Abstract
Auxin controls numerous growth processes in land plants through a gene expression system that modulates ARF transcription factor activity1,2,3. Gene duplications in families encoding auxin response components have generated tremendous complexity in most land plants, and neofunctionalization enabled various unique response outputs during development1,3,4. However, it is unclear what fundamental biochemical principles underlie this complex response system. By studying the minimal system in Marchantia polymorpha, we derive an intuitive and simple model where a single auxin-dependent A-ARF activates gene expression. It is antagonized by an auxin-independent B-ARF that represses common target genes. The expression patterns of both ARF proteins define developmental zones where auxin response is permitted, quantitatively tuned or prevented. This fundamental design probably represents the ancestral system and formed the basis for inflated, complex systems.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All materials generated in this study are freely available upon request from the corresponding author. All data are available in the main text or the supplementary materials. The crystallographic data are available from the RCSB PDB (accession number 6SDG), and the RNA-seq data are available from the NCBI SRA under the project accession number PRJNA554398 (http://www.ncbi.nlm.nih.gov/bioproject/554398).
Code availability
The Python script used to calculate the residues in dimer interface is available at http://www.protein.osaka-u.ac.jp/rcsfp/supracryst/suzuki/jpxtal/Katsutani/InterfaceResidues.py.
References
Du, Y. & Scheres, B. Lateral root formation and the multiple roles of auxin. J. Exp. Bot. 69, 155–167 (2018).
Vanneste, S. & Friml, J. Auxin: a trigger for change in plant development. Cell 136, 1005–1016 (2009).
Weijers, D. & Wagner, D. Transcriptional responses to the auxin hormone. Annu. Rev. Plant Biol. 67, 539–574 (2016).
Mutte, S. K. et al. Origin and evolution of the nuclear auxin response system. eLife 7, e33399 (2018).
Flores-Sandoval, E. et al. Class C ARFs evolved before the origin of land plants and antagonize differentiation and developmental transitions in Marchantia polymorpha. New Phytol. 218, 1612–1630 (2018).
Finet, C., Berne-Dedieu, A., Scutt, C. P. & Marlétaz, F. Evolution of the ARF gene family in land plants: old domains, new tricks. Mol. Biol. Evol. 30, 45–56 (2013).
Ulmasov, T., Hagen, G. & Guilfoyle, T. J. Activation and repression of transcription by auxin response factors. Proc. Natl Acad. Sci. USA 96, 5844–5849 (1999).
Vernoux, T. et al. The auxin signalling network translates dynamic input into robust patterning at the shoot apex. Mol. Syst. Biol. 7, 508 (2011).
Piya, S., Shrestha, S. K., Binder, B., Stewart, C. N. Jr. & Hewezi, T. Protein–protein interaction and gene co-expression maps of ARFs and Aux/IAAs in Arabidopsis. Front. Plant Sci. 5, 744 (2014).
Lavy, M. et al. Constitutive auxin response in Physcomitrella reveals complex interactions between Aux/IAA and ARF proteins. eLife 5, e13325 (2016).
Zhao, Z. et al. Hormonal control of the shoot stem-cell niche. Nature 465, 1089–1092 (2010).
Boer, D. R. et al. Structural basis for DNA binding specificity by the auxin-dependent ARF transcription factors. Cell 156, 577–589 (2014).
Hori, K. et al. Klebsormidium flaccidum genome reveals primary factors for plant terrestrial adaptation. Nat. Commun. 5, 3978 (2014).
Bowman, J. L. et al. Insights into land plant evolution garnered from the Marchantia polymorpha genome. Cell 171, 287–304 (2017).
Kato, H. et al. Auxin-mediated transcriptional system with a minimal set of components is critical for morphogenesis through the life cycle in Marchantia polymorpha. PLoS Genet. 11, e1005084 (2015).
Flores-Sandoval, E., Eklund, D. M. & Bowman, J. L. A simple auxin transcriptional response system regulates multiple morphogenetic processes in the liverwort Marchantia polymorpha. PLoS Genet. 11, e1005207 (2015).
Kato, H. et al. The roles of the sole activator-type auxin response factor in pattern formation of Marchantia polymorpha. Plant Cell Physiol. 58, 1642–1651 (2017).
Flores-Sandoval, E., Romani, F. & Bowman, J. L. Co-expression and transcriptome analysis of Marchantia polymorpha transcription factors supports class C ARFs as independent actors of an ancient auxin regulatory module. Front. Plant Sci. 9, 1345 (2018).
Tiwari, S. B., Hagen, G. & Guilfoyle, T. The roles of auxin response factor domains in auxin-responsive transcription. Plant Cell 15, 533–543 (2003).
Choi, H. S., Seo, M. & Cho, H. T. Two TPL-binding motifs of ARF2 are involved in repression of auxin responses. Front. Plant Sci. 9, 372 (2018).
Korasick, D. A. et al. Molecular basis for AUXIN RESPONSE FACTOR protein interaction and the control of auxin response repression. Proc. Natl Acad. Sci. USA 111, 5427–5432 (2014).
Nanao, M. H. et al. Structural basis for oligomerization of auxin transcriptional regulators. Nat. Commun. 5, 3617 (2014).
Sayou, C. et al. A SAM oligomerization domain shapes the genomic binding landscape of the LEAFY transcription factor. Nat. Commun. 7, 11222 (2016).
Ishizaki, K. et al. Development of Gateway binary vector series with four different selection markers for the liverwort Marchantia polymorpha. PLoS ONE 10, e0138876 (2015).
Ishizaki, K., Johzuka-Hisatomi, Y., Ishida, S., Iida, S. & Kohchi, T. Homologous recombination-mediated gene targeting in the liverwort Marchantia polymorpha L. Sci. Rep. 3, 1532 (2013).
Sugano, S. S. et al. Efficient CRISPR/Cas9-based genome editing and its application to conditional genetic analysis in Marchantia polymorpha. PLoS ONE 13, e0205117 (2018).
Nishihama, R., Ishida, S., Urawa, H., Kamei, Y. & Kohchi, T. Conditional gene expression/deletion systems for Marchantia polymorpha using its own Heat-Shock promoter and Cre/loxP-mediated site-specific recombination. Plant Cell Physiol. 57, 271–280 (2016).
Hohlbein, J., Craggs, T. D. & Cordes, T. Alternating-laser excitation: single-molecule FRET and beyond. Chem. Soc. Rev. 43, 1156–1171 (2014).
Leyser, O. Auxin signaling. Plant Physiol. 176, 465–479 (2018).
Rademacher, E. H. et al. A cellular expression map of the Arabidopsis AUXIN RESPONSE FACTOR gene family. Plant J. 68, 597–606 (2011).
Chiyoda, S., Ishizaki, K., Kataoka, H., Yamato, K. T. & Kohchi, T. Direct transformation of the liverwort Marchantia polymorpha L. by particle bombardment using immature thalli developing from spores. Plant Cell Rep. 27, 1467–1473 (2008).
Ishizaki, K., Chiyoda, S., Yamato, K. T. & Kohchi, T. Agrobacterium-mediated transformation of the haploid liverwort Marchantia polymorpha L., an emerging model for plant biology. Plant Cell Physiol. 49, 1084–1091 (2008).
Kubota, A., Ishizaki, K., Hosaka, M. & Kohchi, T. Efficient agrobacterium-mediated transformation of the liverwort Marchantia polymorpha using regenerating thalli. Biosci. Biotechnol. Biochem. 77, 167–172 (2013).
Zhang, Y., Werling, U. & Edelmann, W. SLiCE: a novel bacterial cell extract-based DNA cloning method. Nucleic Acids Res. 40, e55 (2012).
Juanhuix, J. et al. Developments in optics and performance at BL13-XALOC, the macromolecular crystallography beamline at the ALBA Synchrotron. J. Synchrotron Radiat. 21, 679–689 (2014).
Vonrhein, C. et al. Data processing and analysis with the autoPROC toolbox. Acta Crystallogr. D 67, 293–302 (2011).
Tickle, I. J. et al. STARANISO (Global Phasing Ltd., 2018); http://staraniso.globalphasing.org/cgi-bin/staraniso.cgi
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Webb, B. & Sali, A. Comparative protein structure modeling using MODELLER. Curr. Protoc. Bioinformatics 54, 5.6.1–5.6.37 (2016).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
van Zundert, G. C. P. et al. The HADDOCK2.2 web server: user-friendly integrative modeling of biomolecular complexes. J. Mol. Biol. 428, 720–725 (2016).
Truernit, E. et al. High-resolution whole-mount imaging of three-dimensional tissue organization and gene expression enables the study of phloem development and structure in Arabidopsis. Plant Cell 20, 1494–1503 (2008).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Raissig, M. T., Gagliardini, V., Jaenisch, J., Grossniklaus, U. & Baroux, C. Efficient and rapid isolation of early-stage embryos from Arabidopsis thaliana seeds. J. Vis. Exp. 76, e50371 (2013).
Trombetta, J. J. et al. Preparation of single-cell RNA-Seq libraries for next generation sequencing. Curr. Protoc. Mol. Biol. 107, 4.22.1–4.22.17 (2014).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-Seq data with DESeq2. Genome Biol. 15, 550 (2014).
Freire-Rios, A., Radoeva, T., De Rybel, B., Weijers, D. & Borst, J. W. FRET-FLIM for visualizing and quantifying protein interactions in live plant cells. Meth. Mol. Biol. 1497, 135–146 (2017).
Matsuo, N., Minami, M., Maeda, T. & Hiratsuka, K. Dual luciferase assay for monitoring transient gene expression in higher plants. Plant Biotechnol. 18, 71–75 (2001).
Akagi, T., Ikegami, A. & Yonemori, K. DkMyb2 wound-induced transcription factor of persimmon (Diospyros kaki Thunb.), contributes to proanthocyanidin regulation. Planta 232, 1045–1059 (2010).
van Dijk, M. & Bonvin, A. M. 3D-DART: a DNA structure modelling server. Nucleic Acids Res. 37, W235–W239 (2009).
Kalinin, S. et al. A toolkit and benchmark study for FRET-restrained high-precision structural modeling. Nat. Methods 9, 1218–1225 (2012).
Craggs, T. D. et al. Substrate conformational dynamics drive structure-specific recognition of gapped DNA by DNA polymerase. Nucleic Acids Res. 47, 10788–10800 (2019).
Farooq, S. & Hohlbein, J. Camera-based single-molecule FRET detection with improved time resolution. Phys. Chem. Chem. Phys. 17, 27862–27872 (2015).
Cordes, T., Vogelsang, J. & Tinnefeld, P. On the mechanism of Trolox as antiblinking and antibleaching reagent. J. Am. Chem. Soc. 131, 5018–5019 (2009).
Rasnik, I., McKinney, S. A. & Ha, T. Nonblinking and long-lasting single-molecule fluorescence imaging. Nat. Methods 3, 891–893 (2006).
Evans, G. W., Hohlbein, J., Craggs, T., Aigrain, L. & Kapanidis, A. N. Real-time single-molecule studies of the motions of DNA polymerase fingers illuminate DNA synthesis mechanisms. Nucleic Acids Res. 43, 5998–6008 (2015).
Acknowledgements
We thank the XALOC staff at the synchrotron ALBA and V. P. Carrillo, Carrasco, S. Kiryu, M. Katayama and L. Olijslager for experimental support, D. Gadella and K. Hiratsuka for providing materials, and O. Leyser for comments in the manuscript. This work was supported by an EMBO Long-term Fellowship (ALTF 415-2016) to H.K., a PhD fellowship from the Graduate School Experimental Plant Sciences to J.H. and D.W., a VICI grant (no. 865.14.001) from the Netherlands Organization for Scientific Research (NWO) to D.W., grants from the Ministry of Economy and Competitiveness of the Spanish Government (nos BIO2016-77883-C2-2-P and FIS2015-72574-EXP) (AEI/FEDER,EU) to D.R.B., an ALW-open grant (no. ALWOP.402) from the NWO to J.W.B., JSPS/MEXT KAKENHI (grant nos 19K23751 to H.K., 18J12698 to H.S, 19K016166 to Y.Y., 17H06472 to K.I. 18H04836 to R.N. and 25113009, 15K21758 and 19H05675 to T.K.) and SPIRITS 2017 of Kyoto University to R.N.
Author information
Authors and Affiliations
Contributions
H.K., R.N., T.K. and D.W. conceptualized the project. H.K., S.K.M., H.S., I.C., T.R., S.D., M.F., E.H., W.v.d.B. and S.L. conducted the investigation. H.K., S.K.M., I.C. and M.F. conducted the formal analysis. J.H., D.R.B., R.N., T.K. and D.W. supervised the project. K.I., J.H., R.N., T.K., J.W.B. and D.W. acquired the funding. H.K. and D.W. wrote the original draft of the paper. All authors reviewed and edited the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Transgene expression and phenotypes of DBD swap lines.
a, Expression of transgenic transcripts in gemmalings of three independent proARF1-ARF1, proARF1-ARF2 and proARF1-ARF3 lines, measured by qRT-PCR using the common 5’-UTR fragment. Circles indicate each data point for three technical replicates. ND: not detected in Tak-1 wild-type and arf1-4 mutant. b, Ten-day-old gemmalings and c, immature gemmae of Tak-1 wild-type, Mparf1-4, proMpARF1:ARF111 (111), proMpARF1:ARF211 (211) and proMpARF1:ARF311 (311). Bars = 2 mm in (b) and 0.1 mm in (c). The experiment was repeated twice with similar results.
Extended Data Fig. 2 Gemmae phenotypes in Mparf1 and Mparf3 mutants.
Optical medial sections through early- (top row) and late-stage (bottom row) gemmae from Tak-1, Mparf1-4 and Mparf3ge1-1 plants. Medial sections were taken from full 3D stacks of gemmae in which cell walls were labeled using mPS-PI staining. The positions of the future apical notches are indicated by red arrowheads in Tak-1. Note that these indentations are missing in both mutants. Bars are 20 µm in all panels. The experiment was repeated twice with similar results.
Extended Data Fig. 3 Gene Ontology analysis of genes misexpressed in arf1 and arf3 gemmae.
GO categories ‘Biological process’ and ‘Molecular function’ were shown for downregulated genes as directed graphs with least- and most- enriched categories are shown in yellow and red blocks, respectively, as well as in the table underneath the figures. Note the different categories enriched in arf1 and arf3. P-values (table) are derived from a Fisher’s exact test.
Extended Data Fig. 4 Phenotypes induced by Middle Region domain swaps in Tak-1 background.
Ten-day-old gemmalings of Tak-1 wild-type, and 2 independent lines each for proMpARF1:ARF121 (121) and proMpARF1:ARF131 (131) Bar = 1 mm. The experiment was repeated four times with similar results.
Extended Data Fig. 5 Expression of MpTPL and MpARFs for FRET-FLIM.
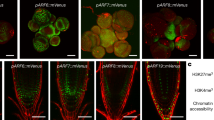
Fluorescence of MpTPL-mNeongreen (FRET donor; top row), either co-expressed with fusions of mScarlet-I (FRET acceptor; middle row), to empty vector or with MpARF1-mScarlet-I, MpARF2-mScarlet-I or MpARF2(LFG-AAA)-mScarlet-I. Bottom panel shows overlay of mNeongreen, mScarlet-I and chloroplast autofluorescence signals. Note that all fusion proteins localize to nuclei. Bar is 5 µm. The experiment was repeated twice with similar results and with 15, 10, 10 and 15 protoplasts (left to right).
Extended Data Fig. 6 Biological significance of the MpARF LFG motifs.
a, Ten-day-old gemmalings of proMpARF1:ARF11Δ1 (11Δ1), proMpARF1:ARF12Δ1 (12Δ1) and proMpARF1:ARF13Δ1 (13Δ1) lines. The experiment was repeated twice with similar results. b, Relative qPCR expression of WIP gene in Tak-1 wild-type, proMpARF1:ARF12Δ1 (12Δ1) and proMpARF1:ARF13Δ1 (13Δ1) gemmalings, 0, 1, 2 and 4 hours after treatment with 10 µM 2,4-D. The experiment was performed once with three technical replicates. c, Relative qPCR expression of YUC2 gene in Tak-1 wild-type, Mparf1-4 and proMpARF1:ARF11Δ1 (11Δ1) gemmalings, after a 1-hour treatment with control media or with 10 µM 2,4-D. n = 3 technical replicates. The experiment was repeated twice with similar results. Bar in (a) = 2 mm.
Extended Data Fig. 7 Docking of MpARF and MpIAA PB1 domains.
Best fit structural models of molecular docking of PB1 domains of MpARF1 (top), MpARF2 (middle) and MpARF3 (bottom) with MpIAA. Residues of positive (K, Lysine) and negative (OPCA) faces are indicated as sticks.
Extended Data Fig. 8 Antagonistic action of MpARF1 and MpARF2.
Fourteen-day-old gemmalings of Tak-1, proEF:MpARF1, proEF:MpARF2, and proEF:MpARF1 proEF:MpARF2 transgenic lines grown on control medium, or on media containing 1, 3 or 10 µM 2,4-D. The proEF:MpARF1 proEF:MpARF2 line was generated by introducing the proEF:MpARF1 cassette into the presented proEF:MpARF2 line. Bar = 1 cm. The experiment was repeated twice with similar results.
Extended Data Fig. 9 Conditional MpARF2 inactivation.
a, Design of transgene for inducible removal of MpARF2. A CRISPR-resistant MpARF2 version (ARF2m; through engineered silent mutations) is flanked by loxP sites and an upstream EF promoter and a downstream tdTomato-NLS gene, in a construct that also harbors a DEX-inducible Cre-GR version driven from a heat-inducible MpHSP17.8A1 promoter (proHSP). Following heat shock, and in the presence of DEX, Cre-GR will recombine the two loxP sites and excise the ARF2m locus. Cells in which recombination occurred are marked by proEF:tdTomato-NLS expression. b, CRISPR/Cas9-induced mutations in the endogenous MpARF2 gene in Mparf2cko genetic background. Two different single-base insertions were recovered, generating two independent Mparf2cko alleles. c, Phenotypes of 8-day-old gemmalings of Tak-1, Mparf2-1cko or Mparf2-2cko lines grown on control medium or with 1 µM DEX after a heat shock on day 1. The experiment was repeated four times with similar results. d, PCR amplicons detecting the endogenous ARF2 (upper band) and ARF2m (lower band) genes in Tak-1 and in 3 independent transgenic lines harboring ARF2m (#11, #14, #18; prior to introduction of the CRISPR/Cas9 construct), either grown on control media, or on 1 µM DEX after a heat shock. The experiment was repeated twice with similar results. e, tdTomato expression (lower panels) and appearance (top panels) of 3-day-old Mparf2cko gemmalings grown on control medium or with 1 µM DEX after a heat shock on day 1. The experiment was repeated four times with similar results. Bars are 1 mm in (c) and 0.2 mm in (e).
Extended Data Fig. 10 Single-molecule FRET analysis of ARF-DNA interaction.
a, Schematic representation of the ER7 DNA oligonucleotide used in single-molecule FRET experiments, harboring two inverted ARF binding sites (indicated in bold lettering and with a red arrow on top- and bottom-strand, respectively). The bottom-strand was 5’ biotinylated (indicated with an orange hexagon, labeled B) to facilitate immobilization on cover slips. FRET-compatible Cy3B (green circle) and ATTO647N (magenta circle) labels were attached immediately upstream of each ARF binding site indicated with yellow). b, Simulations of accessible volumes of the Cy3B and ATTO647N dyes on the ER7 oligonucleotide in the absence (left) or presence (right) of MpARF2 DBD (based on the AtARF2-ER7 complex, PDB ID: 6SDG). Note that dye clouds are constrained upon protein binding, predicting reduced FRET efficiency. c, Histograms showing FRET efficiency (x-axis) and Cy3B/ATTO6437N stoichiometry (y-axis) of the immobilized ER7 oligonucleotide incubated without MpARF1 DBD (left) or with 256 nM MpARF1 DBD (right). Each dot represents a single DNA molecule. Note the shift in FRET efficiency induced by protein binding. d,e, Distribution histograms of FRET efficiency (x-axis) of the labeled and immobilized ER7 oligonucleotide in the presence of increasing concentrations (0-256 nM) of MpARF1 DBD (d) or MpARF2 DBD (e). Each titration series is followed by an incubation without protein to validate recovery to unbound state. Histograms show fits of the two FRET states representing unbound and bound states, and indicate percentages of DNA molecules in each state.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2, and Tables 1 and 2.
Rights and permissions
About this article
Cite this article
Kato, H., Mutte, S.K., Suzuki, H. et al. Design principles of a minimal auxin response system. Nat. Plants 6, 473–482 (2020). https://doi.org/10.1038/s41477-020-0662-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-020-0662-y
This article is cited by
-
Genomic survey and expression analysis of LcARFs reveal multiple functions to somatic embryogenesis in Liriodendron
BMC Plant Biology (2024)
-
Evolution and function analysis of auxin response factors reveal the molecular basis of the developed root system of Zygophyllum xanthoxylum
BMC Plant Biology (2024)
-
Transcriptional regulation of tomato fruit ripening
Physiology and Molecular Biology of Plants (2024)
-
Four class A AUXIN RESPONSE FACTORs promote tomato fruit growth despite suppressing fruit set
Nature Plants (2023)
-
Cis-regulatory elements and transcription factors related to auxin signaling in the streptophyte algae Klebsormidium nitens
Scientific Reports (2023)