Abstract
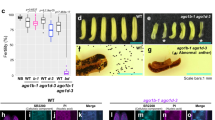
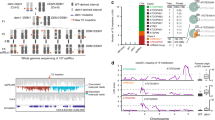
Although plants are able to withstand a range of environmental conditions, spikes in ambient temperature can impact plant fertility causing reductions in seed yield and notable economic losses1,2. Therefore, understanding the precise molecular mechanisms that underpin plant fertility under environmental constraints is critical to safeguarding future food production3. Here, we identified two Argonaute-like proteins whose activities are required to sustain male fertility in maize plants under high temperatures. We found that MALE-ASSOCIATED ARGONAUTE-1 and -2 associate with temperature-induced phased secondary small RNAs in pre-meiotic anthers and are essential to controlling the activity of retrotransposons in male meiocyte initials. Biochemical and structural analyses revealed how male-associated Argonaute activity and its interaction with retrotransposon RNA targets is modulated through the dynamic phosphorylation of a set of highly conserved, surface-located serine residues. Our results demonstrate that an Argonaute-dependent, RNA-guided surveillance mechanism is critical in plants to sustain male fertility under environmentally constrained conditions, by controlling the mutagenic activity of transposons in male germ cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Sequence data (messenger RNA-seq, nanoPARE-seq, sRNA-seq and LTR-seq) that support the findings of this study have been deposited at the European Nucleotide Archive under accession code ERP118841. Mass spectrometry proteomics data have been deposited at the ProteomeXchange Consortium with the dataset identifier PXD013891.
References
Challinor, A. J. et al. A meta-analysis of crop yield under climate change and adaptation. Nat. Clim. Change 4, 287–291 (2014).
Peng, S. et al. Rice yields decline with higher night temperature from global warming. Proc. Natl Acad. Sci. USA 101, 9971–9975 (2004).
Hedhly, A., Hormaza, J. I. & Herrero, M. Global warming and sexual plant reproduction. Trends Plant Sci. 14, 30–36 (2009).
Lobell, D. B. & Asner, G. P. Climate and management contributions to recent trends in U.S. agricultural yields. Science 299, 1032 (2003).
De Storme, N. & Geelen, D. The impact of environmental stress on male reproductive development in plants: biological processes and molecular mechanisms. Plant Cell Environ. 37, 1–18 (2014).
Barnabas, B., Jager, K. & Feher, A. The effect of drought and heat stress on reproductive processes in cereals. Plant Cell Environ. 31, 11–38 (2008).
Begcy, K. & Dresselhaus, T. Epigenetic responses to abiotic stresses during reproductive development in cereals. Plant Reprod. 31, 343–355 (2018).
Ding, J. et al. A long noncoding RNA regulates photoperiod-sensitive male sterility, an essential component of hybrid rice. Proc. Natl Acad. Sci. USA 109, 2654–2659 (2012).
Fan, Y. et al. PMS1T, producing phased small-interfering RNAs, regulates photoperiod-sensitive male sterility in rice. Proc. Natl Acad. Sci. USA 113, 15144–15149 (2016).
Zhou, H. et al. Photoperiod- and thermo-sensitive genic male sterility in rice are caused by a point mutation in a novel noncoding RNA that produces a small RNA. Cell Res. 22, 649–660 (2012).
Johnson, C. et al. Clusters and superclusters of phased small RNAs in the developing inflorescence of rice. Genome Res. 19, 1429–1440 (2009).
Komiya, R. et al. Rice germline-specific Argonaute MEL1 protein binds to phasiRNAs generated from more than 700 lincRNAs. Plant J. 78, 385–397 (2014).
Araki, S. et al. miR2118-dependent U-rich phasiRNA production in rice anther wall development. Nat. Commun. 11, 3115 (2020).
Song, X. et al. Rice RNA-dependent RNA polymerase 6 acts in small RNA biogenesis and spikelet development. Plant J. 71, 378–389 (2012).
Song, X. et al. Roles of DCL4 and DCL3b in rice phased small RNA biogenesis. Plant J. 69, 462–474 (2012).
Xia, R. et al. 24-nt reproductive phasiRNAs are broadly present in angiosperms. Nat. Commun. 10, 627 (2019).
Das, S., Swetha, C., Pachamuthu, K., Nair, A. & Shivaprasad, P. V. Loss of function of Oryza sativa Argonaute 18 induces male sterility and reduction in phased small RNAs. Plant Reprod. 33, 59–73 (2020).
Nonomura, K. et al. A germ cell-specific gene of the ARGONAUTE family is essential for the progression of premeiotic mitosis and meiosis during sporogenesis in rice. Plant Cell 19, 2583–2594 (2007).
Zhai, J. et al. Spatiotemporally dynamic, cell-type-dependent premeiotic and meiotic phasiRNAs in maize anthers. Proc. Natl Acad. Sci. USA 112, 3146–3151 (2015).
Teng, C. et al. Dicer-like 5 deficiency confers temperature-sensitive male sterility in maize. Nat. Commun. 11, 2912 (2020).
Vernoud, V. et al. The HD-ZIP IV transcription factor OCL4 is necessary for trichome patterning and anther development in maize. Plant J. 59, 883–894 (2009).
Dukowic-Schulze, S. et al. The transcriptome landscape of early maize meiosis. BMC Plant Biol. 14, 118 (2014).
Shiboleth, Y. M. et al. The conserved FRNK box in HC-Pro, a plant viral suppressor of gene silencing, is required for small RNA binding and mediates symptom development. J. Virol. 81, 13135–13148 (2007).
Lakatos, L. et al. Small RNA binding is a common strategy to suppress RNA silencing by several viral suppressors. EMBO J. 25, 2768–2780 (2006).
Lisch, D. How important are transposons for plant evolution? Nat. Rev. Genet. 14, 49–61 (2013).
Ye, J. et al. Proteomic and phosphoproteomic analyses reveal extensive phosphorylation of regulatory proteins in developing rice anthers. Plant J. 84, 527–544 (2015).
Walley, J. W. et al. Integration of omic networks in a developmental atlas of maize. Science 353, 814–818 (2016).
Sheu-Gruttadauria, J., Xiao, Y., Gebert, L. F. & MacRae, I. J. Beyond the seed: structural basis for supplementary microRNA targeting by human Argonaute2. EMBO J. 38, e101153 (2019).
Quevillon Huberdeau, M. et al. Phosphorylation of Argonaute proteins affects mRNA binding and is essential for microRNA-guided gene silencing in vivo. EMBO J. 36, 2088–2106 (2017).
Golden, R. J. et al. An Argonaute phosphorylation cycle promotes microRNA-mediated silencing. Nature 542, 197–202 (2017).
Ernst, C., Odom, D. T. & Kutter, C. The emergence of piRNAs against transposon invasion to preserve mammalian genome integrity. Nat. Commun. 8, 1411 (2017).
Naito, K. et al. Unexpected consequences of a sudden and massive transposon amplification on rice gene expression. Nature 461, 1130–1134 (2009).
Studer, A., Zhao, Q., Ross-Ibarra, J. & Doebley, J. Identification of a functional transposon insertion in the maize domestication gene tb1. Nat. Genet. 43, 1160–1163 (2011).
Carpentier, M. C. et al. Retrotranspositional landscape of Asian rice revealed by 3000 genomes. Nat. Commun. 10, 24 (2019).
Dooner, H. K. et al. Spontaneous mutations in maize pollen are frequent in some lines and arise mainly from retrotranspositions and deletions. Proc. Natl Acad. Sci. USA 116, 10734–10743 (2019).
Peng, Y. et al. Elimination of a retrotransposon for quenching genome instability in modern rice. Mol. Plant 12, 1395–1407 (2019).
Li, Y., Segal, G., Wang, Q. & Dooner, H. K. Gene tagging with engineered Ds elements in maize. Methods Mol. Biol. 1057, 83–99 (2013).
McCarty, D. R. et al. Steady-state transposon mutagenesis in inbred maize. Plant J. 44, 52–61 (2005).
Samalova, M., Brzobohaty, B. & Moore, I. pOp6/LhGR: a stringently regulated and highly responsive dexamethasone-inducible gene expression system for tobacco. Plant J. 41, 919–935 (2005).
Nakagawa, T. et al. Improved gateway binary vectors: high-performance vectors for creation of fusion constructs in transgenic analysis of plants. Biosci. Biotechnol. Biochem. https://doi.org/10.1271/bbb.70216 (2007).
Zhai, J. et al. Spatiotemporally dynamic, cell-type-dependent premeiotic and meiotic phasiRNAs in maize anthers. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.1418918112 (2015).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. https://doi.org/10.1186/gb-2013-14-4-r36 (2013).
Jin, Y., Tam, O. H., Paniagua, E. & Hammell, M. TEtranscripts: a package for including transposable elements in differential expression analysis of RNA-seq datasets. Bioinformatics https://doi.org/10.1093/bioinformatics/btv422 (2015).
Schon, M. A., Kellner, M. J., Plotnikova, A., Hofmann, F. & Nodine, M. D. NanoPARE: parallel analysis of RNA 5′ ends from low-input RNA. Genome Res. https://doi.org/10.1101/gr.239202.118 (2018).
MARTIN, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Allen, E. et al. Evolution of microRNA genes by inverted duplication of target gene sequences in Arabidopsis thaliana. Nat. Genet. 36, 1282–1290 (2004).
Wiśniewski, J. R., Zougman, A., Nagaraj, N. & Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods https://doi.org/10.1038/nmeth.1322 (2009).
Thingholm, T. E., Jørgensen, T. J. D., Jensen, O. N. & Larsen, M. R. Highly selective enrichment of phosphorylated peptides using titanium dioxide. Nat. Protoc. https://doi.org/10.1038/nprot.2006.185 (2006).
Acknowledgements
We thank G. Grant and P. Watson for help with plant husbandry, L. M. Costa for discussions and comments on the manuscript and N. Springer and S. Anderson for help with transposon analysis. This research was supported by awards from the US National Science Foundation (nos. 1027445 to A.W.S. and 1649424 and 1754097 to B.C.M.), European Research Council (grant no. 637888 to M.D.N.) and Biotechnology and Biological Sciences Research Council (BBSRC) (nos. BB/L003023/1, BB/N005279/1, BB/N00194X/1 and BB/P02601X/1 to J.G.-M).
Author information
Authors and Affiliations
Contributions
Y.-S.L. and R.M. cultivated plants, harvested samples and collected phenotypic data. R.P., J.G.-M. and J.R. identified transposon insertions. J.D. performed phosphoproteome analysis. R.M. and Y.-S.L. performed immunoprecipitation and sRNA-seq libraries. A.L. and J.C.L. generated constructs and transgenic lines for HC-Pro and HDZIV6 transactivation. R.P. constructed nanoPARE libraries. R.M., A.D., S.T., S.A. and M.D.N. performed bioinformatic analysis. C.d.G. performed molecular dynamics simulation and protein modelling. Y.-S.L., R.M., J.D., A.D., S.T. and C.d.G. prepared figures and tables. A.W.S., J.B., B.C.M., M.D.N., J.R. and J.G.-M. co-ordinated experiments. Y.-S.L., R.M. and J.G.-M. conceived the project. J.G.-M. wrote the manuscript with input from the rest of the authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–13 and source data with supporting blots and/or gels.
Supplementary Table 1
List of differentially expressed Hphasi loci.
Supplementary Table 2
List of differentially expressed genes in pre-meiotic anthers from wild-type and MAGOKD plants.
Supplementary Table 3
List of differentially expressed transposons in pre-meiotic anthers from wild-type and MAGOKD plants.
Supplementary Table 4
List of predicted retrotransposons (LTRs) targeted by Hphasi.
Supplementary Table 5
List of new transposon insertions determined by LTR-seq.
Supplementary Table 6
List of differentially accumulated phosphopeptides from pre-meiotic anthers of wild-type plants exposed to heat stress.
Supplementary Table 7
Conservation of phosphorylated serine and threonine residues in different Argonautes.
Supplementary Table 8
List of oligonucleotides and synthetic DNA constructs.
Supplementary Table 9
Next-generation sequencing library details.
Supplementary Video 1
Electrostatic potential distribution on the molecular surface of the central cleft of native and phosphorylated MAGO2.
Rights and permissions
About this article
Cite this article
Lee, YS., Maple, R., Dürr, J. et al. A transposon surveillance mechanism that safeguards plant male fertility during stress. Nat. Plants 7, 34–41 (2021). https://doi.org/10.1038/s41477-020-00818-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-020-00818-5
This article is cited by
-
Molecular Mechanisms of Male Sterility in Maize
Plant Molecular Biology Reporter (2024)
-
Spatial distribution of three ARGONAUTEs regulates the anther phasiRNA pathway
Nature Communications (2023)
-
The role of LTR retrotransposons in plant genetic engineering: how to control their transposition in the genome
Plant Cell Reports (2023)
-
Mobile ARGONAUTE 1d binds 22-nt miRNAs to generate phasiRNAs important for low-temperature male fertility in rice
Science China Life Sciences (2023)