PBC, treatment-naïve de novo MBC, and post-treatment MBC samples to examine whether the enriched mutations in MBC were acquired resistant or metastasis-specific using available data from treatment records of patients. Of the 11 genes enriched in MBC, SNV/Indels of ARID2 were significantly more frequent in treatment-naïve de novo MBC and post-treatment MBC samples than in treatment-naïve PBC samples in all the subtypes and in the IDC subtype (Fig. 2d). Alterations in ESR1 and NF1 were significantly frequent only in post-treatment MBC samples, indicating that these might reflect treatment selection. ESR1 amplification was significantly more frequent in HR + /HER2− and IDC subtype of post-treatment MBC samples, compared to that in treatment-naïve PBC samples (Supplementary Fig. 12).
Metastatic site-specific analysis
Considering breast cancer organotropism, there may be distinct genetic alterations in MBC cells at each metastatic site20,21. Therefore, we hypothesized that the distinct MBC-enriched altered genes may be associated with particular metastatic sites. To test this hypothesis, we classified the MBC samples into 15 metastatic site sets, including broader categories of pan-metastasis, distant metastasis, and local relapse, and compared the number of samples with altered genes in MBC at specific metastatic site sets with that in PBC using meta-analysis and logistic regression analysis (Supplementary Table 7 and Fig. 3a). These analyses identified 19 genes across 14 metastatic site sets as significantly altered genes in specific metastatic site sets at FDR < 5% (Supplementary Table 8). To investigate the preference of altered genes for specific metastatic sites over other sites, we additionally compared the mutational frequency of 19 genes between MBC samples from metastatic sites using logistic regression analysis (Supplementary Table 9). Of note, four altered genes (ESR1, CDH1, RICTOR, and TP53) showed a preference for at least one specific metastatic site in all subtypes of MBC, indicating that these genes tend to metastasize to specific sites rather than other sites as well as they were enriched in MBC compared with PBC (FDR < 5%; Fig. 3b–c and Supplementary Fig. 13). Intriguingly, ESR1 and TP53 mutations showed a significant preference for more than one metastatic site in the opposite direction, particularly in the liver and lymph nodes (Table 2). In the liver, ESR1 mutations were more frequently observed than other sites while TP53 mutations were less frequently observed than other sites (OR = 3.41 vs. 0.63; 95% CI = 2.67–4.35 vs. 0.52–0.76; FDR = 1.19 × 10−21 vs. 4.18 × 10−5, respectively). The tendency was opposite in the lymph nodes (OR = 0.31 vs. 1.46; 95% CI = 0.19–0.46 vs. 1.17–1.83; FDR = 2.29 × 10−6 vs. 1.67 × 10−2 ). Although it should be elucidated in future studies at scale, the observation was consistent with a recent study that showed TP53 and ESR1 were the most mutually exclusive gene pair in HR + /HER2− MBC (exclusivity score \(\phi =\) 5.4 × 10−9)14. In the analysis of cases with MBC samples from multiple metastatic sites that were biopsied sequentially, we also found that ESR1 mutations were more frequently observed in the liver than bone or ovary (Supplementary Table 10 and Fig. 3d).
a Number of samples across 15 metastatic site sets (pan, local, distant, and 12 specific distant sites). In 12 specific metastatic site sets, 16 samples from the MDA cohort were excluded owing to their vague description of metastatic site (e.g., “distant organ metastasis”). The number of cohorts used in each metastatic site set is presented in parentheses. b Colored genes indicate those are more frequently altered in the corresponding metastatic site than other sites as well as MBC-enriched compared with PBC (FDR < 5%). Asterisk indicates that mutations in ESR1 are significantly less frequently altered in the lymph node than other sites while significantly enriched in MBC compared with PBC (FDR < 5%). c Distribution of mutational frequency of the four genes (SNVs/Indels) significantly altered in all MBC subtype samples across tissues. Metastatic sites with frequently altered genes compared with both other sites and PBC (FDR < 5%) are shown in red, significant genes compared with other sites in green, significant genes compared with PBC in orange, insignificant genes in white, and genes mutated in only one sample in gray. Asterisks indicate an odds ratio <1. d Preference of ESR1 mutations to the liver is represented by comparing mutations between metastatic liver samples and metastatic bone or metastatic ovarian samples from the same patients. Sampling (months) represents months between sample biopsy for sequencing and diagnosis of PBC. e Preference of RICTOR mutations to the bone is presented. Comparison of bone metastasis with PBC or other metastatic (M) sites are shown in the forest plot. P values were adjusted by FDR. Significantly frequent alterations of RICTOR are observed in de novo HR+/HER2− MBC and de novo IDC, as shown in the bottom bar graphs. Asterisks indicate statistical significance at P < 0.05.
To test the effect of intrinsic subtypes of the samples on the associations between the four identified metastatic sites-enriched genes (ESR1, CDH1, RICTOR, and TP53) and their metastatic sites (liver, lymph nodes, bone, chest wall/thorax, brain, ovary, and peritoneum), we performed a logistic regression analysis with a covariate of histology or receptor status. The associations of ESR1 mutations with liver metastasis and RICTOR mutations with bone metastasis were significant, irrespective of the intrinsic subtypes. The preference of altered CDH1 to ovarian and peritoneum metastasis was likely to be related to the preference of ILC subtype for ovary and peritoneum since we observed statistically significant association between ILC subtype and CDH1 mutations for both ovarian and peritoneum metastases (P < 0.01, two-sided Fisher’s exact test), consistent with a previous study22. We also found that RICTOR mutations were prevalent in the bone and were also frequently observed in de novo treatment-naïve MBC samples, particularly in HR+/HER2− and IDC subtypes (Fig. 3e). These results supported MBC organotropism in which distinct mutations may be involved in metastasis to specific metastatic sites.
Driver mutation analysis
For the 19 significantly frequently altered genes (FDR < 5%) in MBC (Supplementary Table 8), we assessed the potential functional impact of each variant using computational algorithms and a knowledge-based approach (pattern-based analysis) to identify candidate driver mutations in these genes (Supplementary Fig. 14 and Supplementary Tables 11–12). Twenty-seven mutations across 14 metastatic site sets were suggested as potential driver mutations (Func score \(\ge\)5), most of which were also suggested as driver mutations with high driver likelihood (>0.5) in a recent study (91%; 20 out of 22 variants analyzed in the study)23. Furthermore, seven of the 27 variants were significantly frequently observed in specific metastatic sites compared with other sites and were enriched in MBC compared with those in PBC (FDR < 5%) (Fig. 4a and Supplementary Tables 13–14). Intriguingly, we identified that ESR1 p.D538G was significantly frequently observed in the liver while less frequent in the lymph nodes than other metastatic sites consistent with our findings from metastatic site-specific analysis at a gene level (Fig. 3) (OR = 4.24 vs. 0.19; 95% CI = 2.91–6.23 vs. 0.07–0.42; FDR = 1.96 × 10−12 vs. 7.71 × 10−3). We additionally confirmed that our candidate driver mutations of MBC were also detected by other tools for driver analysis (MutSigCV and dNdScv) (Fig. 4b)24,25. In breast metastasis, p.Y537 and p.D538 mutations in ESR1 were related to organotropism among many other hotspot mutations (Fig. 4c). We also found that truncated mutations of CDH1 were frequently observed in peritoneum metastasis consistent with the results of metastatic site-specific analyses (Figs. 3b and 4a).
a Significant MBC driver mutations across metastatic sites are shown by comparing 27 candidate driver mutations from MBC of a specific metastatic site with other metastatic sites or PBC. Presented results are from the logistic regression analysis for mutational frequency of 27 candidate driver mutations between MBC samples from different metastatic sites. Truncating mutations are represented by “Trunc” and missense mutations are shown by their amino acid change. Odds ratios and 95% CI for each mutation were shown and asterisks indicate statistical significance at FDR < 0.01. b MBC driver genes of WTSI and INSERM cohort data which had sufficient background mutations from the results of MutSigCV and dNdScv. c Mutation types of ESR1 shown in a lollipop plot. Red-colored variants are candidate driver mutations enriched in MBC samples. X-axis indicates amino acid position.
Discussion
We comprehensively identified the frequently altered genes and driver mutations in MBC across 15 metastatic sites using 261 cancer-related genes with rigorous statistical analyses of 4268 MBC and 5217 PBC samples from eight cohorts. In addition to replicating the findings from recent large-scale genomic studies for MBC, we identified MBC-specific SNVs/Indels in SMACRA4, TSC2, ATRX, and AURKA which were not identified in previous large-scale studies7,10,14,15. We also found preference of mutations in four genes (ESR1, CDH1, RICTOR, and TP53) to specific metastatic sites. Furthermore, ESR1 and TP53 showed a mutually exclusive tendency of organotropism for the liver and lymph nodes, which should be validated in a large independent dataset (Table 3). Driver mutation analysis also supported our findings of organotropism, particularly for ESR1 mutations. Thus, our study characterized metastatic site-specific genetic alterations at a large scale and suggested that distinct genetic alterations may be involved in the different metastatic sites of MBC, supporting breast cancer organotropism.
The novel genes frequently altered in MBC compared with PBC have previously been reported to be involved in metastasis-related mechanisms. For example, down-regulation of SMARCA4 expression inhibited the proliferation, invasiveness, and motility of breast cancer cells in vitro and suppressed metastasis in breast cancer mouse models, suggesting a role of these genes in metastasis4,5. In addition, a high expression level of SMARCA4 or AURKA was associated with poor survival or metastasis-free survival in breast cancer patients26,27. Furthermore, in pancreatic neuroendocrine tumors, aberrant ATRX and DAXX expression was associated with lymph node metastasis and distant metastasis by causing the abnormal lengthening of telomeres, and was also associated with shorter disease-free survival and disease-specific survival28,29. These results suggest that genes identified in this study may be involved in the general metastatic process of breast cancer.
Stephen Paget studied 735 autopsies of female breast cancer patients and proposed the ‘seed-and-soil’ hypothesis that metastasis does not occur by chance but metastatic tumor cells have a preference for specific organs30. To elucidate what genetic factors affect organotropism, it is required to analyze many autopsy samples for multisite metastases, but collecting such samples at scale is challenging31. Our approach may be one of the most feasible approaches to investigating breast cancer organotropism, which combines ‘cohort-wise comparison’ of mutational frequency between MBC samples from different metastatic sites as well as between MBC and PBC samples at scale and ‘comparison of multi-metastatic MBC samples’ from the same patients in a relatively small sample size. However, it is important to note that our analyses may be affected by the fact that the site of sampling was the most clinically accessible site, and may not reflect that most patients with MBC have multiple metastatic sites.
Among the 19 altered genes identified by the metastatic site-specific analysis, four genes showed high mutational frequency at specific tissues, further supporting the organotropism of MBC. RICTOR was most frequently altered in bone-metastatic breast cancer samples (10/201 = 5%), which is consistent with a previous study showing that bone-derived mesenchymal stem cells (MSCs) with aberrant RICTOR expression inhibited breast cancer bone metastasis by repressing osteolytic destruction and cancer-associated fibroblasts32. Considering the plasticity of cells between naïve MSCs and breast cancer cells, the higher mutational frequency of RICTOR may be related to the bone tropism of MBC cells33. Although TP53 was also frequently mutated in various tissues of our MBC samples, this alteration was most frequently identified in the brain (70/86 = 81%). A previous study also observed frequently mutated TP53 in 23 central nervous system-metastatic breast cancer patients, supporting that TP53-mutant cells may preferentially metastasize to the brain34.
There are several limitations to our study. First, we focused on a limited number of genes (261 genes) that were analyzed largely by targeted sequencing. Deep, whole-exome, or whole-genome sequencing in large cohorts is required for unbiased screening of novel genes involved in metastasis and the related mechanisms. Second, we were not able to investigate tumor evolution, but rather could only provide insights into tumor epidemiology, as most of our analyzed samples did not involve primary-metastatic matched pairs like other recent studies7,14,15. Therefore, the recurrently altered genes identified by our analysis of largely unmatched metastatic samples may include somatic mutations that might also exist in primary tumors. Direct comparisons of mutational patterns in primary-metastatic matched pairs at scale are necessary to identify and confirm mutations that are specific to MBC. Third, despite the large sample size, the source of the analyzed datasets was heterogeneous in terms of sequencing technologies, sequencing depths, and mutation callers used. To account for this heterogeneity, we analyzed mutations observed only in concurrently targeted sequencing regions in all cohorts and utilized both meta-analysis and regression analysis adjusted for the cohort. Furthermore, treatment history was available only for a small portion of the samples (610 treatment-naïve PBC samples, 86 treatment-naïve de novo MBC samples, and 692 post-treatment MBC samples). Our findings need to be replicated in further studies with a large number of treatment-naïve samples to determine whether the identified mutations were metastasis-specific or acquired resistant. Finally, our suggested driver mutations should be validated experimentally although they were confirmed using three different approaches (pattern-based analysis, MutSigCV, and dNdScv).
In conclusion, we identified distinct genetic alterations of MBC according to specific metastatic sites using large-scale analyses. The molecular characteristics of MBC cells at specific metastatic sites discovered in our study may be considered as biomarkers or therapeutic targets of MBC patients with specific metastases.
Methods
Cohort description
We used SNVs, Indels, copy number alterations (CNAs), and clinical data from eight different cohorts (Table 1). Details for each cohort are described below. SNVs/Indels and CNAs data were extracted from the selected samples as described below (Supplementary Figs. 1–2 and Supplementary Tables 1 and 2). Because the MSK, WTSI, INSERM, and TCGA cohorts used matched normal-tumor pairs, whereas the others used a stringent process for filtering germline variants, all mutations were regarded as somatic mutations (Supplementary Table 3).
Data from three cohorts [Dana-Farber Cancer Institute (DFCI), MD Anderson Cancer Center (MDA), and Vanderbilt-Ingram Cancer Center (VICC)] were obtained from AACR Genomics Evidence Neoplasia Information Exchange (GENIE) version 8.0 (https://www.synapse.org/#!Synapse:syn7222066/wiki/405659)35. Foundation Medicine Adult Cancer Clinical Dataset (FMAD) data were obtained from dbGAP (phs001179.v1.p1), whereas other data [Wellcome Trust Sanger Institute (WTSI) and INSERM] were obtained from the supplementary data of published studies10,17,36,37,38. Data of the Memorial Sloan Kettering Cancer Center (MSK) cohort were obtained from both a published study by Razavi et al.7 and GENIE version 8.0. We used female patient samples, and all data from each cohort contained a set of metastatic samples and mostly comprised unmatched primary samples, except for WTSI cohort samples which consisted of only paired samples. We excluded patients who had multiple samples from multiple metastatic sites to avoid redundant results. The Cancer Genome Atlas (TCGA)-BRCA dataset was used as the primary cancer data for the INSERM cohort. Although the original analysis of the INSERM study used a previous version of the TCGA-BRCA data, we downloaded a mutation annotation format (MAF) file of the latest version (version: gdc-1.0.0, file date: 20170930, number of analyzed samples = 986) of TCGA-BRCA from the Genomic Data Commons data portal (GDC; https://portal.gdc.cancer.gov/) called by MuTect and input to GRCh37 using maf2vcf.pl and vcf2maf.pl (https://github.com/mskcc/vcf2maf) in the GENIE process internally. The details of each cohort dataset are described below.
All data for DFCI, MDA, and VICC cohorts were obtained from GENIE version 8.035. In “data_clinical_sample.txt,” samples were selected when CANCER_TYPE == “Breast Cancer” and SAMPLE_TYPE== “Metastasis” for metastatic breast cancer (MBC) samples and SAMPLE_TYPE == “Primary” for primary breast cancer (PBC) samples. Using sex information from “data_clinical_patient.txt,” male samples were excluded. Samples analyzed by targeted sequencing with panels for all exonic regions of targeted genes, and not hotspot regions, were included (DFCI-ONCOPANEL-1/2/3, MDA-409-V1, VICC-01-T5A, and VICC-01-T7). Histological information was obtained from ONCOTREE_CODE in “data_clinical_sample.txt,” which described invasive ductal carcinoma as “IDC” and invasive lobular carcinoma as “ILC.” Other types of histology were regarded as “Others.” Information regarding metastatic sites was obtained from SAMPLE_TYPE_DETAILED in “data_clinical_sample.txt” when available. Age information was also obtained from “data_clinical_sample.txt” at the sequencing report. The SNVs and insertions/deletions (Indels) data were obtained from selected samples in “data_mutations_extended.txt.” For DFCI and VICC, available copy number alterations (CNAs) data were selected from the selected samples in “data_CNA.txt.” The CNAs values include: low-level gain (1); high-level amplification (2), which was regarded as amplification; as well as deep loss (−2) and single-copy loss (−1), which were regarded as deletion. We selected high-level amplification (2) and deep loss (−2) deletion for harmonization with data from other cohorts and for minimizing false-positives.
Data for the MSK cohort were obtained from both a published study by Razavi et al.7 (downloaded from cBio Portal at https://cbioportal.org on Dec 19, 2017) and GENIE version 8.0. In these datasets, 670 MBC and 722 PBC samples overlapped, 111 MBC and 100 PBC samples were only available in the published study (Razavi et al.7), and 1190 MBC and 1431 PBC samples were only available in GENIE version 8.0. We combined all available samples from both sources in pan-metastatic analysis, whereas samples from Razavi et al.7 were used in other analyses as well, including metastatic site-specific analyses and survival analysis, owing to the availability of the necessary clinical information such as metastatic sites, receptor status, and survival time and status. In the analysis of mutational frequencies of MBC-enriched genes, we also used information of de novo MBC status in data of Razavi et al.7 Selected MBC and PBC samples were both annotated as PRIMARY_SITE == “Breast” and annotated as SAMPLE_TYPE == “Metastasis” and SAMPLE_TYPE == “Primary,” respectively. Because the target panels of the MSK cohort were all exonic regions of the targeted genes, all samples analyzed by any target panels of MSK were selected (MSK-IMPACT341/410/468). Histological information was obtained from TUMOR_SAMPLE_HISTOLOGY in “data_clinical_sample.txt.” Samples annotated as “Breast Invasive Ductal Carcinoma” were regarded as “IDC,” “Breast Invasive Lobular Carcinoma” as “ILC,” and others as “Others.” Metastatic site and age-related information were obtained from SAMPLE_SITE and INVASIVE_CARCINOMA_DX_AGE in “data_clinical_sample.txt,” respectively. Information on receptor status, overall survival months, and overall survival status were obtained from OVERALL_RECEPTOR_STATUS_PATIENT, OS_MONTHS, and OS_STATUS in “data_clincal_patient.txt,” respectively. The SNV/Indels data were extracted from selected samples in “data_mutations_extended.txt,” and CNAs data were selected from the selected samples in “data_CNA.txt.” There were three levels of CNA values: high-level amplification (2), which is regarded as amplification, as well as deep loss (−2) and single-copy loss (−1.5), which are regarded as deletions. We selected the CNA values described as high-level amplification (2) and deep loss (−2) for harmonization of these data with other cohorts and for minimizing false-positives. In the mutation data of Razavi et al.7, there were different names for the same genes, including KMT2A, KMT2D, MLL, and MLL2. We converted the old name of the gene symbols “MLL” and “MLL2” to “KMT2A” and “KMT2D,” respectively. Moreover, we excluded duplicate variants shown in 9 MBC samples and 15 PBC samples, and we excluded variants in CDKN2B-AS1 annotated by Variant Effect Predictor (VEP), CDKN2A p14ARF.
FMAD data were accessed from dbGAP (phs001179.v1.p1), which includes mutation data (SNVs/Indels) in a MAF file and clinical data36. As the MAF file was aligned to the GRCh38 reference genome, we converted the genome coordinates to GRCh37 using the liftOver tool (https://genome.ucsc.edu/cgi-bin/hgLiftOver) for further analyses. MBC and PBC samples were selected and annotated as classification_of_tumor == “metastasis” and “primary,” respectively. Samples of FMAD data were analyzed using a cancer gene panel for all exonic regions. Information on histology, age at diagnosis, and metastatic sites were used as described in the clinical data. The SNVs/Indels data were extracted from the selected samples.
Data from the WTSI cohort were obtained from the supplementary materials of a published paper for 386 MBC samples of a relapsed cohort analyzed by targeted sequencing and/or whole-genome sequencing10. We used 208 samples with matched normal tissues analyzed by targeted sequencing or both targeted sequencing and whole-genome sequencing for SNVs/Indels analysis, as well as 45 samples analyzed by whole-genome sequencing or both targeted sequencing and whole-genome sequencing for CNA analysis. After excluding samples biopsied from more than one site, we finally selected 53 MBC female samples and 31 PBC female samples that were annotated as SAMPLE_CODE == “DISTANT_METASTASIS,” “LOCAL_RELAPSE” or “SYNC_LN,” and SAMPLE_CODE == “PRIMARY,” respectively. There were 17 metastasis primary paired samples. Information on histology, receptor status, and age was obtained from CODED_HISTO, ER_HER2_PRIMARY, and Age_at_diagnosis, respectively. To determine the receptor status, we evaluated the estrogen receptor as a hormone receptor (HR). Information on metastatic sites was obtained from SAMPLE_CODE. The SNVs/Indels data were obtained from the selected samples as described above, and CNAs data were obtained from the selected samples as described above. Before converting the mutation file to a MAF format file, we corrected Indel variants for harmonization into the MAF format by adjusting the chromosomal start position, end position, reference alleles, and alternative alleles. CNA data were annotated as effect == “AMP” or “HOM_DEL,” and we selected all CNA data because the CNA values included a copy number of more than five as amplification and zero copy number as homo deletion.
Data for the INSERM cohort were acquired from supplementary materials of a published retrospective study of metastatic breast cancer17. In the study, as TCGA-BRCA data were used as the primary breast cancer data for the INSERM cohort, we downloaded and analyzed the latest version of TCGA-BRCA for primary cancer of the INSERM cohort as described below. Raw samples were analyzed by whole-exome sequencing of a single metastatic site, and 196 samples that had mutations in 261 genes remained. Information regarding sex was not available. Information regarding the HR status and metastatic sites was used as described in Table 1, and samples annotated as HER2+ were classified into HR+/HER2+ and HR−/HER2−, as our classification considered more specific subtypes. The SNV/Indel data for the selected samples were used. Before converting the mutation file to a MAF format file, we corrected Indel variants for harmonization to the MAF format by adjusting the chromosomal start position, end position, reference alleles, and alternative alleles. As CNAs data were not available for each sample, this cohort was not included in our CNAs analysis.
TCGA-BRCA data were used as the primary cancer for the INSERM cohort. We used the latest version of the TCGA-BRCA data downloaded from the GDC in a MAF file called by MuTect (version: gdc-1.0.0, file date: 2017-09-30, number of analyzed samples = 986).37,38 As the file was aligned to reference genome version GRCh38, we converted the genome coordinates to GRCh37 using the liftOver tool (https://genome.ucsc.edu/cgi-bin/hgLiftOver). Clinical data for samples in the MAF file were used in our study, and only samples from female patients were included. Age and histological information were obtained from age_at_diagnosis and primary_diagnosis described in the clinical data, respectively. The SNV/Indel data for all samples were used. As CNAs data for INSERM were not available, we did not use the CNAs data of TCGA.
Sample selection
For most of the analyses, except for metastatic site-specific analysis with multi-metastatic samples, we used female patient samples (except for the INSERM cohort which did not provide information regarding sex) biopsied from just one site to avoid redundant counts of genetic alterations in multiple samples from the same patients. We selected samples analyzed by whole-exome sequencing or targeted sequencing of all exonic regions for each gene after excluding samples with only hotspot regions sequenced (Supplementary Tables 1–3). Most of the selected samples were unpaired between primary and metastatic cancer, except 17 paired samples from the WTSI cohort. After variant QC, samples with no variants or too many variants were excluded as described in the Supplementary Information (Supplementary Tables 4–5 and Supplementary Figs. 3–9). The final sample numbers were 4268 MBC and 5217 PBC samples for SNVs/Indels data, and 1805 MBC and 1783 PBC samples for CNAs data. If relevant information was available, the samples were classified into subtypes according to their receptor status, histology, or specific metastatic site. Samples from local recurrence sites were composed of those from ipsilateral and contralateral breast relapse, chest wall, or regional lymph nodes, and other samples were regarded as those of distant metastases. For metastatic site-specific analysis with multi-metastatic samples, we used 48 patient samples from at least two metastatic sites (Supplementary Table 10 and Fig. 3d). For the analysis of mutational frequency based on treatment records, we used data of 610 treatment-naïve PBC samples, 86 treatment-naïve de novo MBC samples and 692 post-treatment MBC samples from Razavi et al. which provided information for de novo status of MBC samples and treatment records for the samples7.
Multi-region samples
To validate organotropism of specific genes, we examined 48 multi-metastatic site samples (Supplementary Table 10) which were excluded in other analyses. After processing and QC with the same pipeline as analyses without multi-metastatic site samples, we checked whether four identified MBC-enriched genes for specific metastatic sites (ESR1, CDH1, RICTOR, and TP53) were observed in multi-region samples.
Data integration
Clinical data and SNVs/Indels data were available for all cohorts, whereas CNAs data were available for only four cohorts DFCI, MSK, VICC, and WTSI. Although eight different cohorts were used in our study, downstream analyses were performed on seven datasets of MBC and PBC because the INSERM and TCGA datasets were compared as metastatic samples and primary samples, respectively (Table 1). Because the analytical pipelines of each cohort varied, we harmonized the data formats and obtained a unified annotation of genes and variants. For FMAD and TCGA cohort data, we matched genome coordinates to GRCh37 from GRCh38 using the liftOver tool (https://genome.ucsc.edu/cgi-bin/hgLiftOver). Other data were aligned to a reference of the GRCh37 version in the original files. Then, gene symbols of SNVs/Indels and CNAs data were transformed into the HGNC symbols when gene symbols in the data from each cohort differed from the HGNC symbols according to the latest version (19,198 protein-coding genes; ftp://ftp.ebi.ac.uk/pub/databases/genenames/new/tsv/locus_groups/protein-coding_gene.txt). For SNVs/Indels data, to generate a uniform mutation file annotated by the same version of an annotation tool, we converted mutation files from each cohort into MAF files using publicly available scripts (vcf2maf.pl and maf2vcf.pl) in GENIE data integration (https://github.com/mskcc/vcf2maf) and bcftools (https://samtools.github.io/bcftools). Each variant in the MAF file was annotated by VEP version 9035. After converting of gene symbols to HGNC symbols as described above, we unified the CNAs format that contained the same gene symbols and the status of amplification or deletion.
Quality control of data for SNVs/Indels data
After integrating the datasets, we preprocessed the data by following QC steps for SNVs/Indels data (Supplementary Fig. 1 and Supplementary Table 4). After VEP annotation, we excluded variants with no reference allele, alternative allele, or duplicated variants in the same sample. We discarded the variants for which gene symbols were different before and after VEP annotation.
We also excluded variants with low coverage (<10 depth) or low variant allele frequency (<0.01 VAF) (Supplementary Figs. 3–5). To analyze functionally effective mutations, mutations annotated as “5′ Flank,” “3′ Flank,” “5′ UTR,” “3′ UTR,” “Intron,” or “Silent” were excluded; the final mutation data of SNVs/Indels were composed of variants classified as “Frame_Shift_Del,” “Frame_Shift_Ins,” “In_Frame_Del”, “In_Frame_Ins,” “Missense_Mutation,” “Nonsense_Mutation,” “Nonstop_Mutation,” “Splice_Region,” “Splice_Site,” or “Translation_Start_Site.” Before exclusion of variant classes, silent mutations were not found in most cohorts (Supplementary Fig. 6).
Finally, we excluded variants observed in outlier samples with a large number of mutations (Supplementary Fig. 7). The final numbers of SNVs and Indels were 21,369 and 20,026 for 4268 MBC samples and 5217 PBC samples, respectively.
QC for CNAs data
After HGNC conversion, we filtered out variants that showed a value of –1 or 1 in the DFCI and VICC cohorts, or a value of –1.5 in the MSK cohort (low CNAs filter) to strictly analyze CNAs generated from targeted sequencing data (Supplementary Figs. 8 and 9). The final numbers of CNAs were 7416 and 5781 for 1805 MBC and 1783 PBC samples, respectively.
Focusing on 261 genes
As most of the cohorts targeted different sets of genes by various targeted sequencing methods, we limited our analyses to concurrently targeted regions to avoid potential bias. For our analyses, we selected a total of 261 genes that were retained in at least 7 of the 11 (60%) targeted sequencing panels used in six cohorts. To investigate whether the 261 genes included important known and novel genes in MBC and PBC, we listed candidate MBC driver genes as well as candidate PBC driver genes identified by previous large-scale analyses, and compared the previously identified genes with the 261 genes (Supplementary Table 6)10,14,15,39. To identify driver genes, Bertucci et al. used MutSigCV for MBC samples analyzed by whole-exome sequencing, whereas Angus et al. and Yates et al. used dNdScv for MBC samples analyzed by whole-genome sequencing and targeted sequencing, respectively10,14,15. Nik-Zainal et al.39 also used dNdScv for PBC samples analyzed by WGS. We regarded the 261 genes as a representative gene set to investigate MBC driver genes, as they included most of the MBC candidate driver genes and PBC candidate driver genes.
Statistical analysis
To identify frequently altered genes in MBC, inverse variance-weighted meta-analyses were conducted based on the numbers of mutation carriers in MBC and PBC. We performed a separate meta-analysis for three mutation types: SNVs/Indels, amplifications, and deletions. Joint ORs and 95% CIs were calculated by assuming a fixed-effects model based on the Cochran-Mantel-Haenszel method when the heterogeneity between cohorts was low (q-value ≥ 0.05), tested by Tarone’s Q test using I2 statistics adjusted by the FDR. When the heterogeneity between cohorts was significant (q-value < 0.05), joint ORs and 95% CIs were calculated by assuming a random-effects model based on the restricted maximum-likelihood (REML) method. Joint P values were adjusted by the FDR, and meta-analyses were conducted using the “metafor” and “meta” packages in R (version 3.5.1).
To validate the results of the meta-analysis for pan-metastases, metastatic site-specific analysis and driver mutation analysis for comparing MBC with PBC, and multivariable logistic regression analyses were conducted with metastasis status as a dependent variable and mutation carrier status as an independent variable after adjustment for cohorts. For the analysis of comparing mutational frequency between MBC samples from different metastatic sites, we compared the mutational frequency of MBC samples from a specific metastatic site with other metastatic sites for 19 MBC-enriched genes (FDR < 5%) using only MSK(R) and FMAD cohort samples (N = 2,160) by multivariable logistic regression analysis with the metastatic site as a dependent variable after adjustment for cohorts. ORs and 95% CIs were estimated, and P values were adjusted by the FDR. We presented organotropism-related genes when they were statistically significant from both analyses (comparing mutational frequency between MBC samples from different metastatic sites and between MBC and PBC samples).
Because our data were composed of eight different cohorts, both strategies (heterogeneity test of meta-analyses and multivariable logistic regression analyses with a covariate of cohorts) were used to determine that our results were unlikely to be cohort-specific.
To investigate the effect of treatment status on 11 metastasis-enriched genetic alterations, we conducted multivariable logistic regression analysis between treatment-naïve PBC samples and treatment-naïve de novo MBC samples or post-treatment MBC samples for all subtypes and specific subtypes according to receptor status or histology.
Analysis of driver mutations in MBC
To investigate whether identified MBC-enriched genes across metastatic sites were driver mutations, we performed two strategies as follows considering data type (targeted sequencing data or WES data). First, we performed pattern-based analysis for all datasets. To investigate the functional effects of the frequently altered genes in MBC at the variant level, we applied the following heuristic strategies to identify candidate driver mutations using the mutational pattern of cancer genes that oncogenes (OGs) tend to be recurrently altered in hotspot regions, whereas tumor suppressor genes (TSGs) tend to be truncated by gaining truncating mutations (nonsense, splice-site, or frameshift insertions/deletions) (Supplementary Fig. 14)40,41. First, the identified MBC-enriched altered genes according to metastatic site sets were classified as OGs, TSGs, or both as annotated in the 723 cancer gene list of Cancer Gene Census (CGC) version 90 of Catalogue of Somatic Mutations in Cancer (COSMIC)42. Genes not annotated in the CGC were discarded. Second, for genes in the CGC, we defined a mutational pattern for each gene. For OGs, we counted the number of MBC samples with mutations in hotspot regions defined by Cancer Hotspots (https://www.cancerhotspots.org)43,44. For TSGs, we counted the number of MBC samples with truncating mutations (nonsense, splice-site, or frameshift Indels). For genes classified as both OGs and TSGs, we considered the mutational patterns of both OG and TSG for the genes by counting samples with hotspot mutations for OGs and those with truncating mutations for TSGs. Third, to consider the functional impacts of the mutations, we scored each mutation by annotation from three functional annotation tools (Mutation Assessor, PolyPhen2, and SIFT) for OGs and by experimental evidence in the literature that supported tumor suppressor function or increased invasiveness/metastatic potential for TSGs. We included mutations with a high functional impact (score \(\ge\)5) observed in more than two carriers in meta-analysis and logistic regression analysis. Finally, for the mutations that satisfied the defined mutational pattern of TSG or OG (N = 27), which were referred to as candidate driver mutations in our study, the numbers of mutation carriers and non-carriers of MBC were compared with those of PBC per gene by Cochran-Mantel-Haenszel inverse variance-weighted meta-analysis. The REML method was used when the heterogeneity between cohorts was significant. Joint P values were adjusted by the false discovery rate for the tested mutation patterns (Supplementary Tables 11–14). Additionally, we compared mutational frequencies of the 27 candidate driver mutations between MBC samples from different metastatic sites by logistic regression analysis adjusted for cohorts to identify MBC-enriched driver mutations preferentially for specific metastatic sites. P values were adjusted by FDR.
In addition to pattern-based analysis, to confirm driver mutations identified by pattern-based analysis, we performed additional driver mutation analyses using MutSigCV and dNdScv for WTSI and INSERM MBC datasets which had sufficient background mutations (synonymous mutations)24,25.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data generated and analyzed during this study are described in the following data record: 10.6084/m9.figshare.1479854145. For DFCI, MDA, MSK(A), and VICC cohorts, the data that support the findings of this study are available in the Synapse repository’s Project GENIE archive, version 8.0 under accession syn7222066 (10.7303/syn22228642)35. For the FMAD cohort, the data are available in the dbGAP repository under accession https://identifiers.org/dbgap:phs001179.v1.p146. For the TCGA cohort, the data are available in the Genomic Data Commons data portal (GDC; https://portal.gdc.cancer.gov/) called by MuTect. For other cohorts (MSK(R), WTSI, and INSERM), the data are available in supplementary materials of published studies7,10,17.
Code availability
Meta-analysis and logistic regression analysis were performed using standard R (v.3.5.1) from the ‘metafor’ package and ‘glm’ function, respectively. For preprocessing, we used public available pipelines (liftOver, maf2vcf.pl, vcf2maf.pl, bcftools).
References
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2018. CA Cancer J. Clin. 68, 7–30 (2018).
Bailey, M. H. et al. Comprehensive characterization of cancer driver genes and mutations. Cell 174, 1034–1035 (2018).
Shah, S. P. et al. The clonal and mutational evolution spectrum of primary triple-negative breast cancers. Nature 486, 395–399 (2012).
Wu, Q. et al. The SWI/SNF ATPases are required for triple negative breast cancer cell proliferation. J. Cell Physiol. 230, 2683–2694 (2015).
Bai, J. et al. BRG1 is a prognostic marker and potential therapeutic target in human breast cancer. PLoS ONE 8, e59772 (2013).
Cao, J. et al. Histone demethylase RBP2 is critical for breast cancer progression and metastasis. Cell Rep. 6, 868–877 (2014).
Razavi, P. et al. The genomic landscape of endocrine-resistant advanced breast cancers. Cancer Cell 34, 427–438 (2018).
Lawrence, M. S. et al. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 505, 495–501 (2014).
Hunter, K. W., Amin, R., Deasy, S., Ha, N. H. & Wakefield, L. Genetic insights into the morass of metastatic heterogeneity. Nat. Rev. Cancer 18, 211–223 (2018).
Yates, L. R. et al. Genomic evolution of breast cancer metastasis and relapse. Cancer Cell 32, 169–184 (2017).
Chen, W., Hoffmann, A. D., Liu, H. & Liu, X. Organotropism: new insights into molecular mechanisms of breast cancer metastasis. NPJ Precis Oncol. 2, 4 (2018).
Smith, H. A. & Kang, Y. Determinants of organotropic metastasis. Annu. Rev. Cancer Biol. 1, 403–423 (2017).
Feng, Y. et al. Breast cancer development and progression: Risk factors, cancer stem cells, signaling pathways, genomics, and molecular pathogenesis. Genes Dis. 5, 77–106 (2018).
Bertucci, F. et al. Genomic characterization of metastatic breast cancers. Nature 569, 560–564 (2019).
Angus, L. et al. The genomic landscape of metastatic breast cancer highlights changes in mutation and signature frequencies. Nat. Genet. 51, 1450–1458 (2019).
Zehir, A. et al. Mutational landscape of metastatic cancer revealed from prospective clinical sequencing of 10,000 patients. Nat. Med. 23, 703–713 (2017).
Lefebvre, C. et al. Mutational profile of metastatic breast cancers: a retrospective analysis. PLoS Med. 13, e1002201 (2016).
Toy, W. et al. ESR1 ligand-binding domain mutations in hormone-resistant breast cancer. Nat. Genet. 45, 1439–1445 (2013).
Robinson, D. R. et al. Activating ESR1 mutations in hormone-resistant metastatic breast cancer. Nat. Genet. 45, 1446–1451 (2013).
Obenauf, A. C. & Massague, J. Surviving at a distance: organ specific metastasis. Trends Cancer 1, 76–91 (2015).
Nguyen, D. X., Bos, P. D. & Massague, J. Metastasis: from dissemination to organ-specific colonization. Nat. Rev. Cancer 9, 274–284 (2009).
Arpino, G., Bardou, V. J., Clark, G. M. & Elledge, R. M. Infiltrating lobular carcinoma of the breast: tumor characteristics and clinical outcome. Breast Cancer Res. 6, R149–R156 (2004).
Priestley, P. et al. Pan-cancer whole-genome analyses of metastatic solid tumours. Nature 575, 210–216 (2019).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Martincorena, I. et al. Universal patterns of selection in cancer and somatic tissues. Cell 171, 1029–1041 (2017).
Siggelkow, W. et al. Expression of aurora kinase A is associated with metastasis-free survival in node-negative breast cancer patients. BMC Cancer 12, 562 (2012).
Guerrero-Martinez, J. A. & Reyes, J. C. High expression of SMARCA4 or SMARCA2 is frequently associated with an opposite prognosis in cancer. Sci. Rep. 8, 2043 (2018).
de Wilde, R. F. et al. Loss of ATRX or DAXX expression and concomitant acquisition of the alternative lengthening of telomeres phenotype are late events in a small subset of MEN-1 syndrome pancreatic neuroendocrine tumors. Mod. Pathol. 25, 1033–1039 (2012).
Singhi, A. D. et al. Alternative lengthening of telomeres and loss of DAXX/ATRX expression predicts metastatic disease and poor survival in patients with pancreatic neuroendocrine tumors. Clin. Cancer Res. 23, 600–609 (2017).
Paget, S. The distribution of secondary growths in cancer of the breast. 1889. Cancer Metastasis Rev. 8, 98–101 (1989).
Iacobuzio-Donahue, C. A. et al. Cancer biology as revealed by the research autopsy. Nat. Rev. Cancer 19, 686–697 (2019).
Liu, Z., Wang, H., He, J., Yuan, X. & Sun, W. Rictor ablation in BMSCs inhibits bone metastasis of TM40D cells by attenuating osteolytic destruction and CAF formation. Int. J. Biol. Sci. 15, 2448–2460 (2019).
Worner, P. M. et al. Breast tumor microenvironment can transform naive mesenchymal stem cells into tumor-forming cells in nude mice. Stem Cells Dev. 28, 341–352 (2019).
Lo Nigro, C. et al. High frequency of complex TP53 mutations in CNS metastases from breast cancer. Br. J. Cancer 106, 397–404 (2012).
Consortium, A. P. G. AACR Project GENIE: powering precision medicine through an international consortium. Cancer Discov. 7, 818–831 (2017).
Hartmaier, R. J. et al. High-throughput genomic profiling of adult solid tumors reveals novel insights into cancer pathogenesis. Cancer Res. 77, 2464–2475 (2017).
Ciriello, G. et al. Comprehensive molecular portraits of invasive lobular breast cancer. Cell 163, 506–519 (2015).
Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012).
Nik-Zainal, S. et al. Landscape of somatic mutations in 560 breast cancer whole-genome sequences. Nature 534, 47–54 (2016).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Cha, S. et al. Clinical application of genomic profiling to find druggable targets for adolescent and young adult (AYA) cancer patients with metastasis. BMC Cancer 16, 170 (2016).
Sondka, Z. et al. The COSMIC Cancer Gene Census: describing genetic dysfunction across all human cancers. Nat. Rev. Cancer 18, 696–705 (2018).
Chang, M. T. et al. Identifying recurrent mutations in cancer reveals widespread lineage diversity and mutational specificity. Nat. Biotechnol. 34, 155–163 (2016).
Gao, J. et al. 3D clusters of somatic mutations in cancer reveal numerous rare mutations as functional targets. Genome Med. 9, 4 (2017).
Cha, S., Lee, E. & Won, H.-H. Metadata record for the article: comprehensive characterization of distinct genetic alterations in metastatic breast cancer across various metastatic sites. figshare https://doi.org/10.6084/m9.figshare.14798541 (2021).
dbGAP. Foundation Medicine Adult Cancer Clinical Dataset (FM-AD). https://identifiers.org/dbgap:phs001179.v1.p1. (2017).
Mayakonda, A., Lin, D. C., Assenov, Y., Plass, C. & Koeffler, H. P. Maftools: efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 28, 1747–1756 (2018).
Acknowledgements
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Ministry of Science and ICT (Information and Communication Technologies) of Korea government (NRF-2019R1A2C4070496) to HW and supported by the Basic Science Research Program (Research Fellow) through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (NRF-2017R1A6A3A11030461) to S.C.; E.L. was supported by the Cornell University Start-up funds and the Nancy and Peter Meinig Family Investigator funds. We thank the participants and investigators of the GENIE, MSK-IMPACT, FMAD, INSERM, WTSI, and TCGA-BRCA cohorts for providing genomic and clinical data.
Author information
Authors and Affiliations
Contributions
S.C. and H.H.W. designed the research. S.C. conducted data analyses. S.C. and H.H.W. wrote the manuscript. E.L. revised the manuscript. All authors approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cha, S., Lee, E. & Won, HH. Comprehensive characterization of distinct genetic alterations in metastatic breast cancer across various metastatic sites. npj Breast Cancer 7, 93 (2021). https://doi.org/10.1038/s41523-021-00303-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41523-021-00303-y
This article is cited by
-
Systematic pan-cancer analysis identifies SLC31A1 as a biomarker in multiple tumor types
BMC Medical Genomics (2023)
-
Interplay between ESR1/PIK3CA codon variants, oncogenic pathway alterations and clinical phenotype in patients with metastatic breast cancer (MBC): comprehensive circulating tumor DNA (ctDNA) analysis
Breast Cancer Research (2023)
-
SWI/SNF complex gene variations are associated with a higher tumor mutational burden and a better response to immune checkpoint inhibitor treatment: a pan-cancer analysis of next-generation sequencing data corresponding to 4591 cases
Cancer Cell International (2022)
-
Comprehensive analyses reveal the carcinogenic and immunological roles of ANLN in human cancers
Cancer Cell International (2022)