Abstract
Biofilm formation is an important problem for many industries. Desulfovibrio vulgaris is the representative sulfate-reducing bacterium (SRB) which causes metal corrosion in oil wells and drilling equipment, and the corrosion is related to its biofilm formation. Biofilms are extremely difficult to remove since the cells are cemented in a polymer matrix. In an effort to eliminate SRB biofilms, we examined the ability of supernatants from Pseudomonas aeruginosa PA14 to disperse SRB biofilms. We found that the P. aeruginosa supernatants dispersed more than 98% of the biofilm. To determine the biochemical basis of this SRB biofilm dispersal, we examined a series of P. aeruginosa mutants and found that mutants rhlA, rhlB, rhlI, and rhlR, defective in rhamnolipids production, had significantly reduced levels of SRB biofilm dispersal. Corroborating these results, purified rhamnolipids dispersed SRB biofilms, and rhamnolipids were detected in the P. aeruginosa supernatants. Hence, P. aeruginosa supernatants disperse SRB biofilms via rhamnolipids. To determine the genetic basis of how the P. aeruginosa supernatants disperse SRB biofilms, a whole transcriptomic analysis was conducted (RNA-seq); based on this analysis, we identified four proteins (DVUA0018, DVUA0034, DVUA0066, and DVUA0084) of the D. vulgaris megaplasmid that influence biofilm formation, with production of DVUA0066 (a putative phospholipase) reducing biofilm formation 5.6-fold. In addition, the supernatants of P. aeruginosa dispersed the SRB biofilms more readily than protease in M9 glucose minimum medium and were also effective against biofilms of Escherichia coli and Staphylococcus aureus.
Similar content being viewed by others
Introduction
Sulfate-reducing bacteria (SRB) are an important type of microorganism causing iron corrosion on metal surfaces under both anaerobic and aerobic conditions.1,2,3 Desulfovibrio vulgaris Hildenborough is a sequenced4 Gram-negative SRB that has been used as an SRB model organism to study biocorrosion and bioremediation of toxic metal ions4 as well as biofilm formation5,6 and bioimmobilization at superfund sites.7 It is also called the “petroleum pest” because it is commonly found in oil fields and causes “souring” of petroleum and damage to topside equipment and pipelines.8
Biofilms are groups of bacteria that are held together in a self-produced extracellular matrix9 and are difficult to remove with antimicrobial agents due to their antibiotic or biocide resistance relative to planktonic cells.10 Desulfovibrio sp. populations in biofilms have a significant role for microbial induced corrosion because of their sulfide production and electron transfer mechanism,5 and biofilms of D. vulgaris have been extensively shown to cause corrosion in many types of steels and other alloys.11 The biofilms of D. vulgaris consists primarily of protein,5 mannose,6 fucose,6 and N-acetylgalactosamine.6 The biofilms of Escherichia coli consist primarily of proteinaceous curli fibres, flagella, and the polysaccharide cellulose,12 and the biofilms of Staphylococcus aureus are largely composed of cytoplasmic proteins13 and extracellular genomic DNA.14
Many Gram-negative bacteria use quorum sensing (QS) molecules or autoinducers to communicate with each other15 and to form biofilms.15 The QS mechanism can control particular processes related to cell density,16 and QS inhibition targeting autoinducers has been used as a method to control biofilm formation.16 The opportunistic pathogen Pseudomonas aeruginosa has four QS systems (Las, Rhl, Pqs, and Iqs).17 Each QS system has its own signal and regulatory protein. For the Las system, LasI synthesizes N-(3-oxododecanoyl)-homoserine lactone (3oxoC12HSL), and LasR is the protein receptor.17 For the Rhl system, RhlI synthesizes N-butyrylhomoserine lactone (C4-HSL), and RhlR is the protein receptor. The third QS system is quinolone-based intercellular signaling; the PQS signal is 2-heptyl-3-hydroxy-4-quinolone, and it is synthesized by the products of the PQS synthesis cluster consisting of pqsABCD, phnAB, and pqsH. PqsR is the protein receptor.17 The fourth system is called Iqs, and the QS molecule is 2-(2-hydroxyphenyl)-thiazole-4-carbaldehyde.17
One aspect of biofilm formation controlled by the Rhl QS system is regulation of the synthesis of rhamnolipids, which are glycolipid biosurfactants composed of rhamnose and 3-(hydroxyalkanoyloxy) alkanoic acid (HAA).18 P. aeruginosa rhamnolipids affect biofilm architecture by participating in the maintenance of biofilm channels19 and by reducing adhesion between cells20; hence, they have been used to disperse biofilms of Bordetella bronchiseptica,21 Bacillus pumilus,22 Staphylococcus aureus,23 Listeria monocytogenes20 and Salmonella enteritidis.20 In addition, rhamnolipids from Lysinibacillus sp. BV152.1 also inhibit biofilm formation of P. aeruginosa PAO1, S. aureus, and Serratia marcescens.24
There are many reasons for forming bacterial biofilms such as a defense against stress (e.g., nutrient deprivation, antibiotics, or pH changes)25 and a mechanism for staying stationary in a nutrient-rich area.25 However, bacteria also must have a means to leave the biofilm (dispersal) due to environmental changes (e.g., fluctuations in oxygen levels, alterations in nutrients, or increasing of toxic products)26; this process involves breaking the matrix to uncement the cells.27
Here, we show that supernatants from P. aeruginosa PA14 (henceforth PA14) disperse the biofilms of SRB, E. coli, and S. aureus. Using QS-related mutants, we determined that the biochemical basis for this dispersal is the presence of rhamnolipids in the supernatants. We also investigated the genetic basis of dispersal of D. vulgaris biofilms via PA14 supernatants by RNA sequencing (RNA-seq) and found that DVUA0018, DVUA0034, DVUA0066, and DVUA0084 encoded by the D. vulgaris megaplasmid are related to biofilm formation.
Results
PA14 wild-type supernatant disperses SRB biofilm
In an effort to investigate whether there are QS compounds utilized by the representative SRB D. vulgaris, we tested whether its supernatants would disperse its own biofilm formed in rich medium. D. vulgaris supernatant did not disperse its own biofilm within 2 h (data not shown). Since there was no negative effect of SRB supernatants on its own biofilm, we investigated the effect of the supernatant of other species (e.g., P. aeruginosa, P. fluorescens, Bacillus subtilis, E. coli). Among these species, the supernatants (concentrated 4×) of P. aeruginosa PA14 and P. aeruginosa PAO1 dispersed SRB biofilm the most (Fig. 1a). The supernatants were obtained from planktonic stationary-phase cultures, and the SRB biofilm grown was in 96-well plates for 24–48 h in modified Baar’s medium. Critically, the PA14 wild-type supernatant dispersed D. vulgaris biofilm more than 92% after 1–2 h of incubation. We utilized short periods of contact of the supernatants with the SRB biofilm to avoid artifacts related with growth of the bacterium.
D. vulgaris biofilm dispersal by supernatants of P. aeruginosa PA14, its quorum sensing mutants, P. aeruginosa PAO1, B. subtilis, P. fluorescens, E. coli TG1, and homoserine lactones. (a) D. vulgaris biofilms were grown for 2 days in modified Baar’s media at 30 °C, and all of the supernatants were concentrated to 4× and contacted with D. vulgaris biofilms for 2 h. * indicates no dispersal. At least two independent cultures were used with three replicates for each culture, and the error bars indicate one standard deviation. (b) D. vulgaris biofilms were grown for 2 days in modified Baar’s medium at 30 °C, and supernatants were concentrated to 4× and contacted with D. vulgaris biofilms for 2 h. Rhamnolipid standards were added at 10 mM. Protease 1 (Savinase) at 0.024 U was used as a positive control. At least two independent cultures were used with three replicates for each culture, and the error bars indicate one standard deviation. (c) D. vulgaris was grown for 24 h to form biofilms in modified Baar’s medium. PA14 wild-type supernatants were used at 1× (so that the effect of the homoserine lactones could be more clearly discerned) with 50 µM of C4-HSL, 3oxoC12HSL, or DMF (negative control) added for 90 min. Savinase (protease) at 0.024 U was also used as a positive control with PBS buffer used as negative control for protease. Two independent cultures and three replicate wells for each condition were used. The error bar indicates one standard deviation. The symbols ** (P < 0.01) indicate significant differences versus the PA14 supernatants + DMF control via one-way ANOVA
Rhamnolipids in the supernatants disperse SRB biofilms
To determine mechanisms behind these strong dispersal results, we hypothesized that the compounds in the P. aeruginosa supernatant may be related to QS since PA14 is the best known strain for QS, and QS controls extracellular compounds like protease28 and rhamnolipids.28 Hence, we tested the supernatant of mutants related to QS as well as those related to virulence factors (e.g., lasI,29 lasB,30 pelA,31 phzM,32 phzS,32 pvdF,33 pchE,34 lecA,35 toxA,36 sodM,37 rhlI,38 and rhlR38). Similar to the supernatant of the PA14 wild-type, the supernatants of the mutants lasI (autoinducer synthase), lasB (elastase), pelA (oligogalacturonide lyase), phzM (pyocyanin biosynthesis), phzS (pyocyanin biosynthesis), pvdF (pyoverdin biosynthesis), pchE (pyochelin biosynthesis), lecA (LecA lectin), toxA (exotoxinA), and sodM (superoxide dismutase) dispersed the D. vulgaris biofilm like the supernatant of PA14 wild-type (Fig. 1b), indicating the Las QS system, Pel polysaccharide, pyocyanin, pyoverdine, pyocheline, LecA lectin, exotoxin A, superoxide dismutase had no role in the biofilm dispersal. To corroborate these QS results, we added both C4-HSL and 3oxoC12HSL to the wild-type supernatants without concentration; i.e., 1×, so the effect of the homoserine lactones could be more clearly discerned, but there was no additional dispersal of the SRB biofilm (Fig. 1c).
In contrast to the other mutants, mutations affecting the regulation and biosynthesis of rhamnolipids, specifically in the gene encoding the autoinducer synthase, rhlI, and the transcriptional regulator, rhlR, had a pronounced decrease in dispersal (Fig. 1b). Since the rhlI and rhlR mutants were found to affect biofilm dispersal, additional mutations in the rhamnolipid pathway were investigated: rhlA, which encodes rhamnosyltransferse 1 subunit A and rhlB, which encodes the catalytic subunit of the rhamnosyltransferase.38,39 The supernatants of both the rhlA and rhlB mutant did not disperse SRB biofilms (Fig. 1b). Therefore, the dispersal compounds are related to rhamnolipids.
To corroborate the 96-well biofilm dispersal results with the PA14 wild-type and rhlA mutant, confocal microscopy was used to visualize the remaining biofilm after treatment with supernatants from these two strains. As shown in Fig. 2, the supernatant from the wild-type strain nearly completely dispersed the SRB biofilm whereas the supernatant from the rhlA mutant had no effect, just like the buffer negative control.
Representative images of D. vulgaris biofilm dispersal via P. aeruginosa supernatants as visualized by confocal microscopy. D. vulgaris biofilms were formed in an 8 chamber cell culture slide for 48 h in modified Baar’s medium. Supernatants were concentrated to 4× and contacted with D. vulgaris biofilms for 20 min: supernatants of (a) P. aeruginosa PA14, (b) P. aeruginosa PA14 rhlA, and (c) PBS buffer (used as a negative control to indicate what undispersed biofilms after the removal planktonic cells and loose biofilm). Scale bar is 50 µm
From the rhamnolipid production pathway (Fig. 3), RhlA synthesizes 3-(3-hydroxyakanoyloxy) alkanoic acids (HAA),40and RhlB uses HAA to make mono-rhamnolipids.41 Therefore, the compounds required for SRB biofilm dispersal are HAA or mono/di-rhamnolipids, respectively. However, the results from mass spectrometry showed that there was no HAA detected in the wild-type supernatant sample. Therefore we conclude that rhamnolipids are the compounds (and not HAA) that are important for biofilm dispersal.
Simplified rhamnolipid biosynthesis pathway in P. aeruginosa.56,57 RhlR binds with N-butanoyl-L-homoserine lactone (C4-HSL) produced by RhlI to form a RhlR–C4-HSL complex. The RhlR–C4-HSL complex interacts with the rhlA promoter to initiate transcription of the rhlAB genes to produce rhamnolipids. HAA is 3-(hydroxyalkanoyloxy) alkanoic acid. P. aeruginosa commonly produces rhamnolipids containing fatty acids with chain lengths between C8 and C14 (n = 1–7).58
Rhamnolipids in the PA14 supernatants disperse SRB biofilms
The genetic results clearly indicate the importance of rhamnolipids for SRB biofilm dispersal via PA14 supernatants. To corroborate these results, the 4× concentrated supernatants of PA14, rhlA, rhlB, rhlI, and rhlR mutants were tested for the presence of rhamnolipids by mass spectrometry (Supplementary Fig. 1). As expected, rhamnolipids were detected only in the PA14 supernatant samples and not from any of the mutant samples. There were four predominant peaks of rhamnolipids shown (Supplementary Fig. 1) which are Rha-C10–C10, Rha-C10–C12/Rha-C12–C10, Rha-Rha-C10–C10, and Rha-Rha-C10–C12/ Rha-Rha-C12–C10. By comparing with the 10 mM rhamnolipids standard from P. aeruginosa, we determined the concentration of rhamnolipids in the PA14 supernatant using the total peak areas is over 100 fold less than the standard (0.1 mM, Supplementary Table 1).
To show conclusively that rhamnolipids in the supernatants are the biochemical means by which the D. vulgaris biofilms are dispersed, commercial, purified P. aeruginosa rhamnolipids (10 mM) were tested. We found that the commercial rhamnolipids disperse D. vulgaris biofilm with approximately 67% dispersal after 2 h (Fig. 1b). The commercial rhamnolipids standard did not disperse the biofilm as well as the PA14 supernatant (90%, Fig. 1b) though there were 100× more rhamnolipids in the 10 mM commercial standard sample than the amount of rhamnolipids in the supernatant (Supplementary Table 1). This implies that the ratio of the rhamnolipids or other compounds in the supernatant are important for SRB biofilm dispersal; the commercial rhamnolipids may also be altered upon purification. We tested whether other components in the supernatant were required by adding the supernatant of the rhlB mutant to the rhamnolipids commercial standard, but there was no increase in biofilm dispersal compared to the rhamnolipids standard alone. This suggests the dispersal is due to the rhamnolipids alone and not some other product in the supernatants.
We also tested whether PA14 supernatants decrease the viability of SRB planktonic cells and found their viability is reduced by 60 ± 8% with wild-type supernatants (5×) and 37 ± 2% by rhlA supernatants (5×, that lack rhamnolipids). The purified commercial rhamnolipids (10 mM) reduced planktonic SRB viability by 17.0 ± 0.6%. These results corroborate the previous report that rhamnolipids from P. aeruginosa PA14 have some toxicity42 and indicate there are other compounds with toxicity in the supernatants. Critically, we tested the effect of PA14 supernatants (concentrated 4×, to match the earlier dispersal conditions) on SRB biofilm cells (which mirrors the conditions used in this manuscript to disperse biofilm cells) and found significantly less toxicity for both wild-type (23 ± 3%) and rhlA supernatants (22.4 ± 0.5%); the purified commercial rhamnolipids reduced viability by 9 ± 2%.
PA14 supernatants disperse D. desulfuricans, E. coli MG1655, and Staphylococcus aureus biofilms
To test the ability of the PA14 supernatants to disperse other bacteria, biofilms of D. desulfuricans, E. coli MG1655, S. aureus, and PA14 were grown in 96-well plates, then treated with supernatants from PA14. The PA14 supernatant dispersed the biofilms of D. desulfuricans, E. coli MG1655, and S. aureus well (by over 90%). Although, there are some species-specific differences, these results show that the PA14 supernatants have a great deal of utility for dispersing a wide range of biofilms.
The supernatant from the rhlR mutant did not disperse the D. desulfuricans and E. coli biofilms well (Fig. 4), providing further evidence of the importance of rhamnolipids for these species. However, there is a rhamnolipid-independent component responsible for dispersing S. aureus biofilms. In addition, the supernatant of PA14 did not disperse its own biofilm (data not shown).
Supernatants of P. aeruginosa PA14 disperse diverse biofilms, and D. vulgaris biofilm formation during production of putative dispersal proteins identified through RNA-seq. The biofilms of D. desulfuricans (a) and E. coli MG1655 (b) were grown for 24 h in modified Baar’s medium and LB, respectively, and the biofilms of S. aureus (c) were grown for 24 h in TSB.59 P. aeruginosa wild-type and rhlR mutant supernatants were concentrated to 4× and contacted with the biofilms for 2 h except for the S. aureus biofilm where 1x supernatant was used instead of 4× and the contact time was 10 min (since S. aureus biofilms were dispersed quickly). At least two independent cultures were used with three replicates for each culture, and the error bars indicate one standard deviation. (d) Biofilms of D. vulgaris were formed for 48 h in modified Baar’s medium at 30 °C (anaerobically) with expression of dvu3313, dvua0018, dvua0034, dvua0066, and dvua0084 identified from the dispersal RNA-seq experiment. Vector pVLT33 was used with the biofilm-specific promoter Pdvu0304 fused to each gene. Normalized biofilm formation percentage (based on OD540nm/OD620nm) is shown relative to the empty plasmid control. Two independent cultures were used with three replicates for each culture, and the error bars indicate one standard deviation. ** P < 0.01 indicates significant differences versus the empty plasmid control group via one-way ANOVA
PA14 supernatant disperses SRB biofilm grown in M9G
Since biofilm matrices can change when the growth medium is altered,43 we tested the ability of the PA14 supernatant to disperse SRB biofilms grown in minimal glucose medium (M9G). The PA14 supernatant (4x concentrated) dispersed 90–100% of the SRB biofilm when formed in modified Baar’s medium and dispersed about 80% of the biofilm formed in M9G. In contrast, the positive control protease, which dispersed 80% of the SRB biofilm formed in modified Baar’s medium, dispersed only about 40% of the biofilm grown in M9G.
Genetic basis of PA14 supernatant dispersal
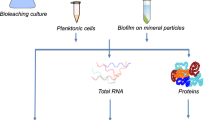
To determine if there was a genetic component to the dispersal of the SRB cells upon the addition of the P. aeruginosa supernatants, gene expression levels of dispersed D. vulgaris biofilm cells after contact with wild-type PA14 supernatants were compared to non-dispersed D. vulgaris cells contacted rhlA PA14 supernatants, via RNA-seq. Hence, the role of rhamnolipids in SRB biofilm dispersal was analyzed.
The genes that had significant differences in expression levels in dispersed cells are shown in Supplementary Table 2 (induced genes) and Supplementary Table 3 (repressed genes). The gene expression levels were quantified as transcript per kilobases million (TPM) for each gene. Based on the results in Supplementary Table 2 and Supplementary Table 3, we cloned one gene (dvua0018) that was induced and four genes (dvua0034, dvua0066, dvua0084, and dvu3313) that were repressed and tested whether production of the proteins impact SRB biofilm formation. These genes were chosen based on these criteria: (i) their high differential expression, (ii) their significant TPM values (which indicates they are actively expressed), and (iii) their position in the genome which indicates they are not in the same operon so we avoided polar affects. To investigate the importance of these SRB proteins in biofilm formation, we utilized a promoter that is induced solely in D. vulgaris biofilms (Pdvu0304, unpublished). We found DVUA0018, DVUA0034, DVUA0066, and DVUA0084 significantly affect biofilm formation, with production of DVUA0066 (a likely phospholipase) reducing biofilm formation 5.6-fold (Fig. 4d). These four proteins are found on the megaplasmid; hence, we identified specific proteins on this megaplasmid that influence biofilm formation.
Discussion
We have demonstrated that the supernatants of PA14 disperse D. vulgaris biofilms effectively and that the biochemical basis of this dispersal is the presence rhamnolipids. The PA14 supernatants dispersed the biofilm better than protease, and they are versatile because they also dispersed D. desulfuricans, E. coli MG1655, and S. aureus biofilms. This is significant since the matrix of D. desulfuricans, E. coli, and S. aureus are primarily proteins,12,13 carbohydrates,44 and some extracellular genomic DNA,14 respectively, which indicates that rhamnolipids are a powerful and general approach for removing biofilms. Though the supernatant from the rhlR mutant did not disperse biofilms of S. aureus as well as the PA14 wild-type supernatants, it dispersed about 70% of the biofilm. Hence, there may be other compounds in the PA14 supernatants beyond rhamnolipids that disperse S. aureus biofilms.
The rhamnolipid standard did not disperse the biofilm as well as the rhamnolipids in the PA14 WT supernatant, possibly because the ratios of the rhamnolipids are critical for biofilm dispersal activity. For example, the ratio between di-rhamnolipids and mono-rhamnolipids affects the mixture properties (e.g., foam thickness, surface electric parameters)45 and changes the emulsification index and antimicrobial properties.46 In addition, the rhamnolipid standard was not pure (90%), and the growth condition used to produce the rhamnolipids is unknown. We also surmise that purifying the rhamnolipids alters their biofilm dispersal properties.
Previously, it was reported that rhamnolipids from P. aeruginosa together with the QS signal 3oxoC12HSL work cooperatively to disperse E. coli biofilms47; however, we found that there was no need for 3oxoC12HSL for supernatants to disperse SRB since the supernatants from the lasI mutant were just as effective as those from the wild-type strain (Fig. 1b). Furthermore, adding either C4-HSL or 3oxoC12HSL had no effect on SRB biofilm dispersal by the wild-type PA14 supernatants (Fig. 1c). Therefore, dispersal of the D. vulgaris biofilm by rhamnolipids is not due to these QS signals from P. aeruginosa.
Based on the whole-transcriptome results (RNA-seq) comparing the expression levels of SRB genes from cells dispersed from biofilms (contacted with PA14 supernatants) vs. their expression in non-dispersed biofilm cells (contacted with PA14 rhlA supernatants that lack rhamnolipids), four proteins (DVUA0018, DVUA0034, DVUA0066, and DVUA0084) were found to impact SRB biofilm formation. Critically, all of them are encoded by the megaplasmid of D vulgaris and production of DVUA0066 reduced biofilm formation by over 5-fold. Hence, by studying the genetic basis of dispersal with rhamnolipids, we identified specific proteins of the megaplasmid involved in biofilm formation. Previously, the megaplasmid (but not specific proteins) has been shown to increase biofilm formation 300%.5 In summary, relatively inexpensive supernatants containing rhamnolipids may be used to disperse industrially-relevant biofilms as well as to provide insights into the genetic basis of biofilm formation.
Methods
Bacterial species and culture conditions
Desulfovibrio vulgaris and Desulfovibrio desulfuricans were grown anaerobically in modified Baar’s medium at 30 °C or minimal medium with 0.4% glucose (M9G)48. PA14 wild-type and its mutants were grown in lysogeny broth (LB)49 or M9G at 37 °C. Gentamicin (Gm) 15 μg/mL was used for culturing the PA14 mutants. Escherichia coli TG1, Escherichia coli MG1655, Bacillus subtilis, and Pseudomonas fluorescens were grown in LB at 37 °C. For the recombinant strains to maintain broad host vector plasmid pVLT33, 50 ng/μL kanamycin was added to LB for E. coli and 400 ng/μL G418 was added to modified Baar’s medium (ATCC medium no. 1249) with 0.025% sodium sulfide (as an oxygen scavenger) for D. vulgaris. All the species are shown in Supplementary Table 4. The PA14 rhlA, rhlB, rhlI, rhlR, and lasI mutants50,51 were verified via PCR using the primers in Supplementary Table 5.
Cloning biofilm dispersal-related genes identified by RNA-seq
To investigate whether proteins encoded by the differentially-expressed genes in the RNA-seq data for SRB treated with P. aeruginosa supernatant samples could affect biofilm formation, dvua0018, dvu3313, dvua0084, dvua0034 and dvua0066 were cloned into broad-host range plasmid pVLT3352 under control of the biofilm-specific promoter Pdvu0304 with ribosome binding site AAGGAG. The primers for construction pVLT33 derivatives are listed in Supplementary Table 5. E. coli TG1 was used for the constructions of all the plasmids. Correct construction of pVLT33 derivatives were confirmed by sequencing with primers pVLT33-SF and pVLT33-SB, which locate at the upstream and downstream of multiple cloning site on plasmid pVLT33 respectively.
To electroporate the pVLT33 derivatives into wild-type D. vulgaris, anaerobically, competent cells (turbidity ~0.3 at 600 nm) were prepared by washing twice with pre-chilled, sterile 10% glycerol. In the anaerobic chamber, plasmid DNA (0.5–1 µg) was added to competent cells (50 µL) by mixing gently, and the solution was transferred to a pre-chilled (0 °C), 1 mm electroporation cuvette (Fisherbrand cat #FB101, Fisher Scientific, USA). Electroporation was performed aerobically (conditions: 25 µF, 200 Ω, and 1.5 kV/cm). The cuvette was returned to the anaerobic chamber, and 1 mL of modified Baar’s medium was added immediately. The cells were mixed gently and transferred to a 1.5 mL Eppendorf tube where they were recovered overnight at 30 °C anaerobically. After the recovery, 50 µL was inoculated into 10 mL modified Baar’s medium (0.2% yeast extract) with G418 (400 µg/mL) or plated on modified Baar’s medium (0.2% yeast extract) 1% agar plate with G418 (400 µg/mL). Genomic DNA from 1 to 2 mL of culture or the colony was isolated using an UltraClean@ Microbial DNA isolation kit (MO BIO cat#12224, Qiagen, USA) for PCR verification. During the PCR verification of the transformation of the cloned genes dvua0018, dvu3313, dvua0084, dvua0034, and dvua0066 into D. vulgaris, primers pVLT33-SB and dvua0018-BamHI-R2, pVLT33-SB and dvu3313-BamHI-R2, pVLT33-SB and dvua0084-BamHI-R2, pVLT33-SB and dvua0034-R, and pVLT33-SB and dvua0066-R were used, respectively (Supplementary Table 5).
PA14 supernatant preparation
P. aeruginosa PA14 wild-type and its isogenic mutants were grown in LB or LB with Gm15 μg/mL for the mutants. The overnight cultures (200 µL) were centrifuged, the cell pellets were resuspended in 50 mL of M9G, and cultured for 4–5 days. The supernatants were collected and filtered using a 0.22 µm filtration system. The samples were concentrated 4× using a speed vacuum.
Biofilm dispersal assay using crystal violet
Biofilm formation was assayed in 96-well polystyrene plates using 0.1% crystal violet staining as described previously53 with some modifications. Diluted overnight cultures (150 µL) at an initial turbidity at 600 nm of 0.05 were inoculated into 96-well-plates with medium, and the bacteria were cultured for 24–48 h at 30 °C without shaking. The planktonic cells were removed from each well, and the biofilm was washed gently using phosphate buffered saline (PBS). The supernatant (150 µL, at the indicated concentrations), chemicals (including the positive control protease Savinase, 0.024 U), or PBS (as a control) were added to the wells, and the plate was incubated at 30 °C for 1–2 h. Then, the supernatants were removed from each well, and the remaining biofilm was washed gently using water. After the crystal violet was added to each well, the wells were rinsed and dried, and ethanol was added to dissolve the crystal violet. The total biofilm remaining in the samples were measured at 540 nm. N-butanoyl-L-homoserine lactone (C4-HSL, Cayman Chemical, Ann Arbor, MI, catalog number 10007898) or N-(3-oxododecanoyl)-homoserine lactone (3oxoC12HSL, Cayman Chemical, catalog number 10007895) were used at 50 µM in dimethylformamide (DMF); DMF was used as the solvent control. At least two independent cultures were used for each strain with three replicates for each culture.
Cell viability
Cell viability was determined by staining SYTO9 and propidium iodide of the LIVE/DEAD BacLight Bacterial Viability Kit (Thermo Fisher Scientific, Waltham, MA). For planktonic cell viability, a D. vulgaris culture (1 mL) was centrifuged, 0.9 mL removed, and the remaining 0.1 mL (with the resuspended cell pellet) was contacted with 0.1 mL of P. aeruginosa PA14 supernatants (10×) for a final 5× supernatant concentration or contacted with 10 mM commercial rhamnolipids. The samples were incubated at 30 °C for 2 h anaerobically. PBS treatment (2 h) was used as a negative control, and ethanol (70%) treatment for 30 min was used as the positive control for cell death. Prior to staining, cells were washed once with 0.85% NaCl and resuspended in 0.1 mL of 0.85% NaCl. For D. vulgaris biofilm cell viability, biofilm cells were formed for 48 h in 96 well plates in 150 μL, washed with 150 μL of PBS buffer (pH 7.4), and contacted with 150 μL of P. aeruginosa PA14 supernatants (4×) and 10 mM commercial rhamnolipids for 2 h anaerobically. Then the liquids were removed and 150 μL of PBS buffer was added. The cells were stained at room temperature for 15 min. The samples were observed under a fluorescence microscope (Zeiss Axioscope.A1). At least three different areas were observed, and two independent cultures were tested.
Confocal microscopy
Diluted overnight cultures (300 µL) at an initial turbidity at 600 nm of 0.1 were inoculated into an 8 chamber cell culture slide (Dot Scientific, Inc, Burton, MI) with modified Baar’s medium, and the bacteria were cultured for 48 h at 30 °C without shaking. The planktonic cells were removed from each well, and the biofilm was washed gently using PBS. Supernatants or PBS was added to the chambers, and the slide was incubated at 30 °C for 20 min. The supernatants or PBS were removed from the chamber. Biofilm cells were washed with 300 µL PBS, then stained with SYTO 9 (5 µL/mL) in the dark for 1 h. Confocal microscopy images were taken using a PlanApo60×/1.4 oil objective lens with Olympus Fluoview 1000 confocal laser microscope. The biofilm structure images were performed using IMARIS software (Bitplane, Zurich, Switzerland), and average biomass and thickness were obtained using COMSTAT image-processing software54. At least three different areas were observed, and two independent cultures were tested.
Mass spectrometric analysis of supernatants
Mass spectrometric analysis was performed on the concentrated supernatants by using a Waters Q-TOF Premier quadrupole/time-of-flight (TOF) mass spectrometer (Waters Corporation (Micromass Ltd.), Manchester, UK). Operation of the mass spectrometer was performed using MassLynx™ software Version 4.1 (http://www.waters.com). Samples were introduced into the mass spectrometer using a Waters 2695 high performance liquid chromatograph. The samples were analyzed using flow injection analysis (FIA), in which the sample is injected into the mobile phase flow and passes directly into the mass spectrometer, where the analytes are ionized and detected. The mobile phase used was 90% acetonitrile (LC-MS grade), and 10% aqueous 10 mM ammonium acetate. The flow rate was 0.15 mL/min. The nitrogen drying gas temperature was set to 300 °C at a flow of 7 L/min. The capillary voltage was 2.8 kV. The mass spectrometer was set to scan from 100 to 1000 m/z in negative ion mode, using electrospray ionization (ESI). The rhamnolipids standard from P. aeruginosa (90%) was purchased from Sigma-Aldrich (St. Louis, MO). Rhamnolipid peaks were identified as disclosed previously55.
Sample preparation for RNA-seq
D. vulgaris was grown anaerobically at 30 °C for 24 h in 400 mL of modified Baar’s medium with 10 g of glass wool to promote biofilm formation. The glass wool was rinsed once in PBS to remove planktonic cells. The glass wool with biofilm cells was incubated in 200 mL of PA14 wild-type or rhlA mutant supernatants (1×) for 30 min anaerobically. The glass wool from the sample with the PA14 wild-type supernatant was removed, and the dispersed biofilm cells in the supernatant were harvested by centrifugation at 4 °C. Since the biofilm cells in the glass wool from the sample with the rhlA mutant supernatant was not dispersed, cells were harvested from the glass wool for RNA-seq. The glass wool was gently washed twice in 200 ml of cold 0.85% NaCl buffer and sonicated for 2 min in 200 mL of cold 0.85% NaCl with 2 mL of RNA later (Thermo Fisher Scientific, Waltham, MA) by using an ultrasonic water bath. The non-dispersed biofilm cells were centrifuged at 8000g at 4 °C. The biofilm cell pellets were washed in 4 mL of cold RNA later and kept at −80°C for RNA isolation. RNA was extracted using the High Pure RNA isolation kit (Roche, Basel, Switzerland). The experiments were performed with two independent cultures. The RNA samples were sent to the Genome Sciences Facility at Pennsylvania State College of Medicine, Hershey, PA to evaluate RNA quality and to perform RNA-seq.
Data availability
The authors declare that data supporting the findings of this study are available within the paper, and the raw RNA sequencing reads have been submitted to the NCBI Sequence Read Archive (SRA) database (http://www.ncbi.nlm.nih.gov) under SRA accession no. SRP154476.
References
Zhang, W., Culley, D. E., Nie, L. & Scholten, J. C. Comparative transcriptome analysis of Desulfovibrio vulgaris grown in planktonic culture and mature biofilm on a steel surface. Appl. Microbiol. Biotechnol. 76, 447–457 (2007).
Pankhania, I. P., Spormann, A. M., Hamilton, W. A. & Thauer, R. K. Lactate conversion to acetate, CO2 and H2 in cell suspensions of Desulfovibrio vulgaris (Marburg)—indications for the involvement of an energy driven reaction. Arch. Microbiol. 150, 26–31 (1988).
Lee, W., Lewandowski, Z., Nielsen, P. H. & Hamilton, W. A. Role of sulfate-reducing bacteria in corrosion of mild-steel—a review. Biofouling 8, 165 (1995).
Heidelberg, J. F. et al. The genome sequence of the anaerobic, sulfate-reducing bacterium Desulfovibrio vulgaris Hildenborough. Nat. Biotechnol. 22, 554–559 (2004).
Clark, M. E., Edelmann, R. E., Duley, M. L., Wall, J. D. & Fields, M. W. Biofilm formation in Desulfovibrio vulgaris Hildenborough is dependent upon protein filaments. Environ. Microbiol. 9, 2844–2854 (2007).
Poosarla, V. G. et al. Dispersal and inhibitory roles of mannose, 2-deoxy-D-glucose and N-acetylgalactosaminidase on the biofilm of Desulfovibrio vulgaris. Environ. Microbiol. Rep. (2017).
Faybishenko, B. et al. In situ long-term reductive bioimmobilization of Cr(VI) in groundwater using hydrogen release compound. Environ. Sci. Technol. 42, 8478–8485 (2008).
Korte, H. L. et al. Independence of nitrate and nitrite inhibition of Desulfovibrio vulgaris Hildenborough and use of nitrite as a substrate for growth. Environ. Sci. Technol. 49, 924–931 (2015).
Lopez, D., Vlamakis, H. & Kolter, R. Biofilms. Cold Spring Harb. Perspect. Biol. 2, a000398 (2010).
Mah, T. F. & O’Toole, G. A. Mechanisms of biofilm resistance to antimicrobial agents. Trends Microbiol. 9, 34–39 (2001).
Street, C. N. & Gibbs, A. Eradication of the corrosion-causing bacterial strains Desulfovibrio vulgaris and Desulfovibrio desulfuricans in planktonic and biofilm form using photodisinfection. Corros. Sci. 52, 1447–1452 (2010).
Hobley, L., Harkins, C., MacPhee, C. E. & Stanley-Wall, N. R. Giving structure to the biofilm matrix: an overview of individual strategies and emerging common themes. Fems Microbiol. Rev. 39, 649–669 (2015).
Foulston, L., Elsholz, A. K. W., DeFrancesco, A. S. & Losick, R. The extracellular matrix of Staphylococcus aureus biofilms comprises cytoplasmic proteins that associate with the cell surface in response to decreasing pH. MBIO 5, 1–9 (2014).
Rice, K. C. et al. The cidA murein hydrolase regulator contributes to DNA release and biofilm development in Staphylococcus aureus. Proc. Natl Acad. Sci. USA 104, 8113–8118 (2007).
Dickschat, J. S. Quorum sensing and bacterial biofilms. Nat. Prod. Rep. 27, 343–369 (2010).
McLean, R. J. C., Pierson, L. S. & Fuqua, C. A simple screening protocol for the identification of quorum signal antagonists. J. Microbiol. Meth. 58, 351–360 (2004).
Lee, J. & Zhang, L. H. The hierarchy quorum sensing network in Pseudomonas aeruginosa. Protein Cell 6, 26–41 (2015).
Desai, J. D. & Banat, I. M. Microbial production of surfactants and their commercial potential. Microbiol. Mol. Biol. Rev. 61, 47 (1997).
Davey, M. E., Caiazza, N. C. & O’Toole, G. A. Rhamnolipid surfactant production affects biofilm architecture in Pseudomonas aeruginosa PAO1. J. Bacteriol. 185, 1027–1036 (2003).
Gomes, M. Z. D. & Nitschke, M. Evaluation of rhamnolipid and surfactin to reduce the adhesion and remove biofilms of individual and mixed cultures of food pathogenic bacteria. Food Control 25, 441–447 (2012).
Irie, Y., O’Toole G, A. & Yuk, M. H. Pseudomonas aeruginosa rhamnolipids disperse Bordetella bronchiseptica biofilms. FEMS Microbiol. Lett. 250, 237–243 (2005).
Dusane, D. H., Nancharaiah, Y. V., Zinjarde, S. S. & Venugopalan, V. P. Rhamnolipid mediated disruption ofmarine Bacillus pumilus biofilms. Colloids Surf. B Biointerfaces 81, 242–248 (2010).
Shiva, E., Carvalho, S. S., Aires, J. W. P. & Nitschke, M. Disruption of Staphylococcus aureus biofilms using rhamnolipid biosurfactants. J. Dairy Sci. 100, 7864–7873 (2017).
Aleksic, I. et al. Anti-biofilm properties of bacterial di-rhamnolipids and their semi-synthetic amide derivatives. Front. Microbiol. 8, 2454 (2017).
Jefferson, K. K. What drives bacteria to produce a biofilm? FEMS Microbiol. Lett. 236, 163–173 (2004).
Kostakioti, M., Hadjifrangiskou, M. & Hultgren, S. J. Bacterial biofilms: development, dispersal, andtherapeutic strategies in the dawn of the postantibiotic era. CSH Perspect. Med. 3, 1–23 (2013).
Kaplan, J. B. Biofilm dispersal: Mechanisms, clinical Implications, and potential therapeutic uses. J. Dent. Res. 89, 205–218 (2010).
Aybey, A. & Demirkan, E. Inhibition of quorum sensing-controlled virulence factors in Pseudomonas aeruginosa by human serum paraoxonase. J. Med. Microbiol. 65, 105–113 (2016).
Storey, D. G., Ujack, E. E., Rabin, H. R. & Mitchell, I. Pseudomonas aeruginosa lasR transcription correlates with the transcription of lasA, lasB, and toxA in chronic lung infections associated with cystic fibrosis. Infect. Immun. 66, 2521–2528 (1998).
Liu, P. V. The roles of various fractions of Pseudomonas aeruginosa in its pathogenesis. 3. Identity of the lethal toxins produced in vitro and in vivo. J. Infect. Dis. 116, 481–489 (1966).
Sakuragi, Y. & Kolter, R. Quorum-sensing regulation of the biofilm matrix genes (pel) of Pseudomonas aeruginosa. J. Bacteriol. 189, 5383–5386 (2007).
Lau, G. W., Hassett, D. J., Ran, H. & Kong, F. The role of pyocyanin in Pseudomonas aeruginosa infection. Trends Mol. Med 10, 599–606 (2004).
Meyer, J. M., Neely, A., Stintzi, A., Georges, C. & Holder, I. A. Pyoverdin is essential for virulence of Pseudomonas aeruginosa. Infect. Immun. 64, 518–523 (1996).
Takase, H., Nitanai, H., Hoshino, K. & Otani, T. Impact of siderophore production on Pseudomonas aeruginosa infections in immunosuppressed mice. Infect. Immun. 68, 1834–1839 (2000).
Winzer, K. et al. The Pseudomonas aeruginosa lectins PA-IL and PA-IIL are controlled by quorum sensing and by RpoS. J. Bacteriol. 182, 6401–6411 (2000).
Gambello, M. J., Kaye, S. & Iglewski, B. H. LasR of Pseudomonas aeruginosa is a transcriptional activator of the alkaline protease gene (Apr) and an enhancer of exotoxin-a expression. Infect. Immun. 61, 1180–1184 (1993).
Hassett, D. J. et al. Quorum sensing in Pseudomonas aeruginosa controls expression of catalase and superoxide dismutase genes and mediates biofilm susceptibility to hydrogen peroxide. Mol. Microbiol. 34, 1082–1093 (1999).
Ochsner, U. A. & Reiser, J. Autoinducer-mediated regulation of rhamnolipid biosurfactant synthesis in Pseudomonas aeruginosa. Proc. Natl Acad. Sci. USA 92, 6424–6428 (1995).
Deziel, E., Lepine, F., Milot, S. & Villemur, R. rhlA is required for the production of a novel biosurfactant promoting swarming motility in Pseudomonas aeruginosa: 3-(3-hydroxyalkanoyloxy)alkanoic acids (HAAs), the precursors of rhamnolipids. Microbiol.-SGM 149, 2005–2013 (2003).
Rehm, B. H., Mitsky, T. A. & Steinbuchel, A. Role of fatty acid de novo biosynthesis in polyhydroxyalkanoic acid (PHA) and rhamnolipid synthesis by pseudomonads: establishment of the transacylase (PhaG)-mediated pathway for PHA biosynthesis in Escherichia coli. Appl. Environ. Microbiol. 67, 3102–3109 (2001).
Abdel-Mawgoud, A. M., Hausmann, R., Lepine, F., Muller, M. M. & Deziel, E. Rhamnolipids: Detection, analysis, biosynthesis, genetic regulation, and bioengineering of production. Microbiol. Monogr. 20, 13–55 (2011).
Bernier, S. P. et al. Pseudomonas aeruginosa-derived rhamnolipids and other detergents modulate colony morphotype and motility in the Burkholderia cepacia complex. J. Bacteriol. 199, 1–15 (2017).
Flemming, H. C. & Wingender, J. The biofilm matrix. Nat. Rev. Microbiol. 8, 623–633 (2010).
Beech, I. B., Gaylarde, C. C., Smith, J. J. & Geesey, G. G. Extracellular polysaccharides from Desulfovibrio desulfuricans and Pseudomonas fluorescens in the presence of mild and stainless-steel. Appl. Microbiol Biotechnol. 35, 65–71 (1991).
Cohen, R. & Exerowa, D. Surface forces and properties of foam films from rhamnolipid biosurfactants. Adv. Colloid Interfac. 134 – 35, 24–34 (2007).
Das, P., Yang, X. P. & Ma, L. Y. Z. Analysis of biosurfactants from industrially viable Pseudomonas strain isolated from crude oil suggests how rhamnolipids congeners affect emulsification property and antimicrobial activity. Front. Microbiol 5, 1–8 (2014).
Bhattacharjee, A., Nusca, T. D. & Hochbaum, A. I. Rhamnolipids mediate an interspecies biofilm dispersal signaling pathway. ACS Chem. Biol. 11, 3068–3076 (2016).
Sambrook, J., Fritsch, E. F. & Maniatis, T. Molecular Cloning, A Laboratory Manual. (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, 1989). Edn. 2.
Sambrook, J. F. & Russell, D. W. Molecular cloning: a laboratory manual. (Cold Spring Harbor Laboratory Press, New York, 2001). Edn. 3rd.
Liberati, N. T. et al. An ordered, nonredundant library of Pseudomonas aeruginosa strain PA14 transposon insertion mutants. Proc. Natl Acad. Sci. USA 103, 2833–2838 (2006).
Price, K. E., Naimie, A. A., Griffin, E. F., Bay, C. & O’Toole, G. A. Tobramycin-treated Pseudomonas aeruginosa PA14 enhances Streptococcus constellatus 7155 biofilm formation in a cystic fibrosis model system. J. Bacteriol. 198, 237–247 (2015).
Delorenzo, V., Eltis, L., Kessler, B. & Timmis, K. N. Analysis of Pseudomonas gene-products using lacIq/Ptrp-lac plasmids and transposons that confer conditional phenotypes. Gene 123, 17–24 (1993).
Fletcher, M. The effects of culture concentration and age, time, and temperature on bacterial attachment to polystyrene. Can. J. Microbiol. 23, 1–6 (1977).
Heydorn, A. et al. Quantification of biofilm structures by the novel computer program COMSTAT. Microbiol.-SGM 146, 2395–2407 (2000).
Deziel, E., Lepine, F., Milot, S. & Villemur, R. Mass spectrometry monitoring of rhamnolipids from a growing culture of Pseudomonas aeruginosa strain 57RP. BBA-Mol. Cell Biol. L 1485, 145–152 (2000).
Dusane, D. H. et al. Quorum sensing: Implications on rhamnolipid biosurfactant production. Biotechnol. Genet. Eng. Rev. 27, 159–184 (2010).
Choi, M. H. et al. Metabolic relationship between polyhydroxyalkanoic acid and rhamnolipid synthesis in Pseudomonas aeruginosa: comparative C-13 NMR analysis of the products in wild-type and mutants. J. Biotechnol. 151, 30–42 (2011).
Wittgens, A. et al. Novel insights into biosynthesis and uptake of rhamnolipids and their precursors. Appl. Microbiol. Biotechnol. 101, 2865–2878 (2017).
Manner, S., Goeres, D. M., Skogman, M., Vuorela, P. & Fallarero, A. Prevention of Staphylococcus aureus biofilm formation by antibiotics in 96-microtiter well plates and drip flow reactors: Critical factors influencing outcomes. Scientific Rep 7, 1–10 (2017).
Acknowledgements
This work was supported by the Dow Chemical Company and funds derived from the Biotechnology Endowed Professorship at the Pennsylvania State University.
Author information
Authors and Affiliations
Contributions
Experiments were designed by T.L.W., T.G., L.Z., D.S.M., B.Y., and T.K.W. The experiments were performed by T.L.W. except for mass spectrometry which was performed by J.M. and some plasmids were constructed and evaluated by T.G. The manuscript was written by T.L.W. and T.K.W.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wood, T.L., Gong, T., Zhu, L. et al. Rhamnolipids from Pseudomonas aeruginosa disperse the biofilms of sulfate-reducing bacteria. npj Biofilms Microbiomes 4, 22 (2018). https://doi.org/10.1038/s41522-018-0066-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41522-018-0066-1
This article is cited by
-
Oligoribonuclease mediates high adaptability of P. aeruginosa through metabolic conversion
BMC Microbiology (2024)
-
Genome mining of Pseudomonas spp. hints towards the production of under-pitched secondary metabolites
3 Biotech (2023)
-
Pseudomonas aeruginosa biofilm dispersion by the mouse antimicrobial peptide CRAMP
Veterinary Research (2022)
-
Rhamnolipids and surfactin inhibit the growth or formation of oral bacterial biofilm
BMC Microbiology (2020)
-
Metal ions weaken the hydrophobicity and antibiotic resistance of Bacillus subtilis NCIB 3610 biofilms
npj Biofilms and Microbiomes (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.