Abstract
Pseudomonas aeruginosa tends to be among the dominant species in multi-species bacterial consortia in diverse environments. To understand P. aeruginosa’s physiology and interactions with co-existing bacterial species in different conditions, we established physiologically reproducible 18 species communities, and found that P. aeruginosa dominated in mixed-species biofilm communities but not in planktonic communities. P. aeruginosa’s H1 type VI secretion system was highly induced in mixed-species biofilm consortia, compared with its monospecies biofilm, which was further demonstrated to play a key role in P. aeruginosa's enhanced fitness over other bacterial species. In addition, the type IV pili and Psl exopolysaccharide were required for P. aeruginosa to compete with other bacterial species in the biofilm community. Our study showed that the physiology of P. aeruginosa is strongly affected by interspecies interactions, and both biofilm determinants and type VI secretion system contribute to higher P. aeruginosa's fitness over other species in complex biofilm communities.
Similar content being viewed by others
Introduction
Biofilms can protect bacteria from hostile environments and also disperse bacteria to colonise in new niches.1,2 Pseudomonas aeruginosa is a model organism for studying biofilm formation in Gram-negative bacteria.1,3 The biofilm mode affords P. aeruginosa protection and renders it more difficult to control.4,5 Type IV pili (T4P), flagella, iron, extracellular DNA and Pel/Psl exopolysaccharides (EPS) are well known factors that contribute to the development of P. aeruginosa monospecies biofilms.6,7,8 The adaptability of P. aeruginosa to various environments, such as water, soil, sewage and plants, suggests its strong competitiveness for ecological niches.6,9,10 For example, P. aeruginosa is a dominant cultivable endophytic bacterium associated with Pennisetum glaucum (millet).11
The broad-spectrum adaptability of P. aeruginosa also hints at its strong competitiveness against various other bacterial species when in mixed-species microbial communities. Several studies have indicated that P. aeruginosa is able to gain fitness over competing species during its survival in either two-species or three-species co-cultures.12,13,14 For example, P. aeruginosa reduces the viability of Staphylococcus aureus and lyses S. aureus cells for iron repletion in planktonic co-cultures.15,16 In biofilms, P. aeruginosa inhibits Staphylococcus epidermidis growth and reduces the initial adhesion of Agrobacterium tumefaciens through the secretion of Psl and Pel exopolysaccharides and small diffusible molecules.17,18,19 P. aeruginosa decreases the swarming motility of P. putida and therefore inhibits its biofilm formation by producing 2-heptyl-3-hydroxy-4-quinolone.20 Quinolones secreted by P. aeruginosa can also reduce biofilm maturation for a broad range of bacteria.21 These studies demonstrate the competitiveness of P. aeruginosa in mixed-species interactions, from metabolic versatility to a strong biofilm formation capacity. Documented strategies in which P. aeruginosa exerts competition involve type VI secretion system (T6SS), generation of antibiotics, iron chelators, pyoverdine, rhamnolipid, pyocyanin, extracellular polysaccharide, fatty acid cis-2-decenoic acid, proteinase and other quorum-sensing system-regulated virulence factors.22,23,24,25,26,27
P. aeruginosa co-exists with many different microbial species in natural settings. However, the main competitive advantage of P. aeruginosa in gaining fitness over other species within complex microbial communities remains unclear. Previous studies indicated that interspecies interactions in mixed-species microbial communities are complicated and involve both cooperation and competition.24,28 Therefore, monitoring the population dynamics of P. aeruginosa in complex microbial communities is important for elucidating how this interaction network is established, which may provide insights into its ecological role in complex mix-species biofilms.
Until recently, there has been a lack of robust tools to monitor the population dynamics in complex microbial communities. The high-throughput digital NanoString nCounter® system has originally been developed to profile the eukaryotic multiplexed gene expression with flexibility and sensitivity.29,30 However, this technology has never been reported for its utilisation on monitoring population dynamics in a complex microbial community. Here we modified the probes to achieve a new application in investigation microbial ecology and evaluated this technology in monitoring population dynamics in a complex laboratory mixed-species microbial community with pairs of tailored probes directly hybridise to the 16 S rRNA of each species instead of mRNA without any enzymatic reaction or DNA synthesis.29 We further investigated the transcriptome of P. aeruginosa competition in this mixed-species community in both planktonic and biofilm modes of growth.
Results
Establishment of the complex microbial community
Instead of proliferating as a single-species culture, P. aeruginosa often grows as commensal species within mixed-species microbial communities containing many other bacterial species in natural environments as well as infection sites.31,32 We presumed that bacteria within the same region of colonisation would have the same chance to co-exist. To obtain a better understanding of P. aeruginosa physiology in complex microbial communities and evaluate the NanoString nCounter® system, we selected bacterial species that potentially co-exist with P. aeruginosa in different environments and can be distinguished from other species by NanoString nCounter® probes. Hence, 18 human pathogens/environmental bacteria (Supplementary Table 1) were established as laboratory planktonic and biofilm communities as some of these species usually share the same niche in humans or in natural environments. Among these species, Acinetobacter baumannii, Burkholderia cenocepacia, Klebsiella pneumoniae and S. aureus can co-infect lungs with P. aeruginosa33,34,35,36; Citrobacter amalonaticus, E. coli and Streptococcus gallolyticus can cause intra-abdominal infections; Chromobacterium violaceum, S. aureus and Stenotrophomonas maltophilia can infect the skin or open wounds; Listeria monocytogenes can colonise a host’s gastrointestinal tract; P. putida has been isolated from infected urinary tract and infected skin; Bacillus subtilis, Elizabethkingia meningoseptica, P. fluorescens and Shewanella oneidensis are widely distributed in natural environments; and P. syringae and Xanthomonas campestris have been associated with plants.37 The abovementioned settings all involve the potential co-existence of P. aeruginosa. Both known and undiscovered natural environments are so complex and unexpected. This laboratory community may assist in understanding P. aeruginosa’s interactions with complex microbial communities. In addition to investigating the ecological behaviour and physiological response of P. aeruginosa in our model system, we also seek insights into how future studies on complex microbial communities can be shaped.
We used 10 times diluted tryptic soy broth (10% TSB) as the growth medium, which supports the growth of all the selected 18 bacterial species at 25 °C (room temperature) (Supplementary Figure 1). Although A. baumannii, C. amalonaticus, C. violaceum, E. meningoseptica, E. coli and K. pneumoniae grew relatively faster than other species, and L. monocytogenes and S. gallolyticus were unable to grow to a high cell density in 10% TSB, the growth curves indicated that 10% TSB was suitable for establishing the mixed-species microbial community, with P. aeruginosa growth comparable to the majority of the species (Supplementary Figure 1).
In addition to the growth rate, the biofilm formation capacity of these bacterial species was also verified in monospecies culture. After 24 h of static incubation, A. baumannii and K. pneumoniae had the highest capacity to form biofilms (Supplementary Figure 2). Among the other 16 species, B. cenocepacia, P. aeruginosa and S. maltophilia have similar biofilm formation capabilities (Supplementary Figure 2). However, B. subtilis, C. violaceum, E. meningoseptica, P. putida, P. syringae and S. gallolyticus formed less biofilms in 10% TSB at 25°C (Supplementary Figure 2). In summary, the biofilm formation capacities of the 18 species differ, with no linear relationship with growth rates. We therefore established the planktonic and biofilm microbial communities by adjusting each species to an OD600 value of 0.01 in 10% TSB medium as inocula.
Physiological reproducibility of the mixed-species microbial communities
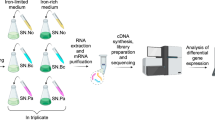
Before investigating the population dynamics and physiology of P. aeruginosa in the mixed-species communities, we employed RNA sequencing-based metatranscriptomics to examine the physiological reproducibility of both the planktonic and biofilm communities. The sequencing reads (7.8–9.2 million per sample) were assigned to 44 classified functional roles (SEED; 51.4 million reads) with unclassified reads in MEGAN6. Similar profiles of functional classifications of genes assigned to each community model by their functional properties were observed among three biological replicates (Fig. 1a), indicating that physiology is highly reproducible in our laboratory model for both planktonic and biofilm communities (Fig. 1a). The proportion of each small bar with different colours represents the ratio of different functional genes in communities. For example, expression level of 'metabolite damage and its repair or mitigation' genes is higher in biofilm communities than in planktonic communities. In contrast, expression levels of genes in 'iron acquisition and metabolism', 'cell division and cell cycle', 'secondary metabolism' are higher in planktonic communities than in biofilm communities (Fig. 1a). In addition, clear separation between the two clusters on the principal coordinates analysis (PCoA) plot confirmed that there is a significant difference between the metatranscriptome profiles of the planktonic community and the biofilm community. The close proximity among replicates of each community indicates the similarity in metatranscriptome profiles of the replicates (Fig. 1b). The relative distanced location among replicates of biofilm community is owing to the small scale chosen to display all points clearly along PC2. Above result suggests a good reproducibility in our model of mixed-species cultures which can serve as a stable system for functional analysis carried out in this study.
Metatranscriptomics comparison between mixed-species planktonic community and biofilm community modes. a Functional classification by SEED. The comparison was based on read counts with a minimum quality score cutoff of 50 and a top percentage of 25. Different colours corresponding to different functions are ranked in the right column. b Clustering by PCoA based on SEED classification using Bray–Curtis dissimilarity
Evaluation of NanoString nCounter® 16S rRNA array for microbial population assay
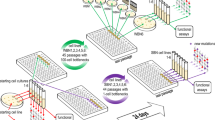
Although the 16 S rRNA gene amplicon sequencing approach is widely used to study the microbial population diversity, it has the drawback of PCR bias. A NanoString nCounter® 16 S rRNA array was developed to detect unique signals from complex hybridised samples at a single-molecule level by utilising special probes captures target 16 S rRNA sequence and omitting the PCR amplification step38 (Supplementary Table 1). To check the feasibility of the designed nCounter® 16 S rRNA array for detecting the 16 S rRNA from different bacterial species in the total RNA mixture, we mixed the total RNA extracted from 14 individual bacterial species at different ratios in three different training groups (Table 1). These were subjected to nCounter® 16 S rRNA array analysis, with the total RNA from the planktonic and biofilm communities together in one batch. The percentage of each species in 16 S rRNA reads according to nCounter® 16 S rRNA array analysis was compared with its ‘actual’ RNA percentage from the training groups to calculate the normalisation factors for each of the 14 species (Table 1). To verify this new technology, the P. aeruginosa probe was tested against the background of other genera in the mixed-species community, with the other three Pseudomonas species excluded. One Gram-positive probe was tested against the background of other genera with L. monocytogenes’s absence. The nCounter® 16 S rRNA array yielded more reads to some species than accurate 16 S rRNA readings, whereas the converse occurred for other species (Table 1). This trend highlighted the need to analyse bacterial population dynamics by multiplying the normalisation factors (the variation fold) (Table 1), when nCounter® 16 S rRNA array is employed to reflect the true percentage of each bacterial species in a mixed-species community.
P. aeruginosa is the dominant species in the mixed-species biofilm community
To maintain the reproducibility of this laboratory model, we used the same mixed-species community. Each individual species culture was normalised to an OD600 reading of 0.01 in the mixture and cultivated for 1 d to establish the planktonic community, and 5 d to establish the biofilm community. Total RNA extracted from these communities were analysed by nCounter® 16 S rRNA array. In the 1-day-old planktonic community, the relative abundance of A. baumannii, C. violaceum and E. meningoseptica increased by twofold or more compared with their inocula, whereas the relative abundance of B. subtilis, P. aeruginosa, S. oneidensis, S. aureus, S. gallolyticus and X. campestris decreased to half or less compared with their inocula (Fig. 2 and Supplementary Table 2). Interestingly, the microbial population dynamics are quite different in the 5-day-old biofilm community compared with the planktonic community; the relative abundance of K. pneumoniae and P. aeruginosa increased to more than fourfold their initial inputs, whereas most of the other species decreased significantly (Fig. 2 and Supplementary Table 2). P. aeruginosa 16 S rRNA constituted ~32.62% of the total 16 S rRNA content, whereas K. pneumoniae constituted ~26.08% of that in 5-day-old biofilms (Fig. 2, and Supplementary Table 2) after normalisation factor adjustment (Table 1). The proportions of all bacterial species (including the four non-normalised ones) are summarised in Supplementary Figure 3.
Normalised population dynamics of each species in the mixed-species microbial communities. The inocula is the mixture of all species mixed at equal OD600 amounts at the starting point for both planktonic and biofilm incubation. Grey bars indicate the proportion of each species in the 24-hour planktonic microbial community. The black bars show the population dynamics of each species in the 5-day biofilm microbial community. Dots show three biological replicates for standard deviation
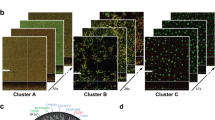
To validate the observed microbial population dynamics, we used GFP-tagged P. aeruginosa to establish the 1-day-old planktonic community and the 5-day-old biofilm community as described above. The abundance of P. aeruginosa was detected and calculated using the fluorescence-proportion standard curve (Supplementary Figure 4). The fluorescence-proportion standard curve is preferred over the colony-forming unit counting approach owing to the difference in the growth rates and the colony sizes of these 18 bacterial species on plates, which contributes to the accuracy of dynamics population counting. Resuspended cells from both planktonic and biofilm communities were used for fluorescence-based proportion tests and next examined using confocal microscopy imaging (Fig. 3b, c). The results of the biological fluorescence-proportion standard curve-based experiment correlated well with the results of the normalised nCounter® 16 S rRNA array (Fig. 3a). The confocal images analysis further verified the proportion of P. aeruginosa cells in the mixed-species biofilm community (Fig. 3c) is much higher than in the planktonic community (Fig. 3b). Together with the results in Fig. 2, we have demonstrated that P. aeruginosa is the dominant species in this microbial biofilm community.
Proportions of P. aeruginosa in the mixed-species microbial communities. a Circular dots indicate three biological replicates of P. aeruginosa normalised proportions in both the planktonic and biofilm communities as measured by NanoString nCounter® 16 S rRNA array; the square dots show three biological replicates of P. aeruginosa proportions in mixed-species microbial communities as calculated by fluorescence-based proportion test. The lines show the average values and standard deviation. Comparison was performed by t test with p < 0.05. b Confocal images of the resuspended 24 h-incubated mixed-species planktonic microbial community containing GFP-tagged P. aeruginosa. c Confocal images of the resuspended 5-day mixed-species biofilm community containing GFP-tagged P. aeruginosa. Green cells show the GFP-tagged P. aeruginosa, red cells show all of other cells stained with Syto62. Scale bar represents 10 µm
Both the H1 type VI secretion system and biofilm formation determinants contributed to the fitness of P. aeruginosa in the biofilm community
As P. aeruginosa constituted up to 32.62% of the biofilm community in contrast to only 3.81% in the planktonic community (Supplementary Table 2), we hypothesised that the important biofilm determinants are likely to contribute to the dominating competence of P. aeruginosa over other species in the biofilm community. To test this hypothesis, we performed a transcriptomics analysis comparing the gene expressions in monospecies P. aeruginosa biofilms with that of the mixed-species biofilm community. Differentially expressed genes of P. aeruginosa in mixed-species biofilm and monospecies biofilm are illustrated as a heat map (Fig. 4a). Further clustering of the transcriptome by principle component analysis revealed that the P. aeruginosa maintained a distinct physiology in monospecific and mixed-species biofilms (Fig. 4b). Based on the negative binomial test with an adjusted P value cutoff of 0.05 and a log2-fold change cutoff of 1,239 genes were upregulated and 171 genes were downregulated in P. aeruginosa cells in the mixed-species biofilm compared to those in monospecific biofilms (Supplementary Table 3).
Transcriptomic comparison of P. aeruginosa between monospecies biofilm cells and the mixed-species biofilm community. a A heat map representing P. aeruginosa genes expression level up- or downregulated more than twofold in the mixed-species biofilm compared with the monospecies biofilm. The Z score shows the standardisation of each gene expression level between the two different groups. Detailed information on this regulation is listed in Supplementary Table 3. b A principal component analysis (PCA) shows that the expression profiles of P. aeruginosa between the monospecies biofilm and in the mixed-species biofilm are different. Each dot indicates one biological replicate with different colours representing the sample source
Among the significantly upregulated genes of P. aeruginosa in the mixed-species biofilm, at least 36 belonged to the T6SS (Table 2), which accounted for 15% of all the upregulated genes. Most of these belong to the Hcp secretion island I (H1) T6SS, whereas some are located in the Hcp secretion island II (H2) T6SS. However, there were only a few genes that belong to H3-T6SS. The ClpV protein is the energy source of T6SS39; the expressions of clpV1, clpV2 and clpV3 belonging to three different sub-type T6SS in P. aeruginosa were all upregulated in the mixed-species biofilm (Table 2). To validate the transcriptomic analysis results, the ΔclpV1, ΔclpV2 and ΔclpV3 mutants of P. aeruginosa were constructed with a GFP tag and thereafter formed biofilms with the other 17 species. Interestingly, only the ΔclpV1 mutant was greatly impaired in terms of fitness to dominate in the mixed-species biofilms; when ClpV1 was complemented into this mutant with multiple copies, the complementation strain showed higher fitness to dominate in the mixed-species biofilm community than both wild-type PAO1 and the deletion mutant (Fig. 5). This finding suggests that the H1-T6SS has a significant effect on fitness gains for P. aeruginosa, enabling it to outcompete other species in the mixed biofilm community (Fig. 5). In addition, confocal images confirmed that parental strain PAO1 gains higher fitness over the other bacterial species than its isogenic H1-T6SS defect mutant ΔclpV1 in the 18 species biofilm communities (Supplementary Figure 5c and 5d). H1-T6SS contains seven effectors to be secreted into its competitor, which are termed Tse1-7.40,41 Both Tse1 and Tse3, which can lyse the target cells by degrading their peptidoglycan,26 were upregulated in the complex microbial biofilm community (Table 2). Furthermore, the expression of the gene encoding the H1-T6SS delivery-dependent proteins VgrG1 was also increased (Table 2).
P. aeruginosa population dynamics in the mixed-species biofilm community affected by H1-T6SS and biofilm formation determinants. Each column shows the population dynamics of GFP-tagged P. aeruginosa parental strain PAO1 and the knockout mutants in the mixed-species microbial planktonic (black) and biofilm (grey) communities at different heights with the standard deviation of three biological replicates (dots). The asterisk indicates statistical significance by t test with p < 0.05
In addition to T6SS, the P. aeruginosa pelF, pelG, flgC and flgD genes, which are important for biofilm formation in monospecies cultures, were also upregulated in the mixed-species biofilms (Table 2). We next investigated the impact of the classic monospecies P. aeruginosa biofilm determinants, e.g., Pel and Psl EPS,42,43,44 T4P8,45 and flagella,8,45 on the fitness of P. aeruginosa in the mixed-species biofilms. The P. aeruginosa deletion mutants ΔpslBCD, ΔpelA, ΔpilA or ΔfliM were used to cultivate mixed-species biofilms with the other 17 bacterial species respectively. The Psl polysaccharide and T4P were required for P. aeruginosa to predominate in the mixed-species biofilm community (Fig. 5). Psl was particularly essential for bacterial colonisation in the mixed-species biofilms,46 and ΔpslBCD was completely outcompeted by other species in the biofilms (Fig. 5), even though its planktonic growth was similar to that of the wild-type PAO1 strain (Supplementary Figure 6).
Quorum sensing is not required for the fitness of P. aeruginosa in the mixed-species biofilm community
Quorum-sensing systems play important roles in biofilm formation both in vitro and in vivo.47,48,49,50,51,52 P. aeruginosa is able to outcompete other microbial species by releasing quorum sensing-regulated virulence factors, such as rhamnolipid, elastase, exotoxins, pyocyanin and pyoverdine.53,54,55,56,57 However, the expression of the P. aeruginosa quorum sensing-related genes, such as lasA, lasB, rhlA, rhlB and pqsH, were downregulated in this biofilm community compared with the monospecies biofilm (Table 2), suggesting that quorum sensing is not important in contributing to the fitness of P. aeruginosa in the mixed-species community biofilm. Intriguingly, deleting the lasR or mvfR quorum-sensing genes in P. aeruginosa even increased this species’ fitness over other species in the mixed-species biofilms, compared to the PAO1 wild-type (Fig. 5). This could possibly be owing to P. aeruginosa H1-T6SS being repressed by LasR and MvfR.58
Discussion
Our study reported on the establishment of physiologically reproducible laboratory planktonic and biofilm communities and their use in investigating how P. aeruginosa competes with multiple other bacterial species. We tracked the population dynamics and profiled the transcriptomes of the established communities and showed that P. aeruginosa maintains distinct transcriptomes in complex microbial communities compared to its monospecies cultures (Fig. 4). The H1-T6SS, Psl and T4P were found to be the key components for P. aeruginosa to gain fitness over other species in the mixed-species biofilm communities (Fig. 5 and Table 2). These pathway-involved genes, such as clpV1 (6.8-fold), pilA (17.1-fold) and the pslBCD operon, were all significantly upregulated in the microbial biofilm community (P. aeruginosa is the dominant species) compared with the planktonic cultures (P. aeruginosa is not the dominant species). Our results highlight the biological significance of biofilm formation for P. aeruginosa to gain fitness in complex microbial communities,59 and we confirmed that H1-T6SS is a powerful weapon for P. aeruginosa interspecies competition within the ecological niche in bacterial communities.39,41 Confocal imaging analysis for dual-species biofilms also showed that parental strain PAO1 had a higher population than the H1-T6SS-defective mutant ΔclpV1 when competing against A. baumannii, C. violaceum, C. amalonaticus and S. maltophilia while E. coli as control (Supplementary Figure 5e–5n). Our findings on P. aeruginosa in mixed-species microbial communities are in accordance with previous studies in pure culture or two-species consortia18,19,26,40 and suggest that Psl, T4P and H1-T6SS are potential targets to control or eliminate P. aeruginosa from complex microbial biofilm communities, such as microbial biofilms on medical devices.
In addition, we demonstrated the sensitivity of the single molecule detection technique, NanoString nCounter® 16 S rRNA array, in monitoring the population dynamics in complex bacterial communities. Our results showed that the accuracy of this technique depends on the specificity and affinity of the probes, which need to be normalised during the calculation of population dynamics.
P. aeruginosa can dominate airway infection microbiota60 and wound communities after the administration of antibiotics,61 suggesting that P. aeruginosa has a competitive physiology in mixed-species microbial communities. Our transcriptomic analysis showed that two of the primary mechanisms for P. aeruginosa maintaining this competitive advantage are H1-T6SS and biofilm formation. This differs from most previous reports in which many quorum-sensing-regulated virulence factors, such as elastase, PQS, pyoverdine and pyocyanin, were used by P. aeruginosa to kill other bacteria.53,54,55,56,57 The reasons for this difference may be as follows: (1) P. aeruginosa las and pqs systems repress H1-T6SS to compete with other bacteria58; (2) quorum-sensing-signalling molecules and virulence factors might function as signals that induce aggressive phenotypes from bacterial competitors as well as hosts via cross-talk, which in turn arrests the growth of P. aeruginosa; (3) the high production of quorum-sensing-regulated virulence factors might consume a great deal of energy to reduce the division of P. aeruginosa cells or even trigger autolysis phenotypes62; and/or (4) the continued fresh medium stream of a flow-tube biofilm cultivation system might wash away the secreted quorum sensing signals and virulence factors and thus reduce the efficacy of quorum sensing. Thus, the cell-contact based T6SS might be the most efficient mechanism for P. aeruginosa to outcompete other species. This is supported by our data that loss of ClpV1 in H1-T6SS will decrease the competitive capacity of P. aeruginosa in our laboratory model (Fig. 5). Recently, Allsopp et al.63 has shown that all three T6SSs in P. aeruginosa are more highly expressed at 25°C than at 37°C, however, expressed H2-T6SS and H3-T6SS did not affect P. aeruginosa competition capacity as much as H1-T6SS at 25°C in our experiment (Fig. 5).
H1-T6SS is well known for its roles in competitiveness and pathogenicity in P. aeruginosa.26,39 In addition to H1-T6SS, Psl and T4P are critical components of P. aeruginosa fitness in the mixed-species biofilm community (Fig. 5). Our result is in accordance with previous research showing that secreted protein A from S. aureus can inhibit P. aeruginosa biofilm formation by binding to Psl and T4P.64 In mixed-species biofilm communities, three P. aeruginosa’s most upregulated genes, PA3661, PA2205 and PA1395, are encoding small lipoproteins (Supplementary Table 3). Wood et al.65 reported P. aeruginosa’s small lipoproteins genes upregulation under cell wall stress, which is regulated by sigma factor AlgU. However, expression of algU changed less than twofold in our study, suggesting that P. aeruginosa may undergoes AlgU-independent stress response in the mixed-species biofilm communities. One limitation of our current study is that we have not elucidated whether P. aeruginosa employs T6SS, Psl and T4P sequentially or simultaneously when outcompeting the other species in biofilm communities. In addition, further study is required to validate whether the T6SS, Psl and T4P are required by P. aeruginosa to outcompete other species in vivo. Because the amounts of bacteria from in vivo samples are usually low and insufficient for transcriptomic analysis, suitable probes for the NanoString nCounter® array can be explored to examine the population dynamics and gene expression of P. aeruginosa from the in vivo communities. Moreover, K. pneumonia is the secondary dominant species in our biofilm co-cultures but not in planktonic co-cultures (Fig. 2), and it constantly co-exists with P. aeruginosa in vitro and in vivo.66 Thus, it would be interesting to ascertain its survival mechanism in future investigations.
Methods
Bacterial strains, media and growth conditions
The bacterial species used in this study are listed in Supplementary Table 1. All strains were revived on TSB agar plates and subsequently in TSB liquid medium at 30 °C. Mixed-species communities were cultivated in 10% TSB at 25 °C. To construct P. aeruginosa gene knockout mutants and green fluorescent protein (GFP)-tagged strains, AB minimal medium67 supplemented with 10 mM citric acid supplemented with appropriate antibiotics was used to select P. aeruginosa. E. coli was grown in LB medium at 37 °C. In total, 60 µg/ml of gentamicin was used to select P. aeruginosa, whereas 15 µg/ml of gentamicin, 10 µg/ml of chloramphenicol and 100 µg/ml of ampicillin were used to select E. coli, when appropriate.
Growth assay and static biofilm cultivation
Overnight cultures of each bacterial species were diluted to OD600 of 0.01 in 10% TSB, and 150 µl diluted culture were loaded into 96-well microtitre plates. The plates were incubated in an infinite M200PRO multimode microplate reader (Tecan Schweiz AG, Männedorf, Switzerland) at 25°C, and the OD600 of each well was measured and recorded hourly. A 10% TSB blank medium was used as a negative control, and its reading has been subtracted from all the culture readings to plot the growth curves for each species.
To examine the static biofilm-forming capacities of each species, bacterial cultures were prepared using the same procedures as for the growth assay. After overnight cultivation, the planktonic cells were discarded from the microtitre plate, and the wells were washed three times using tap water to remove any residual planktonic cells. A 180 µl volume of 0.1% crystal violet (CV) solution was loaded into each well to stain the biofilm for 15 min. The excess stain in the wells was washed out with tap water three times. Finally, 180 µl of 30% acetate acid was loaded into each well to dissolve CV from the stained biofilm. The optical intensity at OD550 was measured by an Infinite M200PRO multimode microplate reader to quantify the biofilms formed by each bacterial species.
Cultivation of the mixed-species planktonic and biofilm communities
Overnight cultures of the 18 individual bacterial species were diluted in 10% TSB medium and normalised to the same ratio (OD600 of each species is ~0.01) to form the initial inoculum for both planktonic cultures in shake flask and biofilm cultures in flow-tube reactors. The planktonic cultures were shaking with 200 rpm at 25°C. The biofilm cultivation was started from initial inoculum attachment to MasterflexTM silicone tube; then fresh 10% TSB medium was continually pumped to flow through these attached cells in silicone tube at 4 ml/h at 25 °C. The 4 ml/h flow will supply enough nutrients for biofilm maturity and take away the planktonic cells in silicone tube. The 24-hour-old planktonic cultures and 5-day-old biofilm cultures were harvested for population dynamic or transcriptomic analysis.
RNA preparation for sequencing
The bacterial cells were first treated with RNA Protect Reagent (Qiagen®, Germany) to maintain the integrity of RNA. The total RNA was extracted from these bacterial cells using a miRNeasy Mini Kit (Qiagen®, Germany) with modifications. A Turbo DNA-free Kit (Thermo Fisher Scientific®, Lithuania) was used to remove genomic DNA contaminants from total RNA. DNA contamination was assessed with a Qubit® dsDNA High Sensitivity assay (PicoGreen dye) and a Qubit® 2.0 Fluorometer (Invitrogen®, Austria) according to manufacturer’s instructions. Ribosomal RNA was depleted with a Ribo-Zero rRNA removal Kit (Illumina, USA). The integrity of the total RNA was assessed with an Agilent TapeStation System (Agilent Technologies, UK).
The double-stranded coplementary DNAs (cDNAs) were reverse-transcribed using a NEBNext RNA first and second strand synthesis module (NEB®, USA). cDNAs were subjected to Illumina’s TruSeq Stranded mRNA protocol. The quantitated libraries were then pooled at equimolar concentrations and sequenced on an Illumina HiSeq2500 sequencer in rapid mode at a read-length of 100 bp paired-ends.
NanoString nCounter® population analysis
One nanogram of purified total RNA from the different bacterial cultures was analysed using a NanoString nCounter® Gene Expression CodeSets platform (NanoString Technologies, Inc., United States) with the customised CodeSets for our selected bacterial species (Supplementary Table 1), according to the manufacturer’s protocol, to obtain raw counts. The quality control assessment and raw counts were normalised and analysed using nSolver™ analysis software version 2.6 (NanoString Technologies, Inc., United States). The samples were analysed using the manufacturer-recommended default parameters. The geometric mean was selected for negative control subtraction; if the normalisation factors were outside the range 0.3–3, the normalised factor and the flag lane were computed by the geometric mean. The experiments were performed in triplicate, and the results are presented as the means ± S.D.
Transcriptomic analysis
The accession number for RNA-seq is SRP128411. The PAO1 genome (NC_002516) was used as the reference for P. aeruginosa transcriptomics analysis, with annotation from the Pseudomonas Genome Database (http://www.pseudomonas.com/). The RNA-Seq raw data were analysed using the “RNA-Seq and expression analysis” application in CLC genomics Workbench 10.0 (QIAGEN). The total gene reads from CLC genomics Workbench 10.0 were subjected to the DESeq2 package for statistical analysis68 by using R/Bioconductor.69 A hierarchical clustering analysis was performed with a negative binomial test using the DESeq2 package. The heatmap.2 package was used to draw a heat map for the differentially expressed genes of P. aeruginosa cells with fold change larger than two and an adjusted p value smaller than 0.05. The principal component analysis plot was generated in R/Bioconductor.
The adapter-trimmed and assembled RNA sequences of the mixed-species microbial community were used as metatranscriptomics data for analysis. The sequences of the mixed-species communities grown in biofilm and planktonic modes were aligned against the NCBI non-redundant protein database using DIAMOND with default settings. The output aligned gene reads in DAA format were uploaded and ‘meganised’ in MEGAN6.11.1 with a minimum bit-score of 50 and a top percentage of 25.70 Functional analyses of these aligned gene reads were performed using SEED classifications. A PCoA was plotted to cluster the samples based on functions. The functional analysis results are illustrated in stacked bar charts.
Fluorescence-based P. aeruginosa population assay
To develop a fast and economical P. aeruginosa population dynamics assay in the mixed-species cultures, a single copy of a GFP tag was inserted into the P. aeruginosa chromosome by using a mini-Tn7-Gm-gfp transposon.71 The other 17 bacterial species without P. aeruginosa were collectively used as a background control. The GFP fluorescence readings of both planktonic and biofilm cultures, with gradient proportion of GFP-tagged P. aeruginosa derived strains, were recorded using a Tecan infinite M200PRO microplate reader with excitation wavelength 485 nm and emission wavelength 535 nm. The fluorescence-proportion standard curves of each tested strain were generated from these GFP readings (Supplementary Figure 4). The proportions of P. aeruginosa derived strains in communities were calculated using these fluorescence-proportion standard curves. All of coefficient of determinations (R2) of these linear regression lines are larger than 0.97, suggesting that the fluorescence-based population assay is a reliable method to detect P. aeruginosa proportions in our mixed-species communities.
Constructions of P. aeruginosa gene deletion and complementation mutants
The upstream fragment of target gene was amplified by primer-1 and primer-2, the downstream fragment of the target gene was amplified by primer-3 and primer-4 (Supplementary Table 4), and then both fragments were fused into a pK18-Gm-mobsacB plasmid72 and transformed into the P. aeruginosa PAO1 wild-type strain to delete the target gene as previously described.72 PCR and sequencing were used to confirm the selected mutant strains (Supplementary Table 5).
To complement the P. aeruginosa clpV1 mutant, a DNA fragment from the clpV1 upstream 22 bps to the downstream 31 bps was amplified by primers clpV1-up and clpV1-down (Supplementary Table 4) and double-digested using BamHI and HindIII. The purified fragment was ligated with the pUCP22 vector transcribed by a lacZ promoter,73 which was digested by the same restriction enzymes to construct the pUCP22-clpV1 complementary plasmid. After sequencing verification, the pUCP22-clpV1 plasmid was transformed into the P. aeruginosa ΔclpV1 mutant by electroporation to construct a clpV1 complementation strain.
Confocal microscopy imaging
P. aeruginosa PAO1-GFP-containing planktonic cultures and biofilm cultures (the same samples as those used for the fluorescence-based population assay) were stained with 5 µm SYTOTM 62 Red fluorescent nucleic acid stain (molecular probe) for 15 min to stain all cells before 5 µl of each sample was aliquoted onto a glass slide and covered with a cover slip. For supplementary figure 5, 5 d biofilm cells were scraped from silicone tube into imaging chamber to stain all cells with SYTOTM 62 Red fluorescent nucleic acid stain, because silicone tube is too thick for directly confocal imaging. The samples were visualised and confocal images acquired using confocal laser scanning microscopy (CLSM; Zeiss LSM 780, Carl Zeiss, Germany) with a 63 × 1.40 DICII objective. Green and red fluorescence were observed upon 488 nm and 633 nm excitation wavelengths, respectively.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request. The accession number for RNA-seq is SRP128411.
References
Hall-Stoodley, L., Costerton, J. W. & Stoodley, P. Bacterial biofilms: from the natural environment to infectious diseases. Nat. Rev. Microbiol. 2, 95–108 (2004).
Flemming, H. C. & Wingender, J. The biofilm matrix. Nat. Rev. Microbiol. 8, 623–633 (2010).
Costerton, J. W., Stewart, P. S. & Greenberg, E. P. Bacterial biofilms: a common cause of persistent infections. Science 284, 1318–1322 (1999).
Mulcahy, L. R., Isabella, V. M. & Lewis, K. Pseudomonas aeruginosa biofilms in disease. Microb. Ecol. 68, 1–12 (2014).
Lewis, K. Riddle of biofilm resistance. Antimicrob. Agents Chemother. 45, 999–1007 (2001).
Harmsen, M., Yang, L., Pamp, S. J. & Tolker-Nielsen, T. An update on Pseudomonas aeruginosa biofilm formation, tolerance, and dispersal. FEMS Immunol. Med. Microbiol. 59, 253–268 (2010).
Banin, E., Vasil, M. L. & Greenberg, E. P. Iron and Pseudomonas aeruginosa biofilm formation. Proc. Natl. Acad. Sci. USA 102, 11076–11081 (2005).
O’Toole, G. A. & Kolter, R. Flagellar and twitching motility are necessary for Pseudomonas aeruginosa biofilm development. Mol. Microbiol. 30, 295–304 (1998).
Hardalo, C. & Edberg, S. C. Pseudomonas aeruginosa: assessment of risk from drinking water. Crit. Rev. Microbiol. 23, 47–75 (1997).
Walker, T. S. et al. Pseudomonas aeruginosa-plant root interactions. Pathogenicity, biofilm formation, and root exudation. Plant Physiol. 134, 320–331 (2004).
Gupta, G., Panwar, J. & Jha, P. N. Natural occurrence of Pseudomonas aeruginosa, a dominant cultivable diazotrophic endophytic bacterium colonizing Pennisetum glaucum (L.) R. Br. Appl. Soil Ecol. 64, 252–261 (2013).
Beaume, M. et al. Metabolic pathways of Pseudomonas aeruginosa involved in competition with respiratory bacterial pathogens. Front. Microbiol. 6, 321 (2015).
Hotterbeekx, A., Kumar-Singh, S., Goossens, H. & Malhotra-Kumar, S. In vivo and In vitro Interactions between Pseudomonas aeruginosa and Staphylococcus spp. Front. Cell Infect. Microbiol 7, 106 (2017).
Hogan, D. A. & Kolter, R. Pseudomonas-Candida interactions: an ecological role for virulence factors. Science 296, 2229–2232 (2002).
Mashburn, L. M., Jett, A. M., Akins, D. R. & Whiteley, M. Staphylococcus aureus serves as an iron source for Pseudomonas aeruginosa during in vivo coculture. J. Bacteriol. 187, 554–566 (2005).
Filkins, L. M. et al. Coculture of Staphylococcus aureus with Pseudomonas aeruginosa Drives S-aureus towards fermentative metabolism and reduced viability in a cystic fibrosis model. J. Bacteriol. 197, 2252–2264 (2015).
Mowat, E. et al. Pseudomonas aeruginosa and their small diffusible extracellular molecules inhibit Aspergillus fumigatus biofilm formation. FEMS Microbiol. Lett. 313, 96–102 (2010).
Qin, Z. Q., Yang, L., Qu, D., Molin, S. & Tolker-Nielsen, T. Pseudomonas aeruginosa extracellular products inhibit staphylococcal growth, and disrupt established biofilms produced by Staphylococcus epidermidis. Microbiology 155, 2148–2156 (2009).
An, D. D., Danhorn, T., Fuqua, C. & Parsek, M. R. Quorum sensing and motility mediate interactions between Pseudomonas aeruginosa and Agrobacterium tumefaciens in biofilm cocultures. Proc. Natl. Acad. Sci. USA 103, 3828–3833 (2006).
Fernandez-Pinar, R., Camara, M., Dubern, J. F., Ramos, J. L. & Espinosa-Urgel, M. The Pseudomonas aeruginosa quinolone quorum sensing signal alters the multicellular behaviour of Pseudomonas putida KT2440. Res. Microbiol. 162, 773–781 (2011).
Reen, F. J. et al. The Pseudomonas quinolone signal (PQS), and its precursor HHQ, modulate interspecies and interkingdom behaviour. FEMS Microbiol. Ecol. 77, 413–428 (2011).
Irie, Y., O’Toole, G. A. & Yuk, M. H. Pseudomonas aeruginosa rhamnolipids disperse Bordetella bronchiseptica biofilms. FEMS Microbiol. Lett. 250, 237–243 (2005).
Davies, D. G. & Marques, C. N. H. A fatty acid messenger is responsible for inducing dispersion in microbial biofilms. J. Bacteriol. 191, 1393–1403 (2009).
Hibbing, M. E., Fuqua, C., Parsek, M. R. & Peterson, S. B. Bacterial competition: surviving and thriving in the microbial jungle. Nat. Rev. Microbiol. 8, 15–25 (2010).
Dietrich, L. E. P., Teal, T. K., Price-Whelan, A. & Newman, D. K. Redox-active antibiotics control gene expression and community behavior in divergent bacteria. Science 321, 1203–1206 (2008).
Russell, A. B. et al. Type VI secretion delivers bacteriolytic effectors to target cells. Nature 475, 343–347 (2011).
Trejo-Hernandez, A., Andrade-Dominguez, A., Hernandez, M. & Encarnacion, S. Interspecies competition triggers virulence and mutability in Candida albicans-Pseudomonas aeruginosa mixed biofilms. ISME J. 8, 1974–1988 (2014).
Elias, S. & Banin, E. Multi-species biofilms: living with friendly neighbors. FEMS Microbiol. Rev. 36, 990–1004 (2012).
Fortina, P. & Surrey, S. Digital mRNA profiling. Nat. Biotechnol. 26, 293–294 (2008).
Veldman-Jones, M. H. et al. Reproducible, quantitative, and flexible molecular subtyping of clinical DLBCL samples using the NanoString nCounter system. Clin. Cancer Res. 21, 2367–2378 (2015).
Gjodsbol, K. et al. Multiple bacterial species reside in chronic wounds: a longitudinal study. Int. Wound J. 3, 225–231 (2006).
Watnick, P. & Kolter, R. Biofilm, city of microbes. J. Bacteriol. 182, 2675–2679 (2000).
Rigatto, M. H., Ribeiro, V. B., Konzen, D. & Zavascki, A. P. Comparison of polymyxin B with other antimicrobials in the treatment of ventilator-associated pneumonia and tracheobronchitis caused by Pseudomonas aeruginosa or Acinetobacter baumannii. Infection 41, 321–328 (2013).
Govan, J. R. & Deretic, V. Microbial pathogenesis in cystic fibrosis: mucoid Pseudomonas aeruginosa and Burkholderia cepacia. Microbiol. Rev. 60, 539–574 (1996).
Falagas, M. E. & Bliziotis, I. A. Pandrug-resistant Gram-negative bacteria: the dawn of the post-antibiotic era? Int. J. Antimicrob. Agents 29, 630–636 (2007).
Hoffman, L. R. et al. Selection for Staphylococcus aureus small-colony variants due to growth in the presence of Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 103, 19890–19895 (2006).
Rahme, L. G. et al. Common virulence factors for bacterial pathogenicity in plants and animals. Science 268, 1899–1902 (1995).
Geiss, G. K. et al. Direct multiplexed measurement of gene expression with color-coded probe pairs. Nat. Biotechnol. 26, 317–325 (2008).
Mougous, J. D. et al. A virulence locus of Pseudomonas aeruginosa encodes a protein secretion apparatus. Science 312, 1526–1530 (2006).
Hood, R. D. et al. A type VI secretion system of Pseudomonas aeruginosa targets, a toxin to bacteria. CellHost. Microbe 7, 25–37 (2010).
Sana, T. G., Berni, B. & Bleves, S. The T6SSs of Pseudomonas aeruginosa strain PAO1 and their effectors:beyond bacterial-cell targeting. Front. Cell Infect. Microbiol. 6, 61 (2016).
Colvin, K. M. et al. The pel polysaccharide can serve a structural and protective role in the biofilm matrix of Pseudomonas aeruginosa. PLoS. Pathog. 7, e1001264 (2011).
Colvin, K. M. et al. The Pel and Psl polysaccharides provide Pseudomonas aeruginosa structural redundancy within the biofilm matrix. Environ. Microbiol. 14, 1913–1928 (2012).
Jennings, L. K. et al. Pel is a cationic exopolysaccharide that cross-links extracellular DNA in the Pseudomonas aeruginosa biofilm matrix. Proc. Natl. Acad. Sci. USA 112, 11353–11358 (2015).
Klausen, M. et al. Biofilm formation by Pseudomonas aeruginosa wild type, flagella and type IV pili mutants. Mol. Microbiol. 48, 1511–1524 (2003).
Periasamy, S. et al. Pseudomonas aeruginosa PAO1 exopolysaccharides are important for mixed species biofilm community development and stress tolerance. Front. Microbiol. 6, 851 (2015).
de Kievit, T. R. Quorum sensing in Pseudomonas aeruginosa biofilms. Environ. Microbiol. 11, 279–288 (2009).
Hentzer, M. et al. Inhibition of quorum sensing in Pseudomonas aeruginosa biofilm bacteria by a halogenated furanone compound. Microbiologym 148, 87–102 (2002).
Hentzer, M. et al. Attenuation of Pseudomonas aeruginosa virulence by quorum sensing inhibitors. EMBO J. 22, 3803–3815 (2003).
O’Loughlin, C. T. et al. A quorum-sensing inhibitor blocks Pseudomonas aeruginosa virulence and biofilm formation. Proc. Natl. Acad. Sci. USA 110, 17981–17986 (2013).
Christensen, L. D. et al. Synergistic antibacterial efficacy of early combination treatment with tobramycin and quorum-sensing inhibitors against Pseudomonas aeruginosa in an intraperitoneal foreign-body infection mouse model. J. Antimicrob. Chemother. 67, 1198–1206 (2012).
Hentzer, M., Eberl, L., Nielsen, J. & Givskov, M. Quorum sensing - a novel target for the treatment of biofilm infections. BioDrugs 17, 241–250 (2003).
Cowell, B. A., Twining, S. S., Hobden, J. A., Kwong, M. S. & Fleiszig, S. M. Mutation of lasA and lasB reduces Pseudomonas aeruginosa invasion of epithelial cells. Microbiology 149, 2291–2299 (2003).
Duan, K., Dammel, C., Stein, J., Rabin, H. & Surette, M. G. Modulation of Pseudomonas aeruginosa gene expression by host microflora through interspecies communication. Mol. Microbiol. 50, 1477–1491 (2003).
Biswas, L., Biswas, R., Schlag, M., Bertram, R. & Gotz, F. Small-colony variant selection as a survival strategy for Staphylococcus aureus in the presence of Pseudomonas aeruginosa. Appl. Environ. Microbiol. 75, 6910–6912 (2009).
Whiley, R. A. et al. Differential potentiation of the virulence of the Pseudomonas aeruginosa cystic fibrosis liverpool epidemic strain by oral commensal Streptococci. J. Infect. Dis. 209, 769–780 (2014).
Korgaonkar, A., Trivedi, U., Rumbaugh, K. P. & Whiteley, M. Community surveillance enhances Pseudomonas aeruginosa virulence during polymicrobial infection. Proc. Natl. Acad. Sci. USA 110, 1059–1064 (2013).
Lesic, B., Starkey, M., He, J., Hazan, R. & Rahme, L. G. Quorum sensing differentially regulates Pseudomonas aeruginosa type VI secretion locus I and homologous loci II and III, which are required for pathogenesis. Microbiology 155, 2845–2855 (2009).
Mah, T. F. et al. A genetic basis for Pseudomonas aeruginosa biofilm antibiotic resistance. Nature 426, 306–310 (2003).
Rogers, G. B., van der Gast, C. J. & Serisier, D. J. Predominant pathogen competition and core microbiota divergence in chronic airway infection. ISME J. 9, 217–225 (2015).
Flanagan, J. L. et al. Loss of bacterial diversity during antibiotic treatment of intubated patients colonized with Pseudomonas aeruginosa. J. Clin. Microbiol. 45, 1954–1962 (2007).
D’Argenio, D. A., Calfee, M. W., Rainey, P. B. & Pesci, E. C. Autolysis and autoaggregation in Pseudomonas aeruginosa colony morphology mutants. J. Bacteriol. 184, 6481–6489 (2002).
Allsopp, L. P. et al. RsmA and AmrZ orchestrate the assembly of all three type VI secretion systems in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 114, 7707–7712 (2017).
Armbruster, C. R. et al. Staphylococcus aureus protein A mediates interspecies interactions at the cell surface of Pseudomonas aeruginosa. mBio 7, e00538–16 (2016).
Wood, L. F. & Ohman, D. E. Use of cell wall stress to characterize sigma(22) (AlgT/U) activation by regulated proteolysis and its regulon in Pseudomonas aeruginosa. Mol. Microbiol. 72, 183–201 (2009).
Lee, K. W. et al. Biofilm development and enhanced stress resistance of a model, mixed-species community biofilm. ISME J. 8, 894–907 (2014).
Pamp, S. J., Gjermansen, M., Johansen, H. K. & Tolker-Nielsen, T. Tolerance to the antimicrobial peptide colistin in Pseudomonas aeruginosa biofilms is linked to metabolically active cells, and depends on the pmr and mexAB-oprM genes. Mol. Microbiol. 68, 223–240 (2008).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Gentleman, R. C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004).
Huson, D. H. et al. MEGAN Community Edition–Interactive exploration and analysis of large-scale microbiome sequencing data. PLoS. Comput. Biol. 12, e1004957 (2016).
Lambertsen, L., Sternberg, C. & Molin, S. Mini-Tn7 transposons for site-specific tagging of bacteria with fluorescent proteins. Environ. Microbiol. 6, 726–732 (2004).
Lee, J. et al. A cell-cell communication signal integrates quorum sensing and stress response. Nat. Chem. Biol. 9, 339–343 (2013).
Olsen, R. H., D., G. & McCombie, W. R. Development of broad-host-range vectors and gene banks: self-cloning of the Pseudomonas aeruginosa PAO chromosome. J. Bacteriol. 150, 60–69 (1982).
Cheng, Y. et al. Population dynamics and transcriptomic responses of Pseudomonas aeruginosa in a complex laboratory microbial community. bioRxiv, https://doi.org/10.1101/351668, (2018).
Acknowledgements
This research was supported by the National Research Foundation and Ministry of Education Singapore under its Research Centre of Excellence Program and AcRF Tier 2 (MOE2016-T2-1-010) from Ministry of Education, Singapore. We appreciate Professor Zhang Lian-Hui to provide us the pK18-Gm-mobsacB plasmid. We thank Dr. Daniela Moses, Mr. Yap Zhei Hwee, Rikky Wenang Purbojati and Mr. Muhammad Hafiz Bin Ismail for help with the RNA-seq experiments. We also appreciate Dr. Sharon Longford for her help on language polishing.
Author information
Authors and Affiliations
Contributions
Y.C. designed and carried out experiments, interpreted results and wrote the manuscript. L.Y. designed the project and revised the manuscript. J.K.H.Y. helped with the confocal microscopy imaging and biofilm bioreactors. Z.C. performed metatranscriptomic analysis. Y.D., L.Z. and Y.D. helped to revise the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cheng, Y., Yam, J.K.H., Cai, Z. et al. Population dynamics and transcriptomic responses of Pseudomonas aeruginosa in a complex laboratory microbial community. npj Biofilms Microbiomes 5, 1 (2019). https://doi.org/10.1038/s41522-018-0076-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41522-018-0076-z
This article is cited by
-
Probiotic supplementation during antibiotic treatment is unjustified in maintaining the gut microbiome diversity: a systematic review and meta-analysis
BMC Medicine (2023)
-
The impact of mass drug administration of antibiotics on the gut microbiota of target populations
Infectious Diseases of Poverty (2022)
-
Shades of grey: host phenotype dependent effect of urbanization on the bacterial microbiome of a wild mammal
Animal Microbiome (2021)
-
Metaproteomics reveals insights into microbial structure, interactions, and dynamic regulation in defined communities as they respond to environmental disturbance
BMC Microbiology (2021)
-
An integrated model system to gain mechanistic insights into biofilm-associated antimicrobial resistance in Pseudomonas aeruginosa MPAO1
npj Biofilms and Microbiomes (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.