Abstract
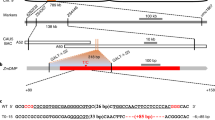
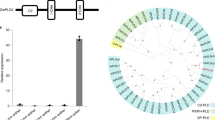
Doubled haploid technology using inducer lines carrying mutations in ZmPLA1/MTL/NLD and ZmDMP1,2,3,4 has revolutionized traditional maize breeding. ZmPLA1/MTL/NLD is conserved in monocots and has been used to extend the system from maize to other monocots5,6,7, but no functional orthologue has been identified in dicots, while ZmDMP-like genes exist in both monocots and dicots4,8,9. Here, we report that loss-of-function mutations in the Arabidopsis thaliana ZmDMP-like genes AtDMP8 and AtDMP9 induce maternal haploids, with an average haploid induction rate of 2.1 ± 1.1%. In addition, to facilitate haploid seed identification in dicots, we established an efficient FAST-Red fluorescent marker-based haploid identification system that enables the identification of haploid seeds with >90% accuracy. These results show that mutations in DMP genes also trigger haploid induction in dicots. The conserved expression patterns and amino acid sequences of ZmDMP-like genes in dicots suggest that DMP mutations could be used to develop in vivo haploid induction systems in dicots.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
The datasets generated and/or analysed during the current study are available from the corresponding author on request. The raw whole-genome sequencing data have been deposited in the NCBI Sequence Read Archive under the accession code PRJNA608056. Source Data for Figs. 1 and 2 are provided with the paper.
References
Gilles, L. M. et al. Loss of pollen‐specific phospholipase NOT LIKE DAD triggers gynogenesis in maize. EMBO J. 36, 707–717 (2017).
Kelliher, T. et al. MATRILINEAL, a sperm-specific phospholipase, triggers maize haploid induction. Nature 542, 105–109 (2017).
Liu, C. et al. A 4-bp insertion at ZmPLA1 encoding a putative phospholipase A generates haploid induction in maize. Mol. Plant 10, 520–522 (2017).
Zhong, Y. et al. Mutation of ZmDMP enhances haploid induction in maize. Nat. Plants 5, 575–580 (2019).
Yao, L. et al. OsMATL mutation induces haploid seed formation in indica rice. Nat. Plants 4, 530–533 (2018).
Liu, C. et al. Extension of the in vivo haploid induction system from diploid maize to hexaploid wheat. Plant Biotechnol. J. 18, 316–318 (2019).
Liu, H. et al. Efficient induction of haploid plants in wheat by editing of TaMTL using an optimized Agrobacterium-mediated CRISPR system. J. Exp. Bot. 71, 1337–1349 (2020).
Takahashi, T. et al. The male gamete membrane protein DMP9/DAU2 is required for double fertilization in flowering plants. Development 145, dev170076 (2018).
Cyprys, P., Lindemeier, M. & Sprunck, S. Gamete fusion is facilitated by two sperm cell-expressed DUF679 membrane proteins. Nat. Plants 5, 253–257 (2019).
Chang, M.-T. & Coe, E. H. in Molecular Genetic Approaches to Maize Improvement (eds Kriz, A. L. & Larkins, B. A.) 127–142 (Springer Berlin Heidelberg, 2009).
Dwivedi, S. L. et al. Haploids: constraints and opportunities in plant breeding. Biotechnol. Adv. 33, 812–829 (2015).
Kalinowska, K. et al. State-of-the-art and novel developments of in vivo haploid technologies. Theor. Appl. Genet. 132, 593–605 (2019).
Coe, E. H. A line of maize with high haploid frequency. Am. Nat. 93, 381–382 (1959).
Ravi, M. & Chan, S. W. L. Haploid plants produced by centromere-mediated genome elimination. Nature 464, 615–618 (2010).
Kelliher, T. et al. Maternal haploids are preferentially induced by CENH3-tailswap transgenic complementation in maize. Front. Plant Sci. 7, 414 (2016).
Watts, A., Kumar, V., Raipuria, R. K. & Bhattacharya, R. C. In vivo haploid production in crop plants: methods and challenges. Plant Mol. Biol. Rep. 36, 685–694 (2018).
Barret, P., Brinkmann, M. & Beckert, M. A major locus expressed in the male gametophyte with incomplete penetrance is responsible for in situ gynogenesis in maize. Theor. Appl. Genet. 117, 581–594 (2008).
Prigge, V. et al. New insights into the genetics of in vivo induction of maternal haploids, the backbone of doubled haploid technology in maize. Genetics 190, 781–793 (2012).
Dong, X. et al. Fine mapping of qhir1 influencing in vivo haploid induction in maize. Theor. Appl. Genet. 126, 1713–1720 (2013).
Liu, C. et al. Fine mapping of qhir8 affecting in vivo haploid induction in maize. Theor. Appl. Genet. 128, 2507–2515 (2015).
Kasaras, A. & Kunze, R. Expression, localisation and phylogeny of a novel family of plant-specific membrane proteins. Plant Biol. 12, 140–152 (2010).
Prigge, V. et al. Doubled haploids in tropical maize: I. Effects of inducers and source germplasm on in vivo haploid induction rates. Crop Sci. 51, 1498–1506 (2011).
Wu, P., Li, H., Ren, J. & Chen, S. Mapping of maternal QTLs for in vivo haploid induction rate in maize (Zea mays L.). Euphytica 196, 413–421 (2014).
Tian, X. et al. Hetero-fertilization together with failed egg–sperm cell fusion supports single fertilization involved in in vivo haploid induction in maize. J. Exp. Bot. 69, 4689–4701 (2018).
Li, L., Xu, X., Jin, W. & Chen, S. Morphological and molecular evidences for DNA introgression in haploid induction via a high oil inducer CAUHOI in maize. Planta 230, 367–376 (2009).
Zhao, X., Xu, X., Xie, H., Chen, S. & Jin, W. Fertilization and uniparental chromosome elimination during crosses with maize haploid inducers. Plant Physiol. 163, 721–731 (2013).
Kelliher, T. et al. One-step genome editing of elite crop germplasm during haploid induction. Nat. Biotechnol. 37, 287–292 (2019).
Nanda, D. K. & Chase, S. S. An embryo marker for detecting monoploids of maize (Zea Mays L.). Crop Sci. 6, 213–215 (1966).
Portemer, V., Renne, C., Guillebaux, A. & Mercier, R. Large genetic screens for gynogenesis and androgenesis haploid inducers in Arabidopsis thaliana failed to identify mutants. Front. Plant Sci. 6, 147 (2015).
Wang, B. et al. Development of a haploid-inducer mediated genome editing system for accelerating maize breeding. Mol. Plant 12, 597–602 (2019).
Rosso, M. G. et al. An Arabidopsis thaliana T-DNA mutagenized population (GABI-Kat) for flanking sequence tag-based reverse genetics. Plant Mol. Biol. 53, 247–259 (2003).
Castel, B., Tomlinson, L., Locci, F., Yang, Y. & Jones, J. D. G. Optimization of T-DNA architecture for Cas9-mediated mutagenesis in Arabidopsis. PLoS ONE 14, e0204778 (2019).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Chen, C., Chen, H., He, Y. & Xia, R. TBtools, a toolkit for biologists integrating various biological data handling tools with a user-friendly interface. Preprint at https://www.biorxiv.org/content/10.1101/289660v3 (2018).
Păcurar, D. I. et al. A collection of INDEL markers for map-based cloning in seven Arabidopsis accessions. J. Exp. Bot. 63, 2491–2501 (2012).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Poplin, R. et al. Scaling accurate genetic variant discovery to tens of thousands of samples. Preprint at https://www.biorxiv.org/content/10.1101/201178v3 (2017).
Kurtz, S. et al. Versatile and open software for comparing large genomes. Genome Biol. 5, R12 (2004).
Tan, E. H. et al. Catastrophic chromosomal restructuring during genome elimination in plants. eLife 4, e06516 (2015).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Acknowledgements
We thank D. Ye and L. Chen for providing the ms1 and ms1-1 mutants, X. Yang for help with the analysis of haploid sequence data, Y. Leng for help with the analysis of seed development, and Z. Li for help with the phylogenetic analysis. This work was supported by the National Key Research and Development Program of China (2016YFD0101200 and 2018YFD0100201), Modern Maize Industry Technology System (CARS-02-04) and National Natural Science Foundation of China (91935303) (to S.C), as well as a China Scholarship Council grant (to B.C).
Author information
Authors and Affiliations
Contributions
Y.Z., B.C., C.L. and S.C. conceived of and designed the experiments. Y.Z., M.L. and B.C. performed most of the experiments. D.W., Y.J., X.Q., Z.L., C.C., Y.W., M.C., J.L., Z.X. and D.C. performed some of the experiments. Y.Z., B.C., S.C., C.L., M.W., W.L. and M.L. analysed the data. Y.Z., B.C., C.L., K.B. and S.C. discussed and wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Thomas Widiez and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–6 and Tables 2, 4, 6 and 7.
Supplementary Table 1
ZmDMP-like genes in plants.
Supplementary Table 3
A summary of CRISPR–Cas9 mutagenesis.
Supplementary Table 5
Genotyping data of haploids.
Supplementary Table 8
SNP data of haploids.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Rights and permissions
About this article
Cite this article
Zhong, Y., Chen, B., Li, M. et al. A DMP-triggered in vivo maternal haploid induction system in the dicotyledonous Arabidopsis. Nat. Plants 6, 466–472 (2020). https://doi.org/10.1038/s41477-020-0658-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-020-0658-7
This article is cited by
-
One-step creation of CMS lines using a BoCENH3-based haploid induction system in Brassica crop
Nature Plants (2024)
-
Mechanism and molecular basis of apomixis in angiosperms
Vegetos (2024)
-
Potential gene editing targets for developing haploid inducer stocks in rice and wheat with high haploid induction frequency
3 Biotech (2024)
-
Design of an Arabidopsis thaliana reporter line to detect heat-sensing and signaling mutants
Plant Methods (2023)
-
Comprehensive identification and functional characterization of GhpPLA gene family in reproductive organ development
BMC Plant Biology (2023)