Abstract
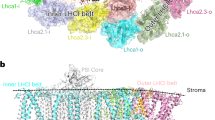
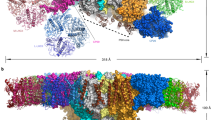
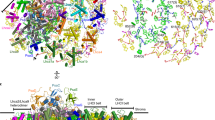
In green algae and plants, state transitions serve as a short-term light-acclimation process in the regulation of the light-harvesting capacity of photosystems I and II (PSI and PSII, respectively). During the process, a portion of light-harvesting complex II (LHCII) is phosphorylated, dissociated from PSII and binds with PSI to form the supercomplex PSI–LHCI–LHCII. Here, we report high-resolution structures of PSI–LHCI–LHCII from Chlamydomonas reinhardtii, revealing the mechanism of assembly between the PSI–LHCI complex and two phosphorylated LHCII trimers containing all four types of LhcbM protein. Two specific LhcbM isoforms, namely LhcbM1 and LhcbM5, directly interact with the PSI core through their phosphorylated amino terminal regions. Furthermore, biochemical and functional studies on mutant strains lacking either LhcbM1 or LhcbM5 indicate that only LhcbM5 is indispensable in supercomplex formation. The results unravel the specific interactions and potential excitation energy transfer routes between green algal PSI and two phosphorylated LHCIIs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The atomic coordinates of the CrPSI–LHCI–LHCII supercomplex have been deposited in the Protein Data Bank with accession codes 7DZ7 (native supercomplex from mutant pph1;pbcp) and 7DZ8 (supercomplex from mutant ΔLhcbM1). The cryo-EM maps of the native and ΔLhcbM1 mutant supercomplexes have been deposited in the Electron Microscopy Data Bank with accession codes EMD-30925 and EMD-30926, respectively. In addition, the atomic coordinates and locally refined cryo-EM maps of LHCII trimers have been deposited in the Protein Data Bank and the Electron Microscopy Data Bank with accession codes 7E0J and EMD-30934 for LHCII-1 in the native supercomplex, 7E0K and EMD-30935 for LHCII-2 in the native supercomplex, 7E0H and EMD-30932 for LHCII-1 in the supercomplex from the ΔLhcbM1 mutant and 7E0I and EMD-30933 for LHCII-2 in the supercomplex from the ΔLhcbM1 mutant. Source data are provided with this paper. All other data generated or analysed are available from the corresponding authors on reasonable request.
References
Dekker, J. P. & Boekema, E. J. Supramolecular organization of thylakoid membrane proteins in green plants. Biochim. Biophys. Acta 1706, 12–39 (2005).
Croce, R. & van Amerongen, H. Light-harvesting and structural organization of photosystem II: from individual complexes to thylakoid membrane. J. Photochem. Photobiol. B 104, 142–153 (2011).
Pan, X., Cao, P., Su, X., Liu, Z. & Li, M. Structural analysis and comparison of light-harvesting complexes I and II. Biochim. Biophys. Acta Bioenerg. 1861, 148038 (2020).
Elrad, D., Niyogi, K. K. & Grossman, A. R. A major light-harvesting polypeptide of photosystem II functions in thermal dissipation. Plant Cell 14, 1801–1816 (2002).
Stauber, E. J. et al. Proteomics of Chlamydomonas reinhardtii light-harvesting proteins. Eukaryot. Cell 2, 978–994 (2003).
Tokutsu, R., Kato, N., Bui, K. H., Ishikawa, T. & Minagawa, J. Revisiting the supramolecular organization of photosystem II in Chlamydomonas reinhardtii. J. Biol. Chem. 287, 31574–31581 (2012).
Ozawa, S. I. et al. Configuration of ten light-harvesting chlorophyll a/b complex I subunits in Chlamydomonas reinhardtii photosystem I. Plant Physiol. 178, 583–595 (2018).
Kubota-Kawai, H. et al. Ten antenna proteins are associated with the core in the supramolecular organization of the photosystem I supercomplex in Chlamydomonas reinhardtii. J. Biol. Chem. 294, 4304–4314 (2019).
Minagawa, J. & Takahashi, Y. Structure, function and assembly of photosystem II and its light-harvesting proteins. Photosynth. Res. 82, 241–263 (2004).
Takahashi, H., Iwai, M., Takahashi, Y. & Minagawa, J. Identification of the mobile light-harvesting complex II polypeptides for state transitions in Chlamydomonas reinhardtii. Proc. Natl Acad. Sci. USA 103, 477–482 (2006).
Natali, A. & Croce, R. Characterization of the major light-harvesting complexes (LHCBM) of the green alga Chlamydomonas reinhardtii. PLoS ONE 10, e0119211 (2015).
Minagawa, J. State transitions—the molecular remodeling of photosynthetic supercomplexes that controls energy flow in the chloroplast. Biochimica Biophysica Acta Bioenerg. 1807, 897–905 (2011).
Rochaix, J. D. Role of thylakoid protein kinases in photosynthetic acclimation. FEBS Lett. 581, 2768–2775 (2007).
Cariti, F. et al. Regulation of light harvesting in Chlamydomonas reinhardtii two protein phosphatases are involved in state transitions. Plant Physiol. 183, 1749–1764 (2020).
Delosme, R., Olive, J. & Wollman, F. A. Changes in light energy distribution upon state transitions: an in vivo photoacoustic study of the wild type and photosynthesis mutants from Chlamydomonas reinhardtii. Biochimica Biophysica Acta Bioenerg. 1273, 150–158 (1996).
Nawrocki, W. J., Santabarbara, S., Mosebach, L., Wollman, F. A. & Rappaport, F. State transitions redistribute rather than dissipate energy between the two photosystems in Chlamydomonas. Nat. Plants 2, 16031 (2016).
Nagy, G. et al. Chloroplast remodeling during state transitions in Chlamydomonas reinhardtii as revealed by noninvasive techniques in vivo. Proc. Natl Acad. Sci. USA 111, 5042–5047 (2014).
Unlu, C., Drop, B., Croce, R. & van Amerongen, H. State transitions in Chlamydomonas reinhardtii strongly modulate the functional size of photosystem II but not of photosystem I. Proc. Natl Acad. Sci. USA 111, 3460–3465 (2014).
Ünlü, C., Polukhina, I. & van Amerongen, H. Origin of pronounced differences in 77 K fluorescence of the green alga Chlamydomonas reinhardtii in state 1 and 2. Eur. Biophys. J. 45, 209–217 (2015).
Finazzi, G. et al. Involvement of state transitions in the switch between linear and cyclic electron flow in Chlamydomonas reinhardtii. EMBO Rep. 3, 280–285 (2002).
Takahashi, H., Clowez, S., Wollman, F. A., Vallon, O. & Rappaport, F. Cyclic electron flow is redox-controlled but independent of state transition. Nat. Commun. 4, 1954 (2013).
Pan, X. et al. Structure of the maize photosystem I supercomplex with light-harvesting complexes I and II. Science 360, 1109–1113 (2018).
Kargul, J. et al. Light-harvesting complex II protein CP29 binds to photosystem I of Chlamydomonas reinhardtii under state 2 conditions. FEBS J. 272, 4797–4806 (2005).
Drop, B., Yadav, K. N. S., Boekema, E. J. & Croce, R. Consequences of state transitions on the structural and functional organization of photosystem I in the green alga Chlamydomonas reinhardtii. Plant J. 78, 181–191 (2014).
Burton-Smith, R. N. et al. Structural determination of the large photosystem II-light-harvesting complex II supercomplex of Chlamydomonas reinhardtii using nonionic amphipol. J. Biol. Chem. 294, 15003–15013 (2019).
Ferrante, P., Ballottari, M., Bonente, G., Giuliano, G. & Bassi, R. LHCBM1 and LHCBM2/7 polypeptides, components of major LHCII complex, have distinct functional roles in photosynthetic antenna system of Chlamydomonas reinhardtii. J. Biol. Chem. 287, 16276–16288 (2012).
Lemeille, S., Turkina, M. V., Vener, A. V. & Rochaix, J. D. Stt7-dependent phosphorylation during state transitions in the green alga Chlamydomonas reinhardtii. Mol. Cell. Proteomics 9, 1281–1295 (2010).
Su, X. et al. Antenna arrangement and energy transfer pathways of a green algal photosystem-I–LHCI supercomplex. Nat. Plants 5, 273–281 (2019).
Suga, M. et al. Structure of the green algal photosystem I supercomplex with a decameric light-harvesting complex I. Nat. Plants 5, 626–636 (2019).
Girolomoni, L. et al. The function of LHCBM4/6/8 antenna proteins in Chlamydomonas reinhardtii. J. Exp. Bot. 68, 628–642 (2017).
Tokutsu, R., Iwai, M. & Minagawa, J. CP29, a monomeric light-harvesting complex II protein, is essential for state transitions in Chlamydomonas reinhardtii. J. Biol. Chem. 284, 7777–7782 (2009).
Iwai, M. et al. Isolation of the elusive supercomplex that drives cyclic electron flow in photosynthesis. Nature 464, 1210–1213 (2010).
Takahashi, H., Okamuro, A., Minagawa, J. & Takahashi, Y. Biochemical characterization of photosystem I-associated light-harvesting complexes I and II isolated from state 2 cells of Chlamydomonas reinhardtii. Plant Cell Physiol. 55, 1437–1449 (2014).
Sheng, X. et al. Structural insight into light harvesting for photosystem II in green algae. Nat. Plants 5, 1320–1330 (2019).
Chen, M. et al. Distinct structural modulation of photosystem I and lipid environment stabilizes its tetrameric assembly. Nat. Plants 6, 314–320 (2020).
Muhleip, A. et al. ATP synthase hexamer assemblies shape cristae of Toxoplasma mitochondria. Nat. Commun. 12, 120 (2021).
Nakane, T., Kimanius, D., Lindahl, E. & Scheres, S. H. W. Characterisation of molecular motions in cryo-EM single-particle data by multi-body refinement in RELION. eLife 7, e36861 (2018).
Liu, Z. F. et al. Crystal structure of spinach major light-harvesting complex at 2.72 Å resolution. Nature 428, 287–292 (2004).
Paulsen, H., Finkenzeller, B. & Kuhlein, N. Pigments induce folding of light-harvesting chlorophyll alpha/beta-binding protein. Eur. J. Biochem. 215, 809–816 (1993).
Croce, R., Weiss, S. & Bassi, R. Carotenoid-binding sites of the major light-harvesting complex II of higher plants. J. Biol. Chem. 274, 29613–29623 (1999).
Novoderezhkin, V. I., Palacios, M. A., van Amerongen, H. & van Grondelle, R. Excitation dynamics in the LHCII complex of higher plants: modeling based on the 2.72 angstrom crystal structure. J. Phys. Chem. B 109, 10493–10504 (2005).
Le Quiniou, C., van Oort, B., Drop, B., van Stokkum, I. H. M. & Croce, R. The high efficiency of photosystem I in the green alga Chlamydomonas reinhardtii is maintained after the antenna size Is substantially increased by the association of light-harvesting complexes II. J. Biol. Chem. 290, 30587–30595 (2015).
Kim, E., Kawakami, K., Sato, R., Ishii, A. & Minagawa, J. Photoprotective capabilities of light-harvesting complex II trimers in the green alga Chlamydomonas reinhardtii. J. Phys. Chem. Lett. 11, 7755–7761 (2020).
Shen, L. et al. Structure of a C2S2M2N2-type PSII-LHCII supercomplex from the green alga Chlamydomonas reinhardtii. Proc. Natl Acad. Sci. USA 116, 21246–21255 (2019).
Huang, Z. et al. Structure of photosystem I-LHCI-LHCII from the green alga Chlamydomonas reinhardtii in state 2. Nat. Commun. 12, 1100 (2021).
Tokutsu, R., Fujimura-Kamada, K., Yamasaki, T., Matsuo, T. & Minagawa, J. Isolation of photoprotective signal transduction mutants by systematic bioluminescence screening in Chlamydomonas reinhardtii. Sci. Rep. 9, 2820 (2019).
Gorman, D. S. & Levine, R. P. Cytochrome f and plastocyanin: their sequence in the photosynthetic electron transport chain of Chlamydomonas reinhardtii. Proc. Natl Acad. Sci. USA 54, 1665–1669 (1965).
Allorent, G. et al. A dual strategy to cope with high light in Chlamydomonas reinhardtii. Plant Cell 25, 545–557 (2013).
Chua, N. H. & Bennoun, P. Thylakoid membrane polypeptides of Chlamydomonas reinhardtii – wild-type and mutant strains deficient in photosystem 2 reaction center. Proc. Natl Acad. Sci. USA 72, 2175–2179 (1975).
Watanabe, A., Kim, E., Burton-Smith, R. N., Tokutsu, R. & Minagawa, J. Amphipol-assisted purification method for the highly active and stable photosystem II supercomplex of Chlamydomonas reinhardtii. FEBS Lett. 593, 1072–1079 (2019).
Wei, X. et al. Structure of spinach photosystem II–LHCII supercomplex at 3.2 Å resolution. Nature 534, 69–74 (2016).
Farber, A., Young, A. J., Ruban, A. V., Horton, P. & Jahns, P. Dynamics of xanthophyll-cycle activity in different antenna subcomplexes in the photosynthetic membranes of higher plants (the relationship between zeaxanthin conversion and nonphotochemical fluorescence quenching). Plant Physiol. 115, 1609–1618 (1997).
Shevchenko, A., Tomas, H., Havlis, J., Olsen, J. V. & Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 1, 2856–2860 (2006).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Wu, C., Huang, X., Cheng, J., Zhu, D. & Zhang, X. High-quality, high-throughput cryo-electron microscopy data collection via beam tilt and astigmatism-free beam-image shift. J. Struct. Biol. 208, 107396 (2019).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Pettersen, E. F. et al. UCSFChimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D. Biol. Crystallogr. 66, 12–21 (2010).
Elrad, D. & Grossman, A. R. A genome’s-eye view of the light-harvesting polypeptides of Chlamydomonas reinhardtii. Curr. Genet. 45, 61–75 (2004).
Acknowledgements
We thank M. Goldschmidt-Clermont at the University of Geneva for the kind gift of the pph1;pbcp strain and insightful discussions; L. H. Chen, X. J. Huang, B. L. Zhu and F. Sun at the Centre for Biological Imaging (CAS) for support with cryo-EM data collection; X. Sheng, D. F. Song and X. Z. Zhang for discussions on cryo-EM data-processing methods; and X. B. Liang for assistance with sample preparation, data collection and storage. We thank K. Fujimura-Kamada and T. Kadowaki for conducting genetical crossing and PCR analysis, and for valuable discussion. We thank T. Mori and Y. Makino for providing technical assistance with LC–MS/MS analysis. We thank A. Watanabe and A. Ishi for technical assistance with supercomplex isolations, and E. Kim for valuable discussions. The project is funded by the National Key R&D Programme of China (no. 2017YFA0503702 to Z.L. and M.L.), the Strategic Priority Research Programme of CAS (nos. XDB27020106 to M.L. and XDB37020101 to Z.L.), the National Natural Science Foundation of China (nos. 31925024 to Z.L. and 31930064 to M.L.), the Basic Frontier Science Research Programme of CAS (no. ZDBS-LY-SM003 to Z.L.) and Grant-in-Aid from Japan Society for the Promotion of Science KAKENHI (nos. 16H06553 to J.M. and JP15H05599 to R.T.). This work was also supported by Functional Genomics Facility, NIBB Core Research Facilities and by Model Plant Research Facility, NIBB Bioresource Centre and the Cooperative Study Programme of National Institute for Physiological Sciences.
Author information
Authors and Affiliations
Contributions
R.T., Z.L., J.M. and M.L. conceived and coordinated the project. X.P. and A.L. collected and processed cryo-EM data. X.P. built and refined the structural models. R.T. performed biochemical and spectroscopic characterization of the PSI–LHCI(–LHCII) supercomplexes from WT and LhcbM mutants. A.L. prepared cryo-EM grids for the supercomplex sample from the pph1;pbcp mutant. C.S. and K.M. prepared cryo-EM grids for the supercomplex samples from WT and the ΔLhcbM1 mutant. T.Y. generated the ΔLhcbM5 mutant. K.T. performed qT quenching measurements. X.P., R.T., A.L., Z.L., J.M. and M.L. analysed the data and wrote the manuscript. All authors discussed and commented on the results and the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Alexey Amunts, Jean-David Rochaix and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Sample preparation and characterization of native CrPSI-LHCI-LHCII supercomplex from the pph1;pbcp mutant strain.
a, Sucrose density gradient of solubilized thylakoid membranes. Each band shown is labeled. b, SDS-PAGE analysis of thylakoids (Lane 1) and the purified PSI-LHCI-LHCII supercomplex (Lane 2). The protein composition of each Coomassie band was indicated based on the mass spectrometry and proteomics data analysis. The bands corresponding to PSII subunits are indicated by asterisk. Representative image selected from 3 biological repeats was shown. c, Room-temperature absorption spectra of PSI-LHCI and PSI-LHCI-LHCII samples. The PSI-LHCI-LHCII sample showed higher peaks around 470 and 650 nm (indicated by arrows), demonstrating that the Chl b (from LHCII) content of this fraction is higher than that of PSI-LHCI complex. The spectra were normalized to the maximum in the red region. d, HPLC analysis of pigment content in PSI-LHCI and PSI-LHCI-LHCII samples. Based on the characteristic absorption spectrum of each peak fraction, the six major pigment peaks separated from the sample are identified as loroxanthin/neoxanthin (Lor/Neo), violaxanthin (Vio), lutein (Lut), chlorophyll b (Chl b), chlorophyll a (Chl a) and β-carotene (BCR).
Extended Data Fig. 2 Single particle cryo-EM analysis and evaluation of CrPSI-LHCI-LHCII supercomplex from the pph1;pbcp mutant strain.
a, Single particle cryo-EM data processing procedure. The class corresponding to PSII-LHCII complex is framed by black box and indicated by asterisk. For the second dataset (shown in right), the PSII-LHCII complexes were excluded during particle picking. b, The gold standard Fourier shell correlation (FSC) curves of the final density map with criterion of 0.143. c, The gold standard Fourier shell correlation (FSC) curves of the refined models versus final maps with criterion of 0.5. d, Angular distribution of particles included in the final 3D reconstruction. e, Local resolution of the cryo-EM map estimated by ResMap.
Extended Data Fig. 3
Cryo-EM data collection, refinement and validation statistics of CrPSI-LHCI-LHCII structures.
Extended Data Fig. 4 Identification of each LhcbM protein in the LHCII-1 structure of CrPSI-LHCI-LHCII supercomplex from the pph1;pbcp mutant strain.
a–c, Map features of characteristic residues and the corresponding sequence (residues are numbered according to LhcbM1) of LhcbM1 (a), LhcbM2/7 (b) and LhcbM3/M4 (c). The resolution and contour level are 3.13 Å and 1.42 rmsd for the masked map of LHCII-1. Unique residues used for identification are indicated by red arrows. a, LhcbM1 possesses unique N-terminal residues (RRt, underlined by a red line) with well-defined density. Moreover, Phe106 shows clear density for its side-chain, which excludes the possibility of all other LhcbM proteins as they have a Thr at the corresponding position. b, LhcbM2/7 contain unique residues Trp32 and Thr98 (residues 41 and 106 in LhcbM1), whereas the corresponding residue of Trp32 is a Phe in Type I and Type II isoforms, and the corresponding residue of Thr98 is a Phe in Type IV isoform, therefore excluding the possibility of all other LhcbM proteins. The density of the Phe147 (residue 155 in LhcbM1) side chain was used to further verify the assignment of LhcbM2/7. c, Type I LhcbM proteins have Phe40 and Trp48 (residues 41 and 49 in LhcbM1) in the N-terminal regions, whereas Type III and Type IV have Trp and Phe in the corresponding positions, and can therefore be excluded. In addition, Phe117 shows clear density for its side chain, which excludes the possibility of Type II LhcbM protein, as LhcbM5 has an Ile at the corresponding position. The well-defined densities of Met213, Met216 and Thr242 further exclude LhcbM6, LhcbM9, LhcbM8 and LhcbM9.
Extended Data Fig. 5 Identification of each LhcbM protein in the LHCII-2 structure of CrPSI-LHCI-LHCII supercomplex from the pph1;pbcp mutant strain.
a–c, Map features of characteristic residues and the corresponding sequence (residues are numbered according to LhcbM1) of LhcbM5 (a), LhcbM2/7 (b) and LhcbM3/M4 (c). The resolution and contour level are 3.09 Å and 1.11 rmsd for the masked map of LHCII-2. Unique residues used for identification are indicated by red arrows. a, LhcbM5 possesses the longest N-terminal region among all LhcbM proteins (underlined by a red line), which shows clear densities in the map. The assignment of LhcbM5 was further verified by the densities of specific residues Phe197 and Phe254 (residue 242 in LhcbM1). The former is absent, while the latter is a residue with small side-chain (Gly or Thr) in all other LhcbM proteins. b, LhcbM2/7 contain unique residues Trp32 and Thr98 (residues 41 and 106 in LhcbM1), whereas the corresponding residue of Trp32 is a Phe in Type I and Type II isoforms, the corresponding residue of Thr98 is a Phe in Type IV isoform, therefore exclude the possibility of all other LhcbM proteins. The density of Phe147 (residues 155 in LhcbM1) side-chain was used to further verify the assignment of LhcbM2/7. c, Type I LhcbM proteins have Phe40 and Trp48 (residues 41 and 49 in LhcbM1) in the N-terminal regions, whereas Type III and Type IV have Trp and Phe in the corresponding positions, therefore can be excluded. In addition, Phe117 shows clear density for its side-chain, which excludes the possibility of Type II LhcbM protein as LhcbM5 has a Ile at the corresponding position. The well-defined densities of Met213, Met216 and Thr242 further exclude LhcbM6, LhcbM9, and LhcbM8 and LhcbM9.
Extended Data Fig. 6 Multi-body refinement of the CrPSI-LHCI-LHCII supercomplex from the pph1:pbcp mutant strain.
a, The three bodies corresponding to LHCII-1, LHCII-2 and PSI-10LHCI are defined by the transparent masks in magenta, cyan and yellow, respectively. b, The contributions of all 18 eigenvectors to the variance. c-e, The flexibility of LHCII-1 and LHCII-2 relative to PSI-LHCI complex in the principal components along the top three eigenvectors (#1-3). The models shown are fitted into the bin 1 and bin 10 maps in each component and then superposed on the PSI-LHCI complex region. The transparent red circles are the estimated anchor points for LHCII-1 and LHCII-2 during the rotational or pivoting movement. In (c) and (e), the views on the left are from stromal side. In (d), the view on the left is from luminal side to exhibit the anchor point. The eye symbols in the left parts of (c-e) indicate the viewing angles for the side views shown on the right.
Extended Data Fig. 7 Single particle cryo-EM data processing of CrPSI-LHCI-LHCII supercomplex from the ΔLhcbM1 mutant strain.
a, Single particle cryo-EM data processing procedure. b, The gold standard Fourier shell correlation (FSC) curves of the final density map with criterion of 0.143. c, The gold standard Fourier shell correlation (FSC) curves of the refined models versus final maps with criterion of 0.5. d, Angular distribution of particles included in the final 3D reconstruction. e, Local resolution of the cryo-EM map estimated by ResMap.
Extended Data Fig. 8 Identification of a Type I isoform replacing LhcbM1 in the structure of PSI-LHCI-LHCII supercomplex from the ΔLhcbM1 mutant strain.
Map features of characteristic residues and the corresponding sequence (residues are numbered according to LhcbM1). The density of the LHCII-1 region of PSI-LHCI-LHCII map is shown, with local resolution of approximately 3.75 Å and contour level of 1.31 rmsd. Type I LhcbM proteins have a Trp48 (residue 49 in LhcbM1) in the N-terminal regions, whereas Type III and Type IV have a Phe in the corresponding position, therefore can be excluded. Residue Tyr52 (residue 53 in LhcbM1) shows clear density for its side-chain, which excludes the possibility of Type II LhcbM protein as LhcbM5 has a Leu at the corresponding position. The map features of Tyr181 further verify the assignment of Type I isoform. Unique residues used for identification are indicated by red arrows.
Extended Data Fig. 9 Comparison of PSI-LHCI-LHCII supercomplex from pph1;pbcp and ΔLhcbM1 mutant strains.
a, Structural comparison of PSI-LHCI-LHCII supercomplex from ΔLhcbM1 mutant and in the native form (from the pph1;pbcp mutant strain), superposed on PsaA. The native supercomplex is shown in white, with LhcbM1 of LHCII-1 highlighted in lime-green. The supercomplex from ΔLhcbM1 mutant is colored orange for the PSI-LHCI moiety and the newly incorporated LhcbM3 of LHCII-1, while other LhcbM proteins are colored the same as in Fig. 1a. The N-terminal tails of LhcbM1 (lime-green) and the newly incorporated LhcbM3 (orange) in the two structures are indicated by arrows in a red square. b,c, Side view of structural comparison of two LHCII trimers in the CrPSI-LHCI-LHCII supercomplex from ΔLhcbM1 mutant and in native form, viewed from the periphery of two trimers (b) and from the PSI-LHCII interface (c). The color codes are the same as in (a). The outmost monomer in LHCII-1 shows the largest shift towards lumen as shown by the arrow in (b).
Extended Data Fig. 10 The variation of chlorophyll-chlorophyll distances at the interface between LHCII-1/LHCII-2 and PSI-LHCI in the first component of the rigid body motion.
The models correspond to the bin1 and bin10 maps of the principal component along eigenvector #1 of the multi-body refinement result. a,b, The interfacial chlorophyll pairs at the stromal layer in bin1 (a) and bin10 (b) states. c,d, The interfacial chlorophyll pairs at the luminal layer in bin1 (c) and bin10 (d) states. The numbers labeled nearby the dash lines indicate the Mg-Mg distances (Å) between two adjacent chlorophylls at the interfaces between LHCII-1/LHCII-2 and PSI-LHCI. The variation of the interfacial chlorophyll-chlorophyll distances in the top three motion components (Eigenvectors #1-3) are also summarized in Supplementary Table 2.
Supplementary information
Supplementary Information
Supplementary Figs. 1–4, Tables 1 and 2, Video 1 and additional Figs. 1 and 2 (source data for Supplementary Fig. 3b,c).
Supplementary Video 1
Analysis of the multi-body refinement result reveals the motion of two LHCII trimers relative to PSI–LHCI.
Source data
Source Data Fig. 4
Unprocessed gel in Fig. 4b.
Source Data Extended Data Fig. 1
Unprocessed gel in Extended Data Fig. 1b.
Rights and permissions
About this article
Cite this article
Pan, X., Tokutsu, R., Li, A. et al. Structural basis of LhcbM5-mediated state transitions in green algae. Nat. Plants 7, 1119–1131 (2021). https://doi.org/10.1038/s41477-021-00960-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-021-00960-8
This article is cited by
-
Macroscale structural changes of thylakoid architecture during high light acclimation in Chlamydomonas reinhardtii
Photosynthesis Research (2024)
-
Structural insights into a unique PSI–LHCI–LHCII–Lhcb9 supercomplex from moss Physcomitrium patens
Nature Plants (2023)
-
Structural insights into the assembly and energy transfer of the Lhcb9-dependent photosystem I from moss Physcomitrium patens
Nature Plants (2023)
-
Uphill energy transfer mechanism for photosynthesis in an Antarctic alga
Nature Communications (2023)
-
Reversible protein phosphorylation in higher plants: focus on state transitions
Biophysical Reviews (2023)