Abstract
Many plants accumulate transitory starch reserves in their leaves during the day to buffer their carbohydrate supply against fluctuating light conditions, and to provide carbon and energy for survival at night. It is universally accepted that transitory starch is synthesized from ADP-glucose (ADPG) in the chloroplasts. However, the consensus that ADPG is made in the chloroplasts by ADPG pyrophosphorylase has been challenged by a controversial proposal that ADPG is made primarily in the cytosol, probably by sucrose synthase (SUS), and then imported into the chloroplasts. To resolve this long-standing controversy, we critically re-examined the experimental evidence that appears to conflict with the consensus pathway. We show that when precautions are taken to avoid artefactual changes during leaf sampling, Arabidopsis thaliana mutants that lack SUS activity in mesophyll cells (quadruple sus1234) or have no SUS activity (sextuple sus123456) have wild-type levels of ADPG and starch, while ADPG is 20 times lower in the pgm and adg1 mutants that are blocked in the consensus chloroplastic pathway of starch synthesis. We conclude that the ADPG needed for starch synthesis in leaves is synthesized primarily by ADPG pyrophosphorylase in the chloroplasts.
Similar content being viewed by others
Main
Many plants accumulate starch reserves in their leaves during the day, which are remobilized at night to provide carbon and energy for survival at night1,2. It is universally accepted that starch is synthesized from ADP-glucose (ADPG) and that in leaves starch is synthesized in the chloroplasts, primarily in mesophyll cells. However, there is disagreement about the subcellular compartmentation and pathway of ADPG synthesis. Studies on isolated chloroplasts3,4 and starch-deficient mutants of Arabidopsis thaliana5,6,7,8,9,10,11 led to a consensus view that ADPG is produced in the chloroplasts by ADPG pyrophosphorylase (AGPase), with its glucose 1-phosphate (Glc1P) substrate being derived from fructose 6-phosphate (Fru6P) that is withdrawn from the Calvin-Benson cycle in the light, and converted sequentially to glucose 6-phosphate (Glc6P) and then Glc1P by the plastidial phosphoglucose isomerase (PGI) and phosphoglucomutase (PGM), respectively. This pathway is consistent with the near absence of starch in source leaves of pgm and adg1 null mutants that lack plastidial PGM or AGPase, respectively5,6,7. ADPG and starch synthesis are thought to be restricted to the chloroplasts in leaves of all species and to the amyloplasts in non-photosynthetic tissues (for example, Arabidopsis seeds), except for the endosperm of cereals and other Poaceae12,13. Here the majority of ADPG is synthesized by a cytosolic AGPase and imported into the amyloplasts via BRITTLE1 (BT1)-type ADPG transporters, probably in exchange for AMP14,15,16,17,18,19. In barley (Hordeum vulgare) and rice (Oryza sativa), alternative splicing of the AGPS small subunit gene in different tissues leads to expression of a chloroplast-targeted AGPS polypeptide in leaves, while the AGPS polypeptide expressed in the endosperm has no plastidial transit peptide and is located in the cytosol20,21. In maize (Zea mays) endosperm, the cytosolic AGPase derives from gene duplication and neo-functionalization during tetraploidization22.
This consensus has been challenged by a proposal that ADPG is synthesized primarily by sucrose synthase (SUS) in the cytosol, and then imported into the chloroplasts for starch synthesis (Fig. 1b)23,24,25,26,27,28,29. In addition to uridine diphosphate (UDP), the preferred substrate, SUS can use ADP as an alternative substrate, producing ADPG instead of UDPG as a product of sucrose cleavage30. The proposed SUS-mediated pathway of ADPG synthesis was based on two lines of evidence. First, if the consensus pathway operates, levels of ADPG and starch would be expected to be much lower in leaves lacking PGM or AGPase (the pgm and adg1 (aps1) mutants, respectively) than in wild-type (WT) leaves. This was claimed not to be the case: ADPG levels in pgm and adg1 mutants were reported to be similar to those in WT plants28,29. Second, in an attempt to discover the effects of reductions in ADPG levels on starch synthesis in either the chloroplast or the cytosol on starch synthesis, Baroja-Fernández et al.24 expressed an adenosine diphosphate sugar pyrophosphatase (ASPP) from Escherichia coli in one or the other of these compartments in WT plants. This enzyme cleaved ADPG into Glc1P and AMP, but could also hydrolyze other ADP-sugars, with ADP-ribose being the preferred substrate31. Expression of ASPP in chloroplasts had little effect on ADPG levels, but reduced starch accumulation by 50%. Expression in the cytosol reduced ADPG levels by 30% and starch levels by 20%24. From these observations, it was argued that a pool of ADPG located in the cytosol is essential for normal rates of starch synthesis. As AGPase is strictly chloroplastic in Arabidopsis leaves32,33, Baroja-Fernández and colleagues proposed that SUS synthesizes ADPG in the cytosol, using ADP as the glucosyl acceptor in the sucrose-cleavage reaction, followed by transfer of ADPG into the chloroplast via a hypothetical transporter in the chloroplast envelope23. However, it remains possible that the reduction in starch levels in plants with cytosolic ASPP is a pleiotropic effect, brought about by ASPP-mediated changes in ADP-sugars other than ADPG.
a, The classical pathway of starch synthesis (modified from a figure in ref. 68) in which the substrate for starch synthesis, ADP-glucose (ADPG), is synthesized in the chloroplasts by ADP-glucose pyrophosphorylase (AGPase). b, The alternative pathway proposed by Baroja-Fernández et al.23 in which ADPG is primarily produced in the cytosol via sucrose synthase (SUS) and imported into the chloroplasts via a so far unknown transporter for starch synthesis. The model proposes that starch is turned over hydrolytically during the day producing glucose, which is phosphorylated by hexokinase (HXK) and converted back into ADPG via plastidial phosphoglucomutase (pPGM) and ADP-glucose pyrophosphorylase for resynthesis of starch. Fru6P, fructose 6-phosphate; GBSS, granule-bound starch synthase; Glc1P, glucose 1-phosphate; Glc6P, glucose 6-phosphate; ISA1, isoamylase 1; ISA2, isoamylase 2; PGI, plastidial phosphoglucose isomerase; PGM, plastidial phosphoglucomutase; SBE2-SBE3, starch branching enzymes 2–3; SPP, sucrose-phosphate phosphatase; SPS, sucrose-phosphate synthase; SSI-SSIV, soluble starch synthases 1–4; Suc6P, sucrose 6´-phosphate; TPT, triose-phosphate translocator.
The proposed SUS pathway for ADPG synthesis has been critically discussed by Okita34 and Neuhaus et al.35. One of their major criticisms was that the SUS pathway, as originally proposed, did not explain the starch-deficient phenotypes of the pgm and adg1 mutants. In response to this criticism, it was proposed that a futile cycle of starch synthesis and degradation operates inside the chloroplast during the day, in parallel with synthesis and import of cytosolic ADPG. In the proposed futile cycle, starch is degraded hydrolytically to glucose, which is then converted via plastidial hexokinase, PGM and plastidial AGPase to ADPG for resynthesis of starch24,27. According to this scheme, loss of either plastidial PGM or AGPase would prevent recycling of the products of starch degradation and thus prevent starch accumulation. Such futile cycling would need to operate under all conditions and throughout the photoperiod to explain the starch-deficient phenotypes of the pgm and adg1 mutants.
The proposal that futile cycling of starch occurs continuously in the light is called into question by several observations. First, recent 13C-labelling studies have shown that there is some turnover of starch in Arabidopsis leaves in the light, but this is limited to certain times of day and prevalent only in long-day conditions36. Second, Streb et al.37 introgressed the pgm mutation into null mutants for key steps in the major pathway of starch degradation in Arabidopsis leaves2,38: glucan, water dikinase (GWD; encoded by STARCH EXCESS 1), phosphoglucan phosphatase (SEX4), isoamylase3 (ISA3) debranching enzyme and the maltose transporter (MEX1). All of the resulting double mutants were as starch-deficient as pgm, consistent with a requirement for plastidial PGM in the primary pathway of starch synthesis. Third, even if the proposed futile cycling of starch and glucose salvage pathways were operating, the SUS pathway does not readily explain the starch-deficient phenotype of Arabidopsis plastidial pgi mutants9. Proponents of the SUS pathway argue that plastidial pgi mutants are starch-deficient because of a defect in the synthesis of isoprenoid-derived phytohormones (for example, gibberellins) that perturbs their growth and seed production, rather than a simple metabolic block in the flow of carbon from Fru6P, an intermediate of the Calvin-Benson cycle, to ADPG39. Fourth, the SUS pathway also does not explain why triose-phosphate translocator (tpt) mutants, which are unable to export triose-phosphates for sucrose biosynthesis, accumulate more starch than WT plants in the light40,41. Likewise, mutants defective in the pathway of sucrose synthesis usually have elevated starch42,43,44,45. If ADPG were synthesized primarily from sucrose via SUS, all of these mutants would be expected to have less starch than WT. Furthermore, f2kp mutants that lack fructose-2,6-bisphosphate have enhanced rates of sucrose synthesis but less starch than WT plants46, not more as would be expected if ADPG and starch were derived from sucrose. Fifth, there is no experimental evidence that Arabidopsis leaf chloroplasts have the capacity to import ADPG from the cytosol. On the contrary, eudicot forms of the plastidial BRITTLE1 adenylate transporter, including the Arabidopsis orthologue AtBT1, have been shown to transport AMP, ADP and ATP but not ADPG, and Arabidopsis bt1 mutants have near-WT levels of starch47. These observations argue against import of ADPG via AtBT1 from the cytosol making a meaningful contribution to starch synthesis in Arabidopsis leaves.
A direct indication that SUS is not required for starch synthesis in mesophyll cells came from a study of a quadruple sus1 sus2 sus3 sus4 (sus1234) mutant. This lacks all four of the SUS isoforms that are expressed in mesophyll cells and has little or no soluble SUS activity in the leaves, yet has WT levels of starch48,49. The main proponents of the SUS pathway confirmed the WT starch levels in the sus1234 mutant, but argued that the result was inconclusive because, unlike Barratt et al.48, they observed high soluble SUS activity in the sus1234 mutant. They suggested that SUS activity had been underestimated by Barratt et al.48 and that Arabidopsis might have other, cryptic forms of SUS25,50.
Barratt et al.48 also questioned whether WT Arabidopsis leaves have sufficient SUS activity to account for their rates of starch synthesis. This was based on a previous report that SUS activity in WT leaves (0.023 µmol min−1 g−1FW)51 was lower than the rates of starch synthesis typically observed under standard laboratory conditions (up to 0.1 µmol [Glc] min−1 g−1 FW). Bieniewska et al.51 measured SUS activity at 25 °C in the direction of sucrose synthesis by the production of [14C]sucrose from UDP-[14C]glucose and fructose. Baroja-Fernández et al.25 contended that SUS activity had been underestimated due to instability of the UDP-glucose substrate under the assay conditions. They reported an activity of 0.185 µmol min−1 g−1 FW in WT leaves, measured at 37 °C in the sucrose cleavage direction using a high-performance liquid chromatography (HPLC)-based assay for UDP-glucose and ADPG, which would be sufficient to mediate observed rates of starch synthesis. The basis for these claims was in turn questioned by studies showing that UDP-glucose is more stable at moderately alkaline pH than claimed by Baroja- Fernández et al.25,52. These conflicting reports leave open the question of whether there is sufficient SUS activity to sustain observed rates of starch synthesis via the proposed cytosolic pathway of ADPG synthesis.
In addition to the genetic evidence favouring the consensus pathway, there are biochemical data that conflict with the proposed SUS pathway. Non-aqueous fractionation of Arabidopsis rosettes showed that ADPG co-localized with the chloroplastic markers, with 100% of the total ADPG being assigned to the chloroplastic fraction in a two-compartment model (chloroplast vs cytosol)53. When WT Arabidopsis rosettes were labelled with 13CO2 under steady-state photosynthetic conditions, the pool of ADPG was labelled more rapidly than sucrose, and had kinetics similar to those of Calvin-Benson cycle intermediates53. These labelling patterns are consistent with direct synthesis of ADPG from intermediates of the Calvin-Benson cycle in the chloroplasts, rather than via sucrose in the cytosol.
Transitory starch is a major end product of photosynthesis, therefore uncertainty about its synthesis is a major obstacle to understanding the regulation of photosynthetic carbon metabolism and how the fixed carbon is allocated between storage and export for growth. To resolve the long-standing controversy surrounding the pathway of starch synthesis in leaves, we critically re-examine the main lines of evidence that ADPG is synthesized by SUS in the cytosol for transitory starch synthesis in the chloroplasts. Furthermore, we generated Arabidopsis mutants that completely lack SUS activity and report that these sextuple (sus1 sus2 sus3 sus4 sus5 sus6) mutants have near wild-type levels of ADPG and starch, and only mild growth phenotypes under standard laboratory growth conditions.
Results
Optimization of ADPG measurement in Arabidopsis leaves
HPLC with UV detection at 260 nm (HPLC-UV) or pulsed amperometric detection (HPLC-PAD) has been widely used to assay ADPG, including all but the most recent study29 claiming support for the SUS pathway. However, plant extracts contain many other types of nucleotide that absorb in the UV range54, and it is difficult to achieve baseline separation of ADPG from other UV-absorbing compounds, potentially leading to overestimation of ADPG. Likewise, HPLC-PAD, as used in Baroja-Fernández et al.25, cannot distinguish ADPG from any co-eluting molecules, so is also prone to overestimation of ADPG. HPLC coupled to tandem mass spectrometry (LC-MS/MS) offers much greater specificity and sensitivity than HPLC-UV or HPLC-PAD55, and has become the method of choice for assaying ADPG.
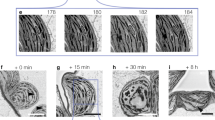
The levels of ADPG in Arabidopsis leaves are generally low (<5 nmol g−1 FW), with an estimated turnover time of <1 s in illuminated leaves56, suggesting that leaf sampling in the light must be done with great care to avoid artefactual decreases in ADPG content below the steady-state level before the tissue is quenched. To assess the sensitivity of ADPG to unintentional shading during leaf sampling, rosettes were harvested from WT, pgm and adg1 Arabidopsis plants in the light either by pouring liquid nitrogen directly onto the plants, or by cutting the hypocotyl from below and then quickly transferring the cut rosette into liquid nitrogen while in the growth chamber, taking care not to shade the leaves at any time during the procedure. In parallel, rosettes were harvested from plants that had been deliberately shaded by transfer from 160 µE m−2 s−1 to <10 µE m−2 s−1 for 2–3 s. This brief shading treatment led to a large (94%) fall in rosette ADPG levels (measured by LC–MS/MS) compared with rosettes that had been harvested in the light without shading (Fig. 2a). The pgm mutant contained much lower levels of ADPG than WT plants, but there was no significant effect of shading (Fig. 2b), while the adg1 mutant had even lower levels than pgm that were decreased even further by shading (Fig. 2c). These results demonstrate the sensitivity of ADPG in WT plants to artefactual changes due to shading during leaf sampling, hence the need for extremely rapid quenching under full illumination. Pouring liquid nitrogen directly onto the plants is inconvenient and potentially hazardous, so for subsequent experiments we cut the hypocotyl from below and immediately transferred the cut rosette into liquid nitrogen within the growth chamber.
a–c, Rosettes were harvested from vegetative wild-type Col-0 plants (a) and from starch-deficient pgm (b) and adg1 (c) mutants in the light (white bars) by pouring liquid nitrogen (liq. N) directly onto the rosette or by cutting the hypocotyl from below and transferring the cut rosette into liquid nitrogen, taking care to avoid all shading. In parallel, other plants were deliberately shaded (grey bars) for 2–3 s before cutting and snap-freezing the rosettes. ADPG was measured by LC–MS/MS. Bars show mean ± s.d. of 5 independently pooled batches of 5 rosettes (n = 5 biological replicates). P values are shown for pairwise comparisons from a one-way analysis of variance (ANOVA) (Holm-Sidak post-hoc test). There were no significant differences between the samples in b (P = 0.285).
Comparison of ADPG levels in wild-type and mutant Arabidopsis plants
WT, pgm, adg1 and sus1234 plants were grown in long-day (16 h photoperiod) conditions, and rosettes were harvested at the end of the night and at intervals during the day for metabolite analysis. There were no significant differences in Glc6P and Glc1P between WT and sus1234 plants (Fig. 3a,b and Supplementary Table 1). In the pgm and adg1 mutants, Glc6P was lower than in WT in the dark, but increased to a greater extent upon illumination and was significantly higher than in WT from about Zeitgeber time (ZT) 4 (that is, 4 h after the start of the light period) onwards (Fig. 3a). Glc1P was lower in the pgm mutant than in WT, but higher in the adg1 mutant than in WT (Fig. 3b). ADPG levels were extremely low (<0.005 nmol g−1 FW) in all genotypes in the dark (Fig. 3c). Upon illumination, ADPG increased rapidly in WT and sus1234 plants, reaching a similar level (0.51–0.71 nmol g−1 FW) in the two genotypes and remaining fairly constant throughout the day with no significant differences between WT and sus1234 plants (Fig. 3c). In contrast, ADPG rose only slightly upon illumination of pgm and adg1, reaching levels of 0.03 nmol g−1 FW and 0.01–0.02 nmol g−1 FW, respectively (Fig. 3c). The sus1234 plants accumulated the same amounts of starch as WT plants, whereas at ZT12 the starch contents of the pgm and adg1 mutants were <1% of WT (Fig. 3d). The lag in net starch accumulation in WT and sus1234 at the beginning of the day is most likely due to starch degradation occurring in parallel with synthesis as previously observed for plants growing in long-day (16 h photoperiod) conditions36.
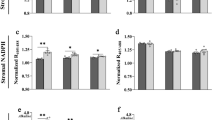
a–d, Wild-type Col-0, the sus1234 mutant (susquad) and two starch-deficient mutants (pgm and adg1) were grown in long-day conditions (16 h photoperiod). At 25 d after germination, rosettes were harvested just before dawn (ZT0.2) and at intervals from ZT0.5 to ZT12 for measurement of: (a) glucose 6-phosphate (Glc6P), (b) glucose 1-phosphate (Glc1P), (c) ADP-glucose (ADPG) and (d) starch. Data are mean ± s.d. of 4 independently pooled batches of 5 rosettes (n = 4 biological replicates). P values for all genotype × genotype comparisons are shown in Supplementary Table 2.
WT plants were labelled with 13CO2 under ambient CO2 conditions to compare the labelling kinetics of ADPG with UDPG. The latter nucleotide-sugar is the substrate for sucrose biosynthesis and potential product of sucrose cleavage by SUS in the cytosol. ADPG was labelled much more rapidly and more completely than UDPG, with very similar labelling kinetics to Calvin-Benson cycle intermediates (Supplementary Fig. 1 and Table 1).
Generation and analysis of sextuple sus123456 mutants
Based on localization of the SUS1–SUS4 isoforms48,49, the quadruple sus1234 mutant is expected to lack SUS activity in the mesophyll cells where transitory starch is synthesized57. The residual SUS activity in this mutant is disputed, with Barratt et al.48 reporting that soluble SUS activities in roots and stems of the quadruple sus1234 mutant were less than 2% of those in WT plants, whereas Baroja-Fernández et al.25 reported soluble SUS activities of 98% and 90% of WT in leaves and stems of the sus1234 mutant. To resolve these discrepancies, we generated two independent sextuple sus1 sus2 sus3 sus4 sus5 sus6 (sus1234516 and sus1234526) mutants by CRISPR/Cas9-mediated gene editing of the SUS5 and SUS6 genes in the sus1234 mutant background. The CRISPR (clustered regularly interspaced short palindromic repeats)/Cas9 (CRISPR-associated protein 9)-generated mutations in the SUS5 (sus5-1 and sus5-2) and SUS6 (sus6) genes gave rise to premature stop codons (Supplementary Fig. 2) within the respective regions coding for the catalytic glucosyltransferase domains of the enzymes. The zygosity of the sus5 and sus6 alleles was tested by genomic PCR using gene-specific primers followed by restriction enzyme digestion of the PCR products (Supplementary Fig. 3). The presence of homozygous T-DNA insertions in the SUS1, SUS2, SUS3 and SUS4 genes was confirmed in both of the gene-edited lines (Supplementary Fig. 3), establishing that the homozygous sextuple mutants (sus1234516 and sus1234526) lacked functional copies of all six Arabidopsis SUS genes.
To confirm loss of SUS activity, we measured activity in developing siliques and in rosette leaves of WT, sus1234 and the two sextuple sus123456 mutants. Developing siliques were chosen initially because all six SUS genes are expressed in siliques and/or seeds, as visualized using the Plant eFP browser (https://bar.utoronto.ca/eplant; Supplementary Fig. 4)58,59, and SUS activity is high and readily quantified in these tissues60. Activity was determined as the UDP-dependent production of UDP-glucose from sucrose, with UDP-glucose being measured enzymatically as previously described61. The soluble SUS activity in WT Col-0 siliques was 86.8 ± 16.7 nmol min−1 g−1 FW (Fig. 4a) and the assay was linear with time for at least 40 min (Fig. 4b). Activities were much lower or undetectable in the sus1234 (1.6 ± 1.9 nmol min−1 g−1 FW; <2% of WT), sus1234516 (−0.2 ± 2.9 nmol min−1 g−1 FW; that is, no detectable activity) and sus1234526 (0.2 ± 2.0 nmol min−1 g−1 FW; <0.2% of WT) mutants (Fig. 4a). The soluble SUS activity in WT Col-0 rosettes was 17.4 ± 5.9 nmol min−1 g−1 FW (Fig. 4a) and the assay was linear with time for at least 60 min (Fig. 4b). Activities were much lower or undetectable in the sus1234 (−4.0 ± 1.9 nmol min−1 g−1 FW; that is, no detectable activity), sus1234516 (0.3 ± 1.9 nmol min−1 g−1 FW; <2% of WT) and sus1234526 (−1.0 ± 2.4 nmol min−1 g−1 FW; that is, no detectable activity) mutants (Fig. 4a). The residual activities detected in extracts of sus1234526 siliques and sus1234516 rosettes were within the range of values obtained with a boiled extract of Col-0 siliques (0.6 ± 1.2 nmol min−1 g−1 FW; Fig. 4), indicating that the apparent activities simply reflect noise at the detection limit of the assay. We conclude that the sextuple sus123456 mutants have no detectable soluble SUS activity. The sus123456 mutants showed WT-like growth under standard laboratory conditions (Fig. 4c,d), indicating that SUS is not essential for plant survival under benign growth conditions.
a, Sucrose synthase activity of developing siliques and rosettes, measured as the UDP-dependent production of UDP-glucose from sucrose. Bars show mean ± s.d. from 6 independently pooled siliques (n = 6 biological replicates) or independently pooled batches of 5 rosettes from Col-0 (n = 10 biological replicates), susquad(n = 4 biological replicates), sussext-1 (n = 4 biological replicates), sussext-2 (n = 3 biological replicates) and Col-0 boiled (n = 3 biological replicates). b, Linearity of sucrose synthase assay. Datapoints are shown as mean ± s.d. of 3 technical replicates from single wild-type Col-0 silique and rosette extracts. c, Seedling morphology at 5 d after germination on 0.5× MS medium. Scale bar, 5 mm. (d) Rosette morphology of plants grown in a 16 h photoperiod for 3 weeks or an 8 h photoperiod for 4 weeks. Scale bars, 1 cm. susquad, sus1234; sussext-1, sus1234516; sussext-2, sus1234526.
WT Col-0, sus1234 and sus1234516 plants were grown under long-day (16 h photoperiod) conditions and rosettes were harvested just before dawn and at intervals during the day for metabolite analysis. In WT Col-0 plants, Glc6P, Glc1P and ADPG increased upon illumination (Fig. 5a–c), as observed previously (Fig. 3), and after a slight lag the plants accumulated starch in a linear manner (Fig. 5d). The levels and diel patterns of Glc1P, Glc6P, ADPG and starch were essentially identical in the sus1234 and sus1234516 plants and not significantly different from those of WT plants (Fig. 5 and Supplementary Table 2). Likewise, there were no significant differences between the mutant and WT Col-0 plants in the levels of soluble sugars (sucrose, glucose and fructose) or intermediates of sucrose synthesis (Fru6P, UDP-glucose, dihydroxyacetone-phosphate; Supplementary Fig. 5 and Table 3). In an independent experiment under the same growth conditions, sus1234526 plants had levels and diel patterns of starch and soluble sugars that were essentially identical to those in WT plants (Supplementary Fig. 6).
a–d, Wild-type Col-0, sus1234 mutant (susquad) and sus1234516 (sussext-1) mutants were grown in long-day conditions (16 h photoperiod). At 25 d after germination, rosettes were harvested just before dawn (ZT0.2) and at intervals from ZT0.5 to ZT12 for measurement of (a) glucose 6-phosphate (Glc6P), (b) glucose 1-phosphate (Glc1P), (c) ADP-glucose (ADPG) and (d) starch. Data are mean ± s.d. from 3 independently pooled batches of 5 rosettes (n = 3 biological replicates). P values for all genotype × genotype comparisons are shown in Supplementary Table 3.
Discussion
According to the consensus pathway, the starch-deficient pgm and adg1 mutants should have little or no ADPG due to loss of essential enzymes for its synthesis in the chloroplasts. Reports of near-WT levels of ADPG in these two mutants28,29 appeared to be inconsistent with expectations. To reassess these reports, we established a robust method for harvesting Arabidopsis rosettes in the light to maintain in vivo levels of metabolites such as ADPG that are present in very small amounts and turn over very quickly (half time <1 s)56. We used LC–MS/MS55,56 to assay ADPG with greater sensitivity and specificity than the HPLC-UV/HPLC-PAD methods that were used for most previous studies. We found that shading of the leaves of WT plants for just a few seconds leads to a rapid decrease in measurable ADPG content, down to levels that resemble those found in the starch-deficient pgm and adg1 mutants. Consequently, unless extreme care is taken during sample collection, ADPG levels are likely to be underestimated and therefore unrepresentative of the levels during steady-state photosynthesis. This issue may well have affected values presented in earlier papers, including some of our own.
Using our optimized methods, we found that ADPG levels in illuminated leaves of the pgm and adg1 mutants were <6% and <4% of WT, respectively, but the same as WT in the sus1234 mutant (Fig. 3c). In contrast, and using an LC–MS-based method similar to our own, Bahaji et al.29 reported that ADPG levels were the same in WT leaves and leaves of the pgm and aps1 mutants (aps1, similar to adg1, lacks a subunit of AGPase and is nearly starchless). In earlier work that employed the HPLC-UV assay for ADPG, these researchers reported that adg1 leaves also had ADPG levels comparable with those of WT leaves28. One possible explanation for this discrepancy between our results and those of Bahaji and colleagues is that their plants were inadvertently shaded during harvest, potentially leading to large losses of ADPG before quenching. Consistent with this idea is the fact that their reported values for ADPG in WT leaves (0.13 nmol g−1 FW)29 are substantially lower than the range we observed in WT leaves (0.5–3 nmol g−1 FW; Figs. 2a, 3c and 5c). Their low value for WT leaves is comparable with the amounts we observed after 2–3 s of shading, and with our shaded values for pgm and adg1. Thus, we suspect that the similar ADPG values for WT and mutant plants reported by Bahaji et al.29, and the discrepancies between their results and ours are due to loss of ADPG during harvesting of WT leaves in their experiments.
Our data are entirely consistent with predictions based on the consensus pathway being the predominant, if not the only, pathway of starch synthesis in leaves. Given the precautions we took to ensure that in vivo metabolite levels were preserved during tissue sampling and the robustness of the LC–MS/MS assay for ADPG, we would argue that our results are more trustworthy than those used by others to question the validity of the consensus pathway.
The soluble SUS activity in WT Col-0 rosettes, measured under linear conditions with an established assay (Fig. 4b), was 17.4 ± 5.9 nmol min−1 g−1 FW (Fig. 4a), which is too low to account for the observed rates of starch synthesis in WT leaves under our experimental conditions (approx. 40 nmol [Glc] min−1 g−1 FW; Figs. 3d and 5d,e). Nevertheless, this does not exclude the possibility of a minor contribution to starch synthesis from ADPG produced via SUS. Therefore, to test whether the proposed alternative pathway of ADPG synthesis via SUS makes any significant contribution to starch synthesis, we re-analysed the sus1234 mutant reported in Barratt et al.48 and also generated two sextuple sus123456 mutants that lack all known isoforms of SUS in Arabidopsis.
The sus1234 mutant lacks SUS expression in mesophyll cells where transitory starch is made in WT plants48,57. The mutant has little or no detectable soluble SUS activity in developing siliques, where activity is high and readily measurable in WT plants, or in rosette leaves (Fig. 4a). The sus1234 mutant should have less starch than WT plants if the SUS pathway supplies a significant amount of ADPG for starch synthesis. However, we confirmed that starch levels in this mutant are indistinguishable from that of WT, as reported in Barratt et al.48 and Baroja-Fernández et al.25, and showed that the mutant also has WT levels of ADPG and of the two other intermediates of the consensus pathway, Glc6P and Glc1P (Figs. 3 and 5). Likewise, sextuple sus123456 mutants that had no detectable SUS activity in developing siliques or rosettes contained WT levels of starch (Fig. 4e and Supplementary Fig. 5a). Key intermediates from the consensus pathway for starch synthesis (Glc6P, Glc1P, ADPG) were also essentially the same as in WT (Fig. 4a–c), as were soluble sugars and intermediates of sucrose synthesis (dihydroxyacetone phosphate, Fru6P, UDPG) (Fig. 5, and Supplementary Figs. 5 and 6).
In principle, it is conceivable that SUS does contribute to ADPG formation and starch synthesis in wild-type leaves, but that in the quadruple and sextuple sus mutants this function can be completely substituted by the consensus pathway via chloroplastic ADPG pyrophosphorylase. However, two observations make this extremely improbable. First, it has previously been shown that essentially all of the ADPG in Arabidopsis leaves is located in the chloroplasts53. Second, pulse-labelling with 13CO2 showed that ADPG is rapidly labelled with similar kinetics to Calvin-Benson cycle intermediates (Supplementary Fig. 1), confirming previous observations53. Labelling of ADPG was both more rapid and more complete than labelling of UDPG, the precursor of sucrose and potential product of sucrose cleavage via SUS (Supplementary Fig. 1). Together, these results show that SUS is not required for transitory starch synthesis in Arabidopsis leaves, and argue strongly against the proposed SUS pathway making any significant contribution to ADPG and starch synthesis in WT leaves.
Methods
Plant materials
Arabidopsis (Arabidopsis thaliana [L.] Heynh) Columbia-0 wild-type and the pgm5, adg17 and sus123448 mutant germplasm were from in-house collections.
Sextuple sus123456 mutants were generated by gene editing of the SUS5 and SUS6 genes in the sus123448 background using CRISPR/Cas9 and two pairs of guide RNA (gRNA) targets: SUS5—GAAATGACATCTGGATCGTT and TGTAGAACTTGGTGAATCTC; SUS6—GGTAAGGGTAGATATCGAAT and
GTTCTTGAAGCACCAGACAA. The CRISPR/Cas9 and gRNA sequences were cloned into the pHEE401E vector as previouslly described62,63. The construct was introduced into the sus1234 quadruple mutant by Agrobacterium-mediated transformation using the floral-dip method64 to generate sextuple sus123456 mutants. The CRISPR/Cas9 construct was subsequently eliminated by crossing the sextuple sus123456 lines with the quadruple sus1234 parent. Progeny were screened by PCR to identify lines that were homozygous for the T-DNA insertions in SUS1–SUS4 and for gene-edited mutant alleles of SUS5 and SUS6 (Supplementary Fig. 3). Mutations in the SUS5 gene were identified by PCR using primers sus5-Fw and sus5-Rv (Supplementary Table 4) and restriction with HinfI. Mutations in the SUS6 gene were identified by PCR using primers sus6-Fw and sus6-Rv (Supplementary Table 4) and restriction with Van91I (Supplementary Fig. 3).
Plants were grown in a 1:1 mixture of compost and vermiculite in 6-cm-diameter pots with a 16 h photoperiod (140 μE m−2 s−1 irradiance provided by white fluorescent tubes) and day/night temperatures of 20 °C/18 °C. For the 13CO2-labelling experiment, plants were grown with a 12 h photoperiod (120 µE m−2 s−1 irradiance). Seedlings were also grown on agar plates containing 0.5× Murashige–Skoog medium.
Enzyme activity measurements
Finely ground frozen leaf tissue (20 mg) was homogenized in 1 ml of ice-cold extraction buffer as previously described65. After centrifugation of the crude extract at 10,000 × g (4 °C) for 5 min, an aliquot (180 µl) of the supernatant was desalted by centrifugation through a MicroSpin column (GE Healthcare Life sciences) equilibrated with extraction buffer. Soluble SUS activity was determined in the sucrose cleavage direction by measuring the UDP-dependent production of UDPG from sucrose, with enzymatic determination of UDPG as previously described61. Reaction mixtures (225 µl) contained: 20 mM PIPES-KOH (pH 6.5), 3 mM MgCl2, 100 mM sucrose and 2 mM UDP, with UDP being omitted from blank reactions. Reactions were started by addition of 25 µl of desalted extract and incubation at 30 °C for 25 min. The reaction was stopped by addition of 250 µl 50 mM Tricine-KOH (pH 8.3) and heating at 100 °C for 2 min. UDPG was measured enzymatically by coupling to reduction of NADP+ and monitoring the increase in absorbance at 340 nm. The initial reaction mixture (500 µl) contained: 50 mM Tricine-HCl (pH 8.0), 4 mM MgCl2, 12.5 mM NADP+, 0.3 U glucose-6-phosphate dehydrogenase (EC 1.1.1.49; from yeast), 0.4 U phosphoglucomutase (EC 5.4.2.2; from rabbit muscle) and 400 µl of the SUS assay reaction. When the A340 was stable, UDPG was determined by addition of sodium pyrophosphate (final concentration 1 mM) and 0.2 U UDP-glucose pyrophosphorylase (EC 2.7.7.9; from yeast). All coupling enzymes were from Sigma-Aldrich.
Metabolite measurements
Rosettes were rapidly quenched in the light by either pouring liquid nitrogen directly onto the rosettes, or by cutting the hypocotyl from below and rapidly transferring the cut rosette into liquid nitrogen in the light, taking care not to shade the leaves at any time. Phosphorylated intermediates were extracted using chloroform-methanol as previously described55 and measured by ion-pair reverse phase LC–MS/MS56 or anion-exchange LC–MS/MS55 (with modifications as previously described66). Soluble sugars were measured in ethanolic extracts and starch was measured in the ethanol-insoluble residue as previously described67.
13CO2 pulse labelling
Four-week-old WT Arabidopsis Col-0 plants were pulse-labelled with 13CO2 (400 µl l−1) at an irradiance of 120 µE m−2 s−1, for 0, 1 or 20 min, and rapidly quenched with liquid nitrogen under ambient illumination as previously described53. Labelling was performed between ZT4 and ZT7. Isotopomers of nucleotide sugars and phosphorylated intermediates were quantified by LC–MS/MS53.
Statistical analysis
Statistical analysis was performed using Microsoft Excel 2019 MSO (https://www.microsoft.com) or Sigmaplot for Windows version 14.5 (Systat Software; http://www.systat.de).
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Original data and Arabidopsis lines are available upon request from the corresponding authors.
References
Stitt, M. & Zeeman, S. C. Starch turnover: pathways, regulation and role in growth. Curr. Opin. Plant Biol. 15, 282–292 (2012).
Smith, A. M. & Zeeman, S. C. Starch: a flexible, adaptable carbon store coupled to plant growth. Annu. Rev. Plant Biol. 71, 217–245 (2020).
Heldt, H. W. et al. Role of orthophosphate and other factors in the regulation of starch formation in leaves and isolated chloroplasts. Plant Physiol. 59, 1146–1155 (1977).
Okita, T. W., Greenberg, E., Kuhn, D. N. & Preiss, J. Subcellular localization of the starch degradative and biosynthetic enzymes of spinach leaves. Plant Physiol. 64, 187–192 (1979).
Caspar, T., Huber, S. C. & Somerville, C. Alterations in growth, photosynthesis, and respiration in a starchless mutant of Arabidopsis thaliana (L.) deficient in chloroplast phosphoglucomutase activity. Plant Physiol. 79, 11–17 (1985).
Lin, T. P., Caspar, T., Somerville, C. & Preiss, J. Isolation and characterization of a starchless mutant of Arabidopsis thaliana (L.) Heynh lacking ADPglucose pyrophosphorylase activity. Plant Physiol. 86, 1131–1135 (1988).
Lin, T. P., Caspar, T., Somerville, C. R. & Preiss, J. A starch deficient mutant of Arabidopsis thaliana with low ADPglucose pyrophosphorylase activity lacks one of the two subunits of the enzyme. Plant Physiol. 88, 1175–1181 (1988).
Ekkehard Neuhaus, H. & Stitt, M. Control analysis of photosynthate partitioning: impact of reduced activity of ADP-glucose pyrophosphorylase or plastid phosphoglucomutase on the fluxes to starch and sucrose in Arabidopsis thaliana (L.) Heynh. Planta 182, 445–454 (1990).
Yu, T.-S., Lue, W.-L., Wang, S.-M. & Chen, J. Mutation of Arabidopsis plastid phosphoglucose isomerase affects leaf starch synthesis and floral initiation. Plant Physiol. 123, 319–326 (2000).
Hädrich, N. et al. Use of TILLING and robotised enzyme assays to generate an allelic series of Arabidopsis thaliana mutants with altered ADP-glucose pyrophosphorylase activity. J. Plant Physiol. 168, 1395–1405 (2011).
Wang, S. M. et al. Characterization of ADG1, an Arabidopsis locus encoding for ADPG pyrophosphorylase small subunit, demonstrates that the presence of the small subunit is required for large subunit stability. Plant J. 13, 63–70 (1998).
Crevillén, P., Ballicora, M. A., Mérida, A., Preiss, J. & Romero, J. M. The different large subunit isoforms of Arabidopsis thaliana ADP-glucose pyrophosphorylase confer distinct kinetic and regulatory properties to the heterotetrameric enzyme. J. Biol. Chem. 278, 28508–28515 (2003).
Figueroa, C. M., Asencion Diez, M. D., Ballicora, M. A. & Iglesias, A. A. Structure, function, and evolution of plant ADP-glucose pyrophosphorylase. Plant Mol. Biol. 108, 307–323 (2022).
Denyer, K., Dunlap, F., Thorbjornsen, T., Keeling, P. & Smith, A. M. The major form of ADP-glucose pyrophosphorylase in maize endosperm is extra-plastidial. Plant Physiol. 112, 779–785 (1996).
Thorbjørnsen, T., Villand, P., Denyer, K., Olsen, O. A. & Smith, A. M. Distinct isoforms of ADPglucose pyrophosphorylase occur inside and outside the amyloplasts in barley endosperm. Plant J. 10, 243–250 (1996).
Shannon, J. C., Pien, F.-M., Cao, H. & Liu, K.-C. Brittle-1, an adenylate translocator, facilitates transfer of extraplastidial synthesized ADP-glucose into amyloplasts of maize endosperms. Plant Physiol. 117, 1235–1252 (1998).
Beckles, D. M., Smith, A. M. & ap Rees, T. A cytosolic ADP-glucose pyrophosphorylase is a feature of graminaceous endosperms, but not of other starch-storing organs. Plant Physiol. 125, 818–827 (2001).
Emes, M. et al. Starch synthesis and carbon partitioning in developing endosperm. J. Exp. Bot. 54, 569–575 (2003).
Kirchberger, S. et al. Molecular and biochemical analysis of the plastidic ADP-glucose transporter (ZmBT1) from Zea mays. J. Biol. Chem. 282, 22481–22491 (2007).
Rösti, S. et al. The gene encoding the cytosolic small subunit of ADP-glucose pyrophosphorylase in barley endosperm also encodes the major plastidial small subunit in the leaves. J. Exp. Bot. 57, 3619–3626 (2006).
Lee, S.-K. et al. Identification of the ADP-glucose pyrophosphorylase isoforms essential for starch synthesis in the leaf and seed endosperm of rice (Oryza sativa L.). Plant Mol. Biol. 65, 531–546 (2007).
Rösti, S. & Denyer, K. Two paralogous genes encoding small subunits of ADP-glucose pyrophosphorylase in maize, Bt2 and L2, replace the single alternatively spliced gene found in other cereal species. J. Mol. Evol. 65, 316–327 (2007).
Baroja-Fernandez, E., Munoz, F. J., Akazawa, T. & Pozueta-Romero, J. Reappraisal of the currently prevailing model of starch biosynthesis in photosynthetic tissues: a proposal involving the cytosolic production of ADP-glucose by sucrose synthase and occurrence of cyclic turnover of starch in the chloroplast. Plant Cell Physiol. 42, 1311–1320 (2001).
Baroja-Fernández, E. et al. Most of ADP × glucose linked to starch biosynthesis occurs outside the chloroplast in source leaves. Proc. Natl Acad. Sci. USA 101, 13080–13085 (2004).
Baroja-Fernández, E. et al. Sucrose synthase activity in the sus1/sus2/sus3/sus4 Arabidopsis mutant is sufficient to support normal cellulose and starch production. Proc. Natl Acad. Sci. USA 109, 321–326 (2012).
Munoz, F. J. et al. Sucrose synthase controls both intracellular ADP glucose levels and transitory starch biosynthesis in source leaves. Plant Cell Physiol. 46, 1366–1376 (2005).
Muñoz, F. J., Baroja-Fernández, E., Morán-Zorzano, M. T., Alonso-Casajús, N. & Pozueta-Romero, J. Cloning, expression and characterization of a Nudix hydrolase that catalyzes the hydrolytic breakdown of ADP-glucose linked to starch biosynthesis in Arabidopsis thaliana. Plant Cell Physiol. 47, 926–934 (2006).
Bahaji, A. et al. Arabidopsis thaliana mutants lacking ADP-glucose pyrophosphorylase accumulate starch and wild-type ADP-glucose content: further evidence for the occurrence of important sources, other than ADP-glucose pyrophosphorylase, of ADP-glucose linked to leaf starch biosynthesis. Plant Cell Physiol. 52, 1162–1176 (2011).
Bahaji, A. et al. HPLC-MS/MS analyses show that the near-starchless aps1 and pgm leaves accumulate wild type levels of ADPglucose: further evidence for the occurrence of important ADPglucose biosynthetic pathway(s) alternative to the pPGI-pPGM-AGP pathway. PLoS ONE 9, e104997 (2014).
Schmölzer, K., Gutmann, A., Diricks, M., Desmet, T. & Nidetzky, B. Sucrose synthase: a unique glycosyltransferase for biocatalytic glycosylation process development. Biotech. Adv. 34, 88–111 (2016).
Moreno-Bruna, B. et al. Adenosine diphosphate sugar pyrophosphatase prevents glycogen biosynthesis in Escherichia coli. Proc. Natl Acad. Sci. USA 98, 8128–8132 (2001).
Ballicora, M. A., Iglesias, A. A. & Preiss, J. ADP-glucose pyrophosphorylase: a regulatory enzyme for plant starch synthesis. Photosynth. Res. 79, 1–24 (2004).
Arrivault, S. et al. Dissecting the subcellular compartmentation of proteins and metabolites in Arabidopsis leaves using non-aqueous fractionation. Mol. Cell. Proteom. 13, 2246–2259 (2014).
Okita, T. W. Is there an alternative pathway for starch synthesis? Plant Physiol. 100, 560–564 (1992).
Neuhaus, H. E., Häusler, R. E. & Sonnewald, U. No need to shift the paradigm on the metabolic pathway to transitory starch in leaves. Trends Plant Sci. 10, 154–156 (2005).
Fernandez, O. et al. Leaf starch turnover occurs in long days and in falling light at the end of the day. Plant Physiol. 174, 2199–2212 (2017).
Streb, S., Egli, B., Eicke, S. & Zeeman, S. C. The debate on the pathway of starch synthesis: a closer look at low-starch mutants lacking plastidial phosphoglucomutase supports the chloroplast-localized pathway. Plant Physiol. 151, 1769–1772 (2009).
Niittylä, T. et al. A previously unknown maltose transporter essential for starch degradation in leaves. Science 303, 87–89 (2004).
Bahaji, A. et al. Plastidial phosphoglucose isomerase is an important determinant of seed yield through its involvement in gibberellin-mediated reproductive development and storage reserve biosynthesis in Arabidopsis. Plant Cell 30, 2082–2098 (2018).
Schneider, A. et al. An Arabidopsis thaliana knock‐out mutant of the chloroplast triose phosphate/phosphate translocator is severely compromised only when starch synthesis, but not starch mobilisation, is abolished. Plant J. 32, 685–699 (2002).
Walters, R. G., Ibrahim, D. G., Horton, P. & Kruger, N. J. A mutant of Arabidopsis lacking the triose-phosphate/phosphate translocator reveals metabolic regulation of starch breakdown in the light. Plant Physiol. 135, 891–906 (2004).
Strand, Å. et al. Decreased expression of two key enzymes in the sucrose biosynthesis pathway, cytosolic fructose‐1,6‐bisphosphatase and sucrose phosphate synthase, has remarkably different consequences for photosynthetic carbon metabolism in transgenic Arabidopsis thaliana. Plant J. 23, 759–770 (2000).
Sun, J., Zhang, J., Larue, C. T. & Huber, S. C. Decrease in leaf sucrose synthesis leads to increased leaf starch turnover and decreased RuBP regeneration‐limited photosynthesis but not Rubisco‐limited photosynthesis in Arabidopsis null mutants of SPSA1. Plant Cell Environ. 34, 592–604 (2011).
Kunz, H.-H. et al. Loss of cytosolic phosphoglucose isomerase affects carbohydrate metabolism in leaves and is essential for fertility of Arabidopsis. Plant Physiol. 166, 753–765 (2014).
Malinova, I. et al. Reduction of the cytosolic phosphoglucomutase in Arabidopsis reveals impact on plant growth, seed and root development, and carbohydrate partitioning. PLoS ONE 9, e112468 (2014).
McCormick, A. J. & Kruger, N. J. Lack of fructose 2,6‐bisphosphate compromises photosynthesis and growth in Arabidopsis in fluctuating environments. Plant J. 81, 670–683 (2015).
Kirchberger, S., Tjaden, J. & Ekkehard Neuhaus, H. Characterization of the Arabidopsis Brittle1 transport protein and impact of reduced activity on plant metabolism. Plant J. 56, 51–63 (2008).
Barratt, D. P. et al. Normal growth of Arabidopsis requires cytosolic invertase but not sucrose synthase. Proc. Natl Acad. Sci. USA 106, 13124–13129 (2009).
Yao, D., Gonzales-Vigil, E. & Mansfield, S. D. Arabidopsis sucrose synthase localization indicates a primary role in sucrose translocation in phloem. J. Exp. Bot. 71, 1858–1869 (2020).
Baroja-Fernández, E., Muñoz, F. J., Bahaji, A., Almagro, G. & Pozueta-Romero, J. Reply to Smith et al.: no evidence to challenge the current paradigm on starch and cellulose biosynthesis involving sucrose synthase activity. Proc. Natl Acad. Sci. USA 109, E777 (2012).
Bieniawska, Z. et al. Analysis of the sucrose synthase gene family in Arabidopsis. Plant J. 49, 810–828 (2007).
Hill, B. L. et al. On the stability of nucleoside diphosphate glucose metabolites: implications for studies of plant carbohydrate metabolism. J. Exp. Bot. 68, 3331–3337 (2017).
Szecowka, M. et al. Metabolic fluxes in an illuminated Arabidopsis rosette. Plant Cell 25, 694–714 (2013).
Dawson R. M. C., Elliott, D. C., Elliot W. H. & Jones K. M. Data for Biochemical Research 3rd edn 103–114 (Oxford Univ. Press, 1986).
Lunn, J. E. et al. Sugar-induced increases in trehalose 6-phosphate are correlated with redox activation of ADPglucose pyrophosphorylase and higher rates of starch synthesis in Arabidopsis thaliana. Biochem. J. 397, 139–148 (2006).
Arrivault, S. et al. Use of reverse‐phase liquid chromatography, linked to tandem mass spectrometry, to profile the Calvin cycle and other metabolic intermediates in Arabidopsis rosettes at different carbon dioxide concentrations. Plant J. 59, 826–839 (2009).
Smith, A. M., Kruger, N. J. & Lunn, J. E. Source of sugar nucleotides for starch and cellulose synthesis. Proc. Natl Acad. Sci. USA 109, E776 (2012).
Winter, D. et al. An “Electronic Fluorescent Pictograph” browser for exploring and analyzing large-scale biological data sets. PLoS ONE 2, e718 (2007).
Waese, J. et al. ePlant: visualizing and exploring multiple levels of data for hypothesis generation in plant biology. Plant Cell 29, 1806–1821 (2017).
Fallahi, H. et al. Localization of sucrose synthase in developing seed and siliques of Arabidopsis thaliana reveals diverse roles for SUS during development. J. Exp. Bot. 59, 3283–3295 (2008).
Dancer, J. E. & Ap Rees, T. Relationship between pyrophosphate: fructose-6-phosphate 1-phosphotransferase, sucrose breakdown, and respiration. J. Plant Physiol. 135, 197–206 (1989).
Xing, H.-L. et al. A CRISPR/Cas9 toolkit for multiplex genome editing in plants. BMC Plant Biol. 14, 327 (2014).
Wang, Z.-P. et al. Egg cell-specific promoter-controlled CRISPR/Cas9 efficiently generates homozygous mutants for multiple target genes in Arabidopsis in a single generation. Genome Biol. 16, 144 (2015).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium‐mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Gibon, Y. et al. A robot-based platform to measure multiple enzyme activities in Arabidopsis using a set of cycling assays: comparison of changes of enzyme activities and transcript levels during diurnal cycles and in prolonged darkness. Plant Cell 16, 3304–3325 (2004).
Figueroa, C. M. et al. Trehalose 6–phosphate coordinates organic and amino acid metabolism with carbon availability. Plant J. 85, 410–423 (2016).
Stitt, M., Lilley, R. M., Gerhardt, R. & Heldt, H. W. Metabolite levels in specific cells and subcellular compartments of plant-leaves. Methods Enzymol. 174, 518–552 (1989).
Stitt, M., Sulpice, R. & Keurentjes, J. Metabolic networks: how to identify key components in the regulation of metabolism and growth. Plant Physiol. 152, 428–444 (2010).
Acknowledgements
We thank C. Abel for help with plant cultivation. This work was supported by a PhD fellowship (to M.M.F.F.F.) from the International Max Planck Research School ‘Primary Metabolism and Plant Growth’, the European Commission FP7 collaborative project TiMet (Contract No. 245143 to A.M.S. and M.S.), the Swedish Research Council for Sustainable Development (Formas) (project grant 2016-01322 to T.N.) and the Max Planck Society (H.I., S.A., R.F., M.S. and J.E.L.).
Funding
Open access funding provided by Max Planck Society.
Author information
Authors and Affiliations
Contributions
W.W. generated the sextuple sus mutants and performed the morphological phenotyping. M.M.F.F.F. performed the harvesting trials and, with H.I., collected samples for metabolite analysis. M.M.F.F.F., S.A., H.I. and R.F. measured metabolites. S.A. performed the 13CO2 labelling experiment. H.I. measured SUS activity. T.N., M.S., A.M.S. and J.E.L. conceived the study. J.E.L. and T.N. drafted the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Nicholas Kruger and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–6 and Tables 2–4.
Supplementary Table 1
13C-labelling data from wild-type Arabidopsis Col-0 leaves.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fünfgeld, M.M.F.F., Wang, W., Ishihara, H. et al. Sucrose synthases are not involved in starch synthesis in Arabidopsis leaves. Nat. Plants 8, 574–582 (2022). https://doi.org/10.1038/s41477-022-01140-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-022-01140-y
This article is cited by
-
Overexpression of ZmSUS1 increased drought resistance of maize (Zea mays L.) by regulating sucrose metabolism and soluble sugar content
Planta (2024)
-
Genetic evidence for functions of Chloroplast CA in Pyropia yezoensis: decreased CCM but increased starch accumulation
Advanced Biotechnology (2024)
-
Recent finding on sucrose synthase research: not the only key for starch and cellulose synthesis
Physiology and Molecular Biology of Plants (2023)