Abstract
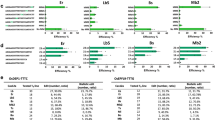
Clustered regularly interspaced short palindromic repeats (CRISPR)–Cas12b is a newly emerged genome engineering system. Here, we compared Cas12b from Alicyclobacillus acidoterrestris (Aac), Alicyclobacillus acidiphilus (Aa), Bacillus thermoamylovorans (Bth) and Bacillus hisashii (Bh) for genome engineering in rice, an important crop. We found AaCas12b was more efficient than AacCas12b and BthCas12b for targeted mutagenesis, which was further demonstrated in multiplexed genome editing. We also engineered the Cas12b systems for targeted transcriptional repression and activation. Our work establishes Cas12b as the third promising CRISPR system, after Cas9 and Cas12a, for plant genome engineering.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The 29 Gateway compatible vectors for the CRISPR–Cas12b systems are available from Addgene: pYPQ290 (no. 129670), pYPQ291 (no. 129671), pYPQ292 (no. 129672), pYPQ290-D570A (no. 129673), pYPQ290-D977A (no. 129674), pYPQ290-E848A (no. 129675), pYPQ291-D573A (no. 129676), pYPQ291-D951A (no. 129677), pYPQ291-E827A (no. 129678), pYPQ292-D570A (no. 129679), pYPQ292-D977A (no. 129680), pYPQ292-E848A (no. 129681), pYPQ290-D570A-SRDX (no. 129682), pYPQ291-D573A-SRDX (no. 129683), pYPQ292-D570A-SRDX (no. 129684), pYPQ141-ZmUbi-RZ-Aac (no. 129685), pYPQ141-ZmUbi-RZ-Bth (no. 129686), pYPQ141-ZmUbi-RZ-Aa1.2.3 (no. 136372), pYPQ141-ZmUbi-RZ-Aa1.2 (no. 136373), pYPQ141-ZmUbi-RZ-Aa3.8.3 (no. 136374), pYPQ141-ZmUbi-RZ-Aa3.8.4 (no. 136375), pYPQ141-ZmUbi-RZ-Aa3.8 (no. 136376), pYPQ141-ZmUbi-RZ-Aac.3 (no. 136377), pYPQ141-ZmUbi-RZ-Bh (no. 136378), pYPQ239A (dFnCas12a)-TV (no. 136379), pYPQ292 (AaCas12b)-D570-TV (no. 136380), pYPQ292 (AaCas12b)-D570-TV-MS2-TV (no. 136381), pYPQ292 (AaCas12b)-D570-TV-MS2-VPR (no. 136382) and pYPQ293 (BhCas12b_v4) (no. 136383). The high-throughput sequencing data sets have been submitted to the National Center for Biotechnology information (NCBI) database under Sequence Read Archive (SRA) BioProject ID PRJNA553352.
References
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 163, 759–771 (2015).
Zhang, Y., Malzahn, A., Sretenovic, S. & Qi, Y. The emerging and uncultivated potential of CRISPR technology in plant science. Nat. Plants 5, 778–791 (2019).
Shmakov, S. et al. Discovery and functional characterization of diverse class 2 CRISPR-Cas systems. Mol. Cell 60, 385–397 (2015).
Teng, F. et al. Repurposing CRISPR-Cas12b for mammalian genome engineering. Cell Discov. 4, 63 (2018).
Strecker, J. et al. Engineering of CRISPR–Cas12b for human genome editing. Nat. Commun. 10, 212 (2019).
Yang, H., Gao, P., Rajashankar, K. R. & Patel, D. J. PAM-dependent target DNA recognition and cleavage by C2c1 CRISPR-Cas endonuclease. Cell 167, 1814–1828 (2016).
Liu, L. et al. C2c1-sgRNA complex structure reveals RNA-guided DNA cleavage mechanism. Mol. Cell 65, 310–322 (2017).
Wu, D., Guan, X., Zhu, Y., Ren, K. & Huang, Z. Structural basis of stringent PAM recognition by CRISPR-C2c1 in complex with sgRNA. Cell Res. 27, 705–708 (2017).
Tang, X. et al. A CRISPR–Cpf1 system for efficient genome editing and transcriptional repression in plants. Nat. Plants 3, 17018 (2017).
Zhong, Z. et al. Plant genome editing using FnCpf1 and LbCpf1 nucleases at redefined and altered PAM sites. Mol. Plant 11, 999–1002 (2018).
Jain, I. et al. Defining the seed sequence of the Cas12b CRISPR-Cas effector complex. RNA Biol. 16, 413–422 (2019).
Paul, J. W. 3rd & Qi, Y. CRISPR/Cas9 for plant genome editing: accomplishments, problems and prospects. Plant Cell Rep. 35, 1417–1427 (2016).
Fu, Y., Sander, J. D., Reyon, D., Cascio, V. M. & Joung, J. K. Improving CRISPR–Cas nuclease specificity using truncated guide RNAs. Nat. Biotechnol. 32, 279–284 (2014).
Lowder, L. G. et al. A CRISPR/Cas9 toolbox for multiplexed plant genome editing and transcriptional regulation. Plant Physiol. 169, 971–985 (2015).
Lowder, L. G. et al. Robust transcriptional activation in plants using multiplexed CRISPR-Act2.0 and mTALE-act systems. Mol. Plant 11, 245–256 (2018).
Teng, F. et al. Artificial sgRNAs engineered for genome editing with new Cas12b orthologs. Cell Discov. 5, 23 (2019).
Li, Z. et al. A potent Cas9-derived gene activator for plant and mammalian cells. Nat. Plants 3, 930–936 (2017).
Chavez, A. et al. Highly efficient Cas9-mediated transcriptional programming. Nat. Methods 12, 326–328 (2015).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Tang, X. et al. A single transcript CRISPR-Cas9 system for efficient genome editing in plants. Mol. Plant 9, 1088–1091 (2016).
You, Q. et al. CRISPRMatch: An automatic calculation and visualization tool for high-throughput CRISPR genome-editing data analysis. Int J. Biol. Sci. 14, 858–862 (2018).
Liu, W. et al. DSDecode: A web-based tool for decoding of sequencing chromatograms for genotyping of targeted mutations. Mol. Plant 8, 1431–1433 (2015).
Bae, S., Park, J. & Kim, J. S. Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 30, 1473–1475 (2014).
Acknowledgements
This work was supported by University of Maryland startup funds, the National Science Foundation Plant Genome Research Program grant (award no. IOS-1758745), the Biotechnology Risk Assessment Grant Program competitive grant (award no. 2018-33522-28789) from the US Department of Agriculture, Foundation for Food and Agriculture Research grant (award no. 593603) and Syngenta Biotechnology to Y.Q. It was also supported by the National Transgenic Major Project (award nos. 2019ZX08010003-001-002 and 2018ZX08020-003), the National Science Foundation of China (award no. 31771486), the Sichuan Youth Science and Technology Foundation (award no. 2017JQ0005) and the Science Strength Promotion Program of UESTC to Yong Z. M.M was supported by a scholarship from China Scholarship Council. A.M was supported by a scholarship from Cosmos Club Foundation.
Author information
Authors and Affiliations
Contributions
Y.Q. and Yong Z. designed the experiments. M.M, Yingxiao Z. and C.P. generated all the constructs. M.M., Q.R., Y.H., S.L., Z.Z., J.W. and X.Z. performed the transient assays in protoplasts. Q.R. and Y.H. prepared samples for high-throughput sequencing. M.M. generated stable transgenic rice and analysed the plants. C.P. conducted transcriptional repression and activation assays. Y.Q., Yong Z., A.M. and J.W. wrote the paper with input from other authors. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Sang Gyu Kim and Huawei Zhang and the other, anonymous, reviewer for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary methods, Supplementary Figs. 1–20 and Supplementary Tables 1 and 2.
Rights and permissions
About this article
Cite this article
Ming, M., Ren, Q., Pan, C. et al. CRISPR–Cas12b enables efficient plant genome engineering. Nat. Plants 6, 202–208 (2020). https://doi.org/10.1038/s41477-020-0614-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-020-0614-6
This article is cited by
-
Genome editing based trait improvement in crops: current perspective, challenges and opportunities
The Nucleus (2024)
-
CRISPR/Cas-mediated germplasm improvement and new strategies for crop protection
Crop Health (2024)
-
Targeted genome editing for cotton improvement: prospects and challenges
The Nucleus (2024)
-
Developing a CRISPR/FrCas9 system for core promoter editing in rice
aBIOTECH (2024)
-
The applications of CRISPR/Cas-mediated genome editing in genetic hearing loss
Cell & Bioscience (2023)