Abstract
Plants, like other multicellular lifeforms, are colonized by microorganisms. How plants respond to their microbiota is currently not well understood. We used a phylogenetically diverse set of 39 endogenous bacterial strains from Arabidopsis thaliana leaves to assess host transcriptional and metabolic adaptations to bacterial encounters. We identified a molecular response, which we termed the general non-self response (GNSR) that involves the expression of a core set of 24 genes. The GNSR genes are not only consistently induced by the presence of most strains, they also comprise the most differentially regulated genes across treatments and are predictive of a hierarchical transcriptional reprogramming beyond the GNSR. Using a complementary untargeted metabolomics approach we link the GNSR to the tryptophan-derived secondary metabolism, highlighting the importance of small molecules in plant–microbe interactions. We demonstrate that several of the GNSR genes are required for resistance against the bacterial pathogen Pseudomonas syringae. Our results suggest that the GNSR constitutes a defence adaptation strategy that is consistently elicited by diverse strains from various phyla, contributes to host protection and involves secondary metabolism.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Transcriptomics raw data are available at the European Bioinformatics Institute (EBI) under the following identifier: PRJEB40488. Metabolomics raw data, together with relevant metadata, are available at Metabolights79 under the following identifier: MTBLS2522. Source data are provided with this paper.
Code availability
The code used in the analysis of the data can be found in the following repositories. The scripts for RNA-seq raw data analysis according to the standard operating procedure of the FGCZ is available at their GitHub repository: https://github.com/uzh/ezRun. The code used for metabolomics raw data analysis based on the in-house pipeline can be found at: https://gitlab.ethz.ch/emzed2-plant-metabolomics-workflows. The scripts for all major downstream data processing/figure generation are available at the project’s GitLab repository: https://gitlab.ethz.ch/emzed2-plant-metabolomics-workflows.
References
Vorholt, J. A. Microbial life in the phyllosphere. Nat. Rev. Microbiol. 10, 828–840 (2012).
Lindow, S. E. & Brandl, M. T. Microbiology of the phyllosphere. Appl. Environ. Microbiol. 69, 1875–1883 (2003).
Müller, D. B., Vogel, C., Bai, Y. & Vorholt, J. A. The plant microbiota: systems-level insights and perspectives. Annu. Rev. Genet 50, 211–234 (2016).
Hacquard, S., Spaepen, S., Garrido-Oter, R. & Schulze-Lefert, P. Interplay between innate immunity and the plant microbiota. Annu. Rev. Phytopathol. 55, 565–589 (2017).
Ditt, R. F. et al. The Arabidopsis thaliana transcriptome in response to Agrobacterium tumefaciens. Mol. Plant Microbe Interact. 19, 665–681 (2006).
Thilmony, R., Underwood, W. & He, S. Y. Genome-wide transcriptional analysis of the Arabidopsis thaliana interaction with the plant pathogen Pseudomonas syringae pv. tomato DC3000 and the human pathogen Escherichia coli O157:H7. Plant J. 46, 34–53 (2006).
Verhagen, B. W. et al. The transcriptome of rhizobacteria-induced systemic resistance in Arabidopsis. Mol. Plant Microbe Interact. 17, 895–908 (2004).
van de Mortel, J. E. et al. Metabolic and transcriptomic changes induced in Arabidopsis by the rhizobacterium Pseudomonas fluorescens SS101. Plant Physiol. 160, 2173–2188 (2012).
Vogel, C., Bodenhausen, N., Gruissem, W. & Vorholt, J. A. The Arabidopsis leaf transcriptome reveals distinct but also overlapping responses to colonization by phyllosphere commensals and pathogen infection with impact on plant health. New Phytol. 212, 192–207 (2016).
D’Auria, J. C. & Gershenzon, J. The secondary metabolism of Arabidopsis thaliana: growing like a weed. Curr. Opin. Plant Biol. 8, 308–316 (2005).
Sato, F. in Encyclopedia of Life Sciences (ELS, 2014); https://doi.org/10.1002/9780470015902.a0001812.pub2
Huang, A. C. et al. A specialized metabolic network selectively modulates Arabidopsis root microbiota. Science 364, eaau6389 (2019).
Voges, M., Bai, Y., Schulze-Lefert, P. & Sattely, E. S. Plant-derived coumarins shape the composition of an Arabidopsis synthetic root microbiome. Proc. Natl Acad. Sci. USA 116, 12558–12565 (2019).
Badri, D. V., Chaparro, J. M., Zhang, R., Shen, Q. & Vivanco, J. M. Application of natural blends of phytochemicals derived from the root exudates of Arabidopsis to the soil reveal that phenolic-related compounds predominantly modulate the soil microbiome. J. Biol. Chem. 288, 4502–4512 (2013).
Pastorczyk, M. et al. The role of CYP71A12 monooxygenase in pathogen-triggered tryptophan metabolism and Arabidopsis immunity. New Phytol. 225, 400–412 (2020).
Clay, N. K., Adio, A. M., Denoux, C., Jander, G. & Ausubel, F. M. Glucosinolate metabolites required for an Arabidopsis innate immune response. Science 323, 95–101 (2009).
Schlaeppi, K., Abou-Mansour, E., Buchala, A. & Mauch, F. Disease resistance of Arabidopsis to Phytophthora brassicae is established by the sequential action of indole glucosinolates and camalexin. Plant J. 62, 840–851 (2010).
Rajniak, J., Barco, B., Clay, N. K. & Sattely, E. S. A new cyanogenic metabolite in Arabidopsis required for inducible pathogen defence. Nature 525, 376–379 (2015).
Nongbri, P. L. et al. Indole-3-acetaldoxime-derived compounds restrict root colonization in the beneficial interaction between Arabidopsis roots and the endophyte Piriformospora indica. Mol. Plant Microbe Interact. 25, 1186–1197 (2012).
Bai, Y. et al. Functional overlap of the Arabidopsis leaf and root microbiota. Nature 528, 364–369 (2015).
Vogel, C., Innerebner, G., Zingg, J., Guder, J. & Vorholt, J. A. Forward genetic in planta screen for identification of plant-protective traits of Sphingomonas sp. strain Fr1 against Pseudomonas syringae DC3000. Appl. Environ. Microbiol. 78, 5529–5535 (2012).
Hruz, T. et al. Genevestigator v3: a reference expression database for the meta-analysis of transcriptomes. Adv. Bioinforma. 2008, 420747 (2008).
Bethke, G. et al. Arabidopsis PECTIN METHYLESTERASEs contribute to immunity against Pseudomonas syringae. Plant Physiol. 164, 1093–1107 (2014).
Maekawa, T., Kracher, B., Vernaldi, S., Ver Loren van Themaat, E. & Schulze-Lefert, P. Conservation of NLR-triggered immunity across plant lineages. Proc. Natl Acad. Sci. USA 109, 20119–20123 (2012).
Liu, S., Kracher, B., Ziegler, J., Birkenbihl, R. P. & Somssich, I. E. Negative regulation of ABA signaling by WRKY33 is critical for Arabidopsis immunity towards Botrytis cinerea 2100. eLife 4, e07295 (2015).
Hacquard, S. et al. Survival trade-offs in plant roots during colonization by closely related beneficial and pathogenic fungi. Nat. Commun. 7, 11362 (2016).
Hiruma, K. et al. Root endophyte Colletotrichum tofieldiae confers plant fitness benefits that are phosphate status dependent. Cell 165, 464–474 (2016).
Stotz, H. U. et al. Role of camalexin, indole glucosinolates, and side chain modification of glucosinolate-derived isothiocyanates in defense of Arabidopsis against Sclerotinia sclerotiorum. Plant J. 67, 81–93 (2011).
Howard, B. E. et al. High-throughput RNA sequencing of Pseudomonas-infected Arabidopsis reveals hidden transcriptome complexity and novel splice variants. PLoS ONE 8, e74183 (2013).
Li, B. et al. Phosphorylation of trihelix transcriptional repressor ASR3 by MAP KINASE4 negatively regulates Arabidopsis immunity. Plant Cell 27, 839–856 (2015).
Zhan, X. et al. An Arabidopsis PWI and RRM motif-containing protein is critical for pre-mRNA splicing and ABA responses. Nat. Commun. 6, 8139 (2015).
Hooper, C. M., Castleden, I. R., Tanz, S. K., Aryamanesh, N. & Millar, A. H. SUBA4: the interactive data analysis centre for Arabidopsis subcellular protein locations. Nucleic Acids Res. 45, D1064–D1074 (2017).
Tanz, S. K. et al. SUBA3: a database for integrating experimentation and prediction to define the SUBcellular location of proteins in Arabidopsis. Nucleic Acids Res. 41, D1185–D1191 (2013).
Hull, A. K., Vij, R. & Celenza, J. L. Arabidopsis cytochrome P450s that catalyze the first step of tryptophan-dependent indole-3-acetic acid biosynthesis. Proc. Natl Acad. Sci. USA 97, 2379–2384 (2000).
Muller, T. M., Bottcher, C. & Glawischnig, E. Dissection of the network of indolic defence compounds in Arabidopsis thaliana by multiple mutant analysis. Phytochemistry 161, 11–20 (2019).
Glawischnig, E. Camalexin. Phytochemistry 68, 401–406 (2007).
Wei, G. & Shirsat, A. H. Extensin over-expression in Arabidopsis limits pathogen invasiveness. Mol. Plant Pathol. 7, 579–592 (2006).
Chassot, C., Nawrath, C. & Metraux, J. P. Cuticular defects lead to full immunity to a major plant pathogen. Plant J. 49, 972–980 (2007).
Yu, Z. et al. The Brassicaceae-specific secreted peptides, STMPs, function in plant growth and pathogen defense. J. Integr. Plant Biol. 62, 403–420 (2020).
Roux, M. et al. The Arabidopsis leucine-rich repeat receptor-like kinases BAK1/SERK3 and BKK1/SERK4 are required for innate immunity to hemibiotrophic and biotrophic pathogens. Plant Cell 23, 2440–2455 (2011).
Teixeira, P. J. P., Colaianni, N. R., Fitzpatrick, C. R. & Dangl, J. L. Beyond pathogens: microbiota interactions with the plant immune system. Curr. Opin. Microbiol. 49, 7–17 (2019).
Lebeis, S. L. The potential for give and take in plant–microbiome relationships. Front. Plant Sci. 5, 287 (2014).
Hu, F. P. & Young, J. M. Biocidal activity in plant pathogenic Acidovorax, Burkholderia, Herbaspirillum, Ralstonia and Xanthomonas spp. J. Appl. Microbiol. 84, 263–271 (1998).
van der Wolf, J. & De Boer, S. H. in Principles of Plant–Microbe Interactions: Microbes for Sustainable Agriculture (ed. Lugtenberg, B.) 65–77 (Springer, 2015).
Bull, C. T. et al. Comprehensive list of names of plant pathogenic bacteria, 1980–2007. J. Plant Pathol. 92, 551–592 (2010).
Shigeto, J. et al. Simultaneously disrupting AtPrx2, AtPrx25 and AtPrx71 alters lignin content and structure in Arabidopsis stem. J. Integr. Plant Biol. 57, 349–356 (2015).
Barth, C. & Jander, G. Arabidopsis myrosinases TGG1 and TGG2 have redundant function in glucosinolate breakdown and insect defense. Plant J. 46, 549–562 (2006).
Muller, T. M. et al. TRANSCRIPTION ACTIVATOR-LIKE EFFECTOR NUCLEASE-mediated generation and metabolic analysis of camalexin-deficient cyp71a12 cyp71a13 double knockout lines. Plant Physiol. 168, 849–858 (2015).
Hara, M., Yatsuzuka, Y., Tabata, K. & Kuboi, T. Exogenously applied isothiocyanates enhance glutathione S-transferase expression in Arabidopsis but act as herbicides at higher concentrations. J. Plant Physiol. 167, 643–649 (2010).
Yu, K. et al. A feedback regulatory loop between G3P and lipid transfer proteins DIR1 and AZI1 mediates azelaic-acid-induced systemic immunity. Cell Rep. 3, 1266–1278 (2013).
Cecchini, N. M., Steffes, K., Schlappi, M. R., Gifford, A. N. & Greenberg, J. T. Arabidopsis AZI1 family proteins mediate signal mobilization for systemic defence priming. Nat. Commun. 6, 7658 (2015).
Bowling, S. A., Clarke, J. D., Liu, Y., Klessig, D. F. & Dong, X. The cpr5 mutant of Arabidopsis expresses both NPR1-dependent and NPR1-independent resistance. Plant Cell 9, 1573–1584 (1997).
Li, X., Clarke, J. D., Zhang, Y. & Dong, X. Activation of an EDS1-mediated R-gene pathway in the snc1 mutant leads to constitutive, NPR1-independent pathogen resistance. Mol. Plant Microbe Interact. 14, 1131–1139 (2001).
Schlaeppi, K. & Bulgarelli, D. The plant microbiome at work. Mol. Plant Microbe Interact. 28, 212–217 (2015).
El-Esawi, M. A., Al-Ghamdi, A. A., Ali, H. M. & Ahmad, M. Overexpression of AtWRKY30 transcription factor enhances heat and drought stress tolerance in wheat (Triticum aestivum L.). Genes 10, 163 (2019).
Scarpeci, T. E., Zanor, M. I., Mueller-Roeber, B. & Valle, E. M. Overexpression of AtWRKY30 enhances abiotic stress tolerance during early growth stages in Arabidopsis thaliana. Plant Mol. Biol. 83, 265–277 (2013).
Schlesier, B., Breton, F. & Mock, H. P. A hydroponic culture system for growing Arabidopsis thaliana plantlets under sterile conditions. Plant Mol. Biol. Rep. 21, 449–456 (2003).
Fan, J., Crooks, C. & Lamb, C. High-throughput quantitative luminescence assay of the growth in planta of Pseudomonas syringae chromosomally tagged with Photorhabdus luminescens luxCDABE. Plant J. 53, 393–399 (2008).
King, E. O., Ward, M. K. & Raney, D. E. Two simple media for the demonstration of pyocyanin and fluorescin. J. Lab. Clin. Med. 44, 301–307 (1954).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinf. 12, 323 (2011).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
McCarthy, D. J., Chen, Y. & Smyth, G. K. Differential expression analysis of multifactor RNA-seq experiments with respect to biological variation. Nucleic Acids Res. 40, 4288–4297 (2012).
Tian, T. et al. agriGO v2.0: a GO analysis toolkit for the agricultural community, 2017 update. Nucleic Acids Res. 45, W122–W129 (2017).
Mulleder, M., Bluemlein, K. & Ralser, M. A high-throughput method for the quantitative determination of free amino acids in Saccharomyces cerevisiae by hydrophilic interaction chromatography–tandem mass spectrometry. Cold Spring Harb. Protoc. 2017, pdb.prot089094 (2017).
Rost, H. L. et al. OpenMS: a flexible open-source software platform for mass spectrometry data analysis. Nat. Methods 13, 741–748 (2016).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Kanehisa, M., Sato, Y., Furumichi, M., Morishima, K. & Tanabe, M. New approach for understanding genome variations in KEGG. Nucleic Acids Res. 47, D590–D595 (2019).
Kanehisa, M. KEGG bioinformatics resource for plant genomics and metabolomics. Methods Mol. Biol. 1374, 55–70 (2016).
Kim, S. et al. PubChem 2019 update: improved access to chemical data. Nucleic Acids Res. 47, D1102–D1109 (2019).
Schlapfer, P. et al. Genome-wide prediction of metabolic enzymes, pathways, and gene clusters in plants. Plant Physiol. 173, 2041–2059 (2017).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Csardi, G. & Nepusz, T. The igraph software package for complex network research. InterJournal 1695, 1–9 (2006).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Oksanen, J. et al. vegan: Community ecology. R package version 2.5-6 (2013).
Bocker, S., Letzel, M. C., Liptak, Z. & Pervukhin, A. SIRIUS: decomposing isotope patterns for metabolite identification. Bioinformatics 25, 218–224 (2009).
Bocker, S. & Duhrkop, K. Fragmentation trees reloaded. J. Cheminform. 8, 5 (2016).
Duhrkop, K., Shen, H., Meusel, M., Rousu, J. & Bocker, S. Searching molecular structure databases with tandem mass spectra using CSI:FingerID. Proc. Natl Acad. Sci. USA 112, 12580–12585 (2015).
Wang, M. et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat. Biotechnol. 34, 828–837 (2016).
Haug, K. et al. MetaboLights: a resource evolving in response to the needs of its scientific community. Nucleic Acids Res. 48, D440–D444 (2020).
Acknowledgements
We thank C. Zipfel, W. Gruissem and C. Sánchez-Rodríguez for helpful discussions and A. Imboden for reproduction of plant lines. We thank E. Sattely for providing the cyp71a12 and fox1 mutant lines. We also thank L. Opitz and C. Aquino Fournier from the Functional Genomics Center Zurich (FGCZ) for their work on the RNA sequencing part of the experiment. This work was funded through a European Research Council advanced grant (PhyMo; no. 668991), ETH Zurich, a grant from the German Research Foundation (DECRyPT, no. SPP2125) and the NCCR Microbiomes, funded by the Swiss National Science Foundation.
Author information
Authors and Affiliations
Contributions
B.A.M., C.M.V. and J.A.V. designed the study. B.A.M. led and conducted the experimental work. P.K. designed the metabolomics pipeline. P.K. and C.M.F carried out omics raw data analysis. B.A.M., C.M.F. and S.S. performed the data analysis. L.H., M.B.-M., B.E. and M.S. contributed to the generation of the plant material for the transcriptomics and metabolomics experiments. S.P. helped with the design of the pathogen assays. B.A.M. and J.A.V. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Alisdair Fernie and Susannah Tringe for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Phenotype of Arabidopsis inoculated with the 39 bacterial strains.
Representative phenotypes of 3-week-old Arabidopsis wild-type plants (Col-0), 9 days after inoculation with individual bacterial strains.
Extended Data Fig. 2 Cluster analysis of DEGs and DAMs of Arabidopsis in response to bacterial treatments.
a-b, Heat-maps of total DEGs (|log2FC|> 1. FDR < 0.01) (a) and DAMs (|log2FC|> 1, FDR < 0.05) (b). Strains and DEGs/ DAMs are clustered using Ward’s method. Top colour bars indicate phylogeny. c-d, Principle component analysis (PCA) plot depicting distances based on gene expression counts (for DEGs) or metabolite areas under the curve (for DAMs). Colours represent bacterial phyla/classes. Selected treatments are annotated with strain name and replicate number. Data from five independent biological replicates (R1-R5), each representing 18–24 plants.
Extended Data Fig. 3 Correlation between phyllosphere colonization and host response intensity.
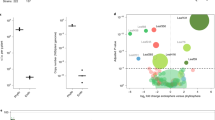
a-c, Linear regression of cfu*gfw-1 against nDEGs or nDAMs and nDEGs against nDAMs in the respective conditions. Axis scaled to log10 for improved readability. Dots are annotated by the strain used in the treatment. Colours represent bacterial phyla/class analogous to Supplementary Fig. 2. Coefficients of variation from the linear regression (R²) value and Spearman’s ranked correlation values (ρ) are provided. Data from five independent biological replicates, each representing 18–24 plants.
Extended Data Fig. 4 Hierarchy heatmap of DAMs.
Sorted heat-maps of total DAMs (log2FC > 1, FDR < 0.05). Conditions were sorted by strains causing the weakest to strongest host response based on the number of DAMs (x axis, left to right) and amount of times a metabolite feature was differentially regulated from most frequent to least frequent (y axis, top to bottom). Top colour bars indicate bacterial phyla/classes analogous to Extended Data Fig. 2. Data from five independent biological replicates, each representing 18–24 plants.
Extended Data Fig. 5 Genevestigator analysis of GNSR genes.
Selected query results for GNSR genes in Genevestigator category ‘Perturbations’. Names adjusted to match names used in this study a, Results for mRNA seq datasets from various experiments (|FC|> 2, p-value < 0.01)) for biotic, chemical, hormonal, nutritional, photoperiod, temperature or other abiotic perturbations. b, Results for micro-array datasets from various experiments (|FC|> 1.5, p-value < 0.001) for biotic, chemical, elicitor, light intensity, nutritional and other abiotic stress perturbations. All p-values were computed by two-sided t-test implemented in limma (for micro-array data) and by Voom’s algorithm (two-sided) for RNAseq data and are adjusted by Benjamini–Hochberg to compute the threshold under which the p-values are considered sufficiently small22 (https://genevestigator.com/userdocs/manual/GENEVESTIGATOR_UserManual.pdf).
Extended Data Fig. 6 Functional enrichment and subcellular location analysis.
Analysis of subcellular locations using SUBA4 prediction scores (left, coloured) and summarized associated GO analysis of GNSR genes based on AgriGO2 functional enrichment (right, grey).
Extended Data Fig. 7 Tryptophan-derived secondary metabolism in Arabidopsis.
Simplified version of the TDSM with the three main branches, important intermediates and enzymes. GNSR genes are highlighted in green, genes homologous to GNSR genes are highlighted in yellow.
Extended Data Fig. 8 Targeted metabolomics on the cyp71a12 mutant.
Heatmap of log2-transformed metabolite fold changes in the cyp71a12 mutant against the respective wild-type conditions. Only changes in 349, 365 and 383 are significant (two-sided t-test, Benjamini–Hochberg adjusted p-value < 0.05). Data from ten technical replicates, each representing 1 plant.
Extended Data Fig. 9 Phyllosphere colonization by bacteria native to the potting soil used in this experiment.
a, Median cfu*gfw-1 with 95% confidence interval of 30 different plants (n = 30) in one experiment. b, Representative picture of bacterial colonies extracted from leaves, grown on R2A agar with 0.5% (v/v) methanol and supplemented with cycloheximide to prevent fungal growth.
Extended Data Fig. 10 Gene expression correlation of GNSR and transcription factor encoding genes.
Network graph showing significant, positive correlations (Spearman correlation, ρ ≥ 0.85, best two correlations) between GNSR genes and transcription factors of the defence-associated MYB and WRKY families. GNSR genes are displayed in red, transcription factor genes in blue. WRKY30 in yellow as it is both a GNSR and a transcription factor gene.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10.
Supplementary Table 1
Genes differentially expressed (|log2FC|> 1, FDR < 0.01) in plants colonized by At-LSPHERE strains
Supplementary Table 2
Metabolite features differentially abundant (|log2FC|> 1, FDR < 0.05) in plants colonized by At-LSPHERE strains
Supplementary Table 3
GO-terms associated with GNSR genes
Supplementary Table 4
Results of linear regression analysis between transcriptional response intensity [nDEGs] and individual gene fold-changes. All genes with adj. R²-values > 0.75
Supplementary Table 5
rho-values of Spearmans ranked correlation between GNSR genes and metabolite features
Supplementary Table 6–16
m/z values of fragment ions obtained during MS2 measurements of GNSR associated compounds
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
About this article
Cite this article
Maier, B.A., Kiefer, P., Field, C.M. et al. A general non-self response as part of plant immunity. Nat. Plants 7, 696–705 (2021). https://doi.org/10.1038/s41477-021-00913-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-021-00913-1
This article is cited by
-
Microbiome homeostasis on rice leaves is regulated by a precursor molecule of lignin biosynthesis
Nature Communications (2024)
-
MAPK Cascades in Plant Microbiota Structure and Functioning
Journal of Microbiology (2024)
-
The Arabidopsis holobiont: a (re)source of insights to understand the amazing world of plant–microbe interactions
Environmental Microbiome (2023)
-
Modules in robust but low-efficiency phyllosphere fungal networks drive saponin accumulation in leaves of different Panax species
Environmental Microbiome (2023)
-
Spatial metatranscriptomics resolves host–bacteria–fungi interactomes
Nature Biotechnology (2023)