Abstract
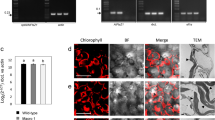
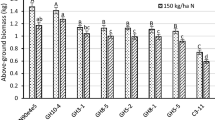
High accumulation of heterologous proteins expressed from the plastid genome has sometimes been reported to result in compromised plant phenotypes. Comparisons of transplastomic plants to wild-type (WT) are typically made in environmentally controlled chambers with relatively low light; little is known about the performance of such plants under field conditions. Here, we report on two plastid-engineered tobacco lines expressing the bacterial cellulase Cel6A. Field-grown plants producing Cel6A at ~20% of total soluble protein exhibit no loss in biomass or Rubisco content and only minor reductions in photosynthesis compared to WT. These experiments demonstrate that, when grown in the field, tobacco possesses sufficient metabolic flexibility to accommodate high levels of recombinant protein by increasing total protein synthesis and accumulation and/or by reallocating unneeded endogenous proteins. Based on current tobacco cultivation practices and readily achievable recombinant protein yields, we estimate that specific proteins could be obtained from field-grown transgenic tobacco plants at costs three orders of magnitude less than current cell culture methods.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data generated or analysed during this study are included in this published article and its Supplementary information.
References
Palomares, L. A., Kuri-Breña, F. & Ramírez, O. T. in Biotechnology Vol. 5 (eds Doelle, H., Rokem, J. & Berovic, M.) Ch. 43 (EOLSS Publishers, 2002).
Twyman, R. M., Stoger, E., Schillberg, S., Christou, P. & Fischer, R. Molecular farming in plants: host systems and expression technology. Trends Biotechnol. 21, 570–578 (2003).
Doran, P. M. Foreign protein production in plant tissue cultures. Curr. Opin. Biotechnol. 11, 199–204 (2000).
Xu, J., Towler, M. & Weathers, P. in Bioprocessing of Plant in Vitro Systems (eds Pavlov, A. & Bley, T.) 1–40 (Springer, 2016).
Shaaltiel, Y. et al. Production of glucocerebrosidase with terminal mannose glycans for enzyme replacement therapy of Gaucher’s disease using a plant cell system. Plant Biotechnol. J. 5, 579–590 (2007).
Zimran, A. et al. Pivotal trial with plant cell-expressed recombinant glucocerebrosidase, taliglucerase alfa, a novel enzyme replacement therapy for Gaucher disease. Blood 118, 5767–5773 (2011).
Yao, J., Weng, Y., Dickey, A. & Wang, K. Y. Plants as factories for human pharmaceuticals: applications and challenges. Int. J. Mol. Sci. 16, 28549–28565 (2015).
Holtz, B. R. et al. Commercial‐scale biotherapeutics manufacturing facility for plant‐made pharmaceuticals. Plant Biotechnol. J. 13, 1180–1190 (2015).
Farsad, A., Malekzadeh-Shafaroudi, S., Zibaee, S., Fotouhi, F. & Moshtaghi, N. Transient expression of HA1 antigen of H5N1 influenza virus in Tob. (Nicotiana tab. L.) via agro-infiltration. J. Agr. Sci. Tech. 19, 439–451 (2018).
Fischer, R., Vasilev, N., Twyman, R. M. & Schillberg, S. High-value products from plants: the challenges of process optimization. Curr. Opin. Biotechnol. 32, 156–162 (2015).
Bock, R. Engineering plastid genomes: methods, tools, and applications in basic research and biotechnology. Annu. Rev. Plant Biol. 66, 211–241 (2015).
Svab, Z. & Maliga, P. Exceptional transmission of plastids and mitochondria from the transplastomic pollen parent and its impact on transgene containment. Proc. Natl Acad. Sci. USA 104, 7003–7008 (2007).
Golczyk, H. et al. Chloroplast DNA in mature and senescing leaves: a reappraisal. Plant Cell 26, 847–854 (2014).
Shaver, J. M., Oldenburg, D. J. & Bendich, A. J. Changes in chloroplast DNA during development in tobacco, Medicago truncatula, pea, and maize. Planta 224, 72–82 (2006).
Leelavathi, S. & Reddy, V. S. Chloroplast expression of His-tagged GUS-fusions: a general strategy to overproduce and purify foreign proteins using transplastomic plants as bioreactors. Mol. Breed. 11, 49–58 (2003).
Staub, J. M. & Maliga, P. Long regions of homologous DNA are incorporated into the tobacco plastid genome by transformation. Plant Cell 4, 39–45 (1992).
Oey, M., Lohse, M., Kreikemeyer, B. & Bock, R. Exhaustion of the chloroplast protein synthesis capacity by massive expression of a highly stable protein antibiotic. Plant J. 57, 436–445 (2009).
Long, S., Humphries, S. & Falkowski, P. G. Photoinhibition of photosynthesis in nature. Annu. Rev. Plant Biol. 45, 633–662 (1994).
Arlen, P. A. et al. Field production and functional evaluation of chloroplast‐derived interferon‐α2b. Plant Biotechnol. J. 5, 511–525 (2007).
Bhat, M. Cellulases and related enzymes in biotechnology. Biotechnol. Adv. 18, 355–383 (2000).
Wilson, D. B. Studies of Thermobifida fusca plant cell wall degrading enzymes. Chem. Rec. 4, 72–82 (2004).
Johnson, E. Integrated enzyme production lowers the cost of cellulosic ethanol. Biofuel. Bioprod. Biorefin. 10, 164–174 (2016).
Kuroda, H. & Maliga, P. Sequences downstream of the translation initiation codon are important determinants of translation efficiency in chloroplasts. Plant Physiol. 125, 430–436 (2001).
Gray, B. N., Ahner, B. A. & Hanson, M. R. High-level bacterial cellulase accumulation in chloroplast-transformed tobacco mediated by downstream box fusions. Biotechnol. Bioeng. 102, 1045–1054 (2009).
De Cosa, B., Moar, W., Lee, S.-B., Miller, M. & Daniell, H. Overexpression of the Bt Cry2Aa2 operon in chloroplasts leads to formation of insecticidal crystals. Nat. Biotechnol. 19, 71 (2001).
Maliga, P. Plastid transformation in higher plants. Annu. Rev. Plant Biol. 55, 289–313 (2004).
Hirose, T. & Sugiura, M. Cis-acting elements and trans-acting factors for accurate translation of chloroplast psbA mRNAs: development of an in vitro translation system from tobacco chloroplasts. EMBO J. 15, 1687–1695 (1996).
Kuroda, H. & Maliga, P. Overexpression of the clpP 5′-untranslated region in a chimeric context causes a mutant phenotype, suggesting competition for a clpP-specific RNA maturation factor in tobacco chloroplasts. Plant Physiol. 129, 1600–1606 (2002).
Scotti, N. & Cardi, T. Transgene-induced pleiotropic effects in transplastomic plants. Biotechnol. Lett. 36, 229–239 (2014).
Bally, J., Job, C., Belghazi, M. & Job, D. Metabolic adaptation in transplastomic plants massively accumulating recombinant proteins. PLoS ONE 6, e25289 (2011).
Bally, J. et al. Plant physiological adaptations to the massive foreign protein synthesis occurring in recombinant chloroplasts. Plant Physiol. 150, 1474–1481 (2009).
Foreman, L. Tobacco Production Costs and Returns in 2004 (US Department of Agriculture, Economic Research Service, 2006).
Scott, G. L. & Warren, J. F. Tobacco Production System (Google Patents, 2016).
Liu, G., Zhang, J. & Bao, J. Cost evaluation of cellulase enzyme for industrial-scale cellulosic ethanol production based on rigorous Aspen Plus modeling. Bioprocess Biosyst. Eng. 39, 133–140 (2016).
Wilken, L. R. & Nikolov, Z. L. Recovery and purification of plant-made recombinant proteins. Biotechnol. Adv. 30, 419–433 (2012).
Rubio-Infante, N. et al. A chloroplast-derived C4V3 polypeptide from the human immunodeficiency virus (HIV) is orally immunogenic in mice. Plant Mol. Biol. 78, 337–349 (2012).
Stephan, A. et al. Simple purification of Nicotiana benthamiana-produced recombinant colicins: high-yield recovery of purified proteins with minimum alkaloid content supports the suitability of the host for manufacturing food additives. Int. J. Mol. Sci. 19, 95 (2017).
Farquhar, Gv, von Caemmerer, Sv & Berry, J. A biochemical model of photosynthetic CO2 assimilation in leaves of C3 species. Planta 149, 78–90 (1980).
Bernacchi, C., Singsaas, E., Pimentel, C., Portis, A. Jr & Long, S. Improved temperature response functions for models of Rubisco‐limited photosynthesis. Plant Cell Environ. 24, 253–259 (2001).
McMurtrie, R. & Wang, Y. P. Mathematical models of the photosynthetic response of tree stands to rising CO2 concentrations and temperatures. Plant Cell Environ. 16, 1–13 (1993).
Caspi, J. et al. Thermobifida fusca exoglucanase Cel6B is incompatible with the cellulosomal mode in contrast to endoglucanase Cel6A. Syst. Synth. Biol. 4, 193–201 (2010).
Caspi, J. et al. Thermobifida fusca family-6 cellulases as potential designer cellulosome components. Biocatal. Biotransformation 24, 3–12 (2006).
Lin, M. T. & Hanson, M. R. Red algal Rubisco fails to accumulate in transplastomic tobacco expressing Griffithsia monilis RbcL and RbcS genes. Plant Direct 2, 1–10 (2018).
Lin, M. T., Stone, W. D., Chaudhari, V. & Hanson, M. R. Enzyme kinetics of tobacco Rubisco expressed in Escherichia coli varies depending on the small subunit composition. Preprint at https://www.biorxiv.org/content/10.1101/562223v1 (2019).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671 (2012).
Acknowledgements
We thank L.V. Richter for providing advice and labororatory training to J.A.S. We also thank D. Drag and B. Harbaugh for managing the field sites and cultivating the plants.
Author information
Authors and Affiliations
Contributions
J.M.M. and S.P.L. supervised planting and harvesting of growth chamber and field trials and made photosynthetic measurements. J.A.S. performed immunoblots, performed statistical analyses and produced all figures. S.P.L., M.R.H. and B.A.A. conceived of the project. All authors participated in writing the manuscript. All authors planned research and discussed results. This work is supported in part by the AFRI NIFA Fellowships Grant Program (2018-67011-28027/project accession No. 1015534) awarded to J.A.S. from the USDA National Institute of Food and Agriculture. Any opinions, findings, conclusions or recommendations expressed in this publication are those of the author(s) and do not necessarily reflect the view of the US Department of Agriculture.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Nature Plants thanks Francis Quétier and other, anonymous, reviewers for their contribution to the peer review of this work.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2, and Supplementary Tables 1–4.
Supplementary Datasets
Data generated for Figs. 1b–d, 2c,e and 3 and Supplementary Figs. 1a–c and 2.
Rights and permissions
About this article
Cite this article
Schmidt, J.A., McGrath, J.M., Hanson, M.R. et al. Field-grown tobacco plants maintain robust growth while accumulating large quantities of a bacterial cellulase in chloroplasts. Nat. Plants 5, 715–721 (2019). https://doi.org/10.1038/s41477-019-0467-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-019-0467-z
This article is cited by
-
Mitigation of deleterious phenotypes in chloroplast-engineered plants accumulating high levels of foreign proteins
Biotechnology for Biofuels (2021)
-
Targeted genome editing of plants and plant cells for biomanufacturing
Transgenic Research (2021)
-
Small subunits can determine enzyme kinetics of tobacco Rubisco expressed in Escherichia coli
Nature Plants (2020)
-
Plant-Based Cellulase Assay Systems as Alternatives for Synthetic Substrates
Applied Biochemistry and Biotechnology (2020)
-
Molecular, biochemical, and proteomic analyses of transplastomic tobacco plants expressing an endoglucanase support chloroplast-based molecular farming for industrial scale production of enzymes
Applied Microbiology and Biotechnology (2019)