Abstract
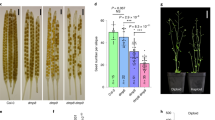
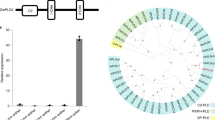
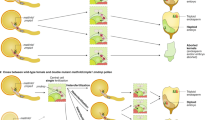
Doubled haploid (DH) breeding based on in vivo haploid induction has led to a new approach for maize breeding1. All modern haploid inducers used in DH breeding are derived from the haploid inducer line Stock6. Two key quantitative trait loci, qhir1 and qhir8, lead to high-frequency haploid induction2. Mutation of the gene MTL/ZmPLA1/NLD in qhir1 could generate a ~2% haploid induction rate (HIR)3,4,5; nevertheless, this mutation is insufficient for modern haploid inducers whose average HIR is ~10%6. Therefore, cloning of the gene underlying qhir8 is important for illuminating the genetic basis of haploid induction. Here, we present the discovery that mutation of a non-Stock6-originating gene in qhir8, namely, ZmDMP, enhances and triggers haploid induction. ZmDMP was identified by map-based cloning and further verified by CRISPR–Cas9-mediated knockout experiments. A single-nucleotide change in ZmDMP leads to a 2–3-fold increase in the HIR. ZmDMP knockout triggered haploid induction with a HIR of 0.1–0.3% and exhibited a greater ability to increase the HIR by 5–6-fold in the presence of mtl/zmpla1/nld. ZmDMP was highly expressed during the late stage of pollen development and localized to the plasma membrane. These findings provide important approaches for studying the molecular mechanism of haploid induction and improving DH breeding efficiency in maize.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data sets generated and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Ren, J. et al. Novel technologies in doubled haploid line development. Plant Biotechnol. J. 15, 1361–1370 (2017).
Prigge, V. et al. New insights into the genetics of in vivo induction of maternal haploids, the backbone of doubled haploid technology in maize. Genetics 190, 781–793 (2012).
Kelliher, T. et al. MATRILINEAL, a sperm-specific phospholipase, triggers maize haploid induction. Nature 542, 105–109 (2017).
Liu, C. et al. A 4-bp insertion at ZmPLA1 encoding a putative phospholipase A generates haploid induction in maize. Mol. Plant 10, 520–522 (2017).
Gilles, L. M. et al. Loss of pollen‐specific phospholipase NOT LIKE DAD triggers gynogenesis in maize. EMBO J. 36, e201796603 (2017).
Hu, H. et al. The genetic basis of haploid induction in maize identified with a novel genome-wide association method. Genetics 202, 1267–1276 (2016).
Coe, E. H. A line of maize with high haploid frequency. Am. Nat. 93, 381–382 (1959).
Lashermes, P., Gaillard, A. & Beckert, M. Gynogenetic haploid plants analysis for agronomic and enzymatic markers in maize (Zea mays L.). Theor. Appl. Genet. 76, 570–572 (1988).
Dong, X. et al. Fine mapping of qhir1 influencing in vivo haploid induction in maize. Theor. Appl. Genet. 126, 1713–1720 (2013).
Jackson, D. No sex please, we’re (in) breeding. EMBO J. 36, 703–704 (2017).
Liu, C. et al. Fine mapping of qhir8 affecting in vivo haploid induction in maize. Theor. Appl. Genet. 128, 2507–2515 (2015).
Yao, L. et al. OsMATL mutation induces haploid seed formation in indica rice. Nat. Plants 4, 530–533 (2018).
Xing, H. et al. A CRISPR/Cas9 toolkit for multiplex genome editing in plants. BMC Plant Biol. 14, 327 (2014).
Yang, Q., Zhang, D. & Xu, M. A sequential quantitative trait locus fine mapping strategy using recombinant-derived progeny. J. Integr. Plant Biol. 54, 228–237 (2012).
Xu, X. et al. Gametophytic and zygotic selection leads to segregation distortion through in vivo induction of a maternal haploid in maize. J. Exp. Bot. 64, 1083–1096 (2013).
Jefferson, R. A., Kavanagh, T. A. & Bevan, M. W. GUS fusions: β‐glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J. 6, 3901–3907 (1987).
Widholm, J. M. The use of fluorescein diacetate and phenosafranine for determining viability of cultured plant cells. Stain Technol. 47, 189–194 (1972).
Herrero, M. P. & Johnson, R. R. High temperature stress and pollen viability of maize. Crop Sci. 20, 796–800 (1980).
Acknowledgements
We acknowledge support from the National Key Research and Development Program of China—Maize heterosis utilization technology and strong heterosis hybrids breeding (2016YFD0101200, 2016YFD0101003 and 2018YFD0100201)—and the Modern Maize Industry Technology System (CARS–02–04).
Author information
Authors and Affiliations
Contributions
S.C., Y.Z. and C.L. conceived and designed the experiments. Y.Z., C.L., X.Q. and Y.J. performed the experiments. Y.Z., S.C., C.L. and X.Q. analysed the data. C.L., S.C., Y.Z., D.W., Y.W., Z.L., C.C., B.C. X.T., J.Li, M.C., X.D., X.X., L.L., W.Li., W.Liu., W.J. and J.Lai. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Nature Plants thanks Jose Seguí-Simarro and other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–6 and Supplementary Tables 1–4
Rights and permissions
About this article
Cite this article
Zhong, Y., Liu, C., Qi, X. et al. Mutation of ZmDMP enhances haploid induction in maize. Nat. Plants 5, 575–580 (2019). https://doi.org/10.1038/s41477-019-0443-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-019-0443-7
This article is cited by
-
One-step creation of CMS lines using a BoCENH3-based haploid induction system in Brassica crop
Nature Plants (2024)
-
Potential gene editing targets for developing haploid inducer stocks in rice and wheat with high haploid induction frequency
3 Biotech (2024)
-
Mechanism and molecular basis of apomixis in angiosperms
Vegetos (2024)
-
Gene editing tool kit in millets: present status and future directions
The Nucleus (2024)
-
Applications of CRISPR/Cas genome editing in economically important fruit crops: recent advances and future directions
Molecular Horticulture (2023)