Abstract
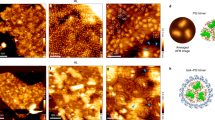
Little is known about how the photosynthetic machinery is arranged in time and space during the biogenesis of thylakoid membranes. Using in situ cryo-electron tomography to image the three-dimensional architecture of the cyanobacterium Synechocystis, we observed that the tips of multiple thylakoids merge to form a substructure called the ‘convergence membrane’. This high-curvature membrane comes into close contact with the plasma membrane at discrete sites. We generated subtomogram averages of 70S ribosomes and array-forming phycobilisomes, then mapped these structures onto the native membrane architecture as markers for protein synthesis and photosynthesis, respectively. This molecular localization identified two distinct biogenic regions in the thylakoid network: thylakoids facing the cytosolic interior of the cell that were associated with both marker complexes, and convergence membranes that were decorated by ribosomes but not phycobilisomes. We propose that the convergence membranes perform a specialized biogenic function, coupling the synthesis of thylakoid proteins with the integration of cofactors from the plasma membrane and the periplasmic space.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
References
Nelson, N. & Junge, W. Structure and energy transfer in photosystems of oxygenic photosynthesis. Annu. Rev. Biochem. 84, 659–683 (2015).
Eberhard, S., Finazzi, G. & Wollmann, F.-A. The dynamics of photosynthesis. Annu. Rev. Genet. 42, 463–515 (2008).
Umena, Y., Kawakami, K., Shen, J. R. & Kamiya, N. Crystal structure of oxygen-evolving photosystem II at a resolution of 1.9 Å. Nature 473, 55–60 (2011).
Hasan, S. S., Yamashita, E., Baniulis, D. & Cramer, W. A. Quinone-dependent proton transfer pathways in the photosynthetic cytochrome b6f complex. Proc. Natl Acad. Sci. USA 110, 4297–4302 (2013).
Jordan, P. et al. Three-dimensional structure of cyanobacterial photosystem I at 2.5 Å resolution. Nature 411, 909–917 (2001).
Heinz, S., Liauw, P., Nickelsen, J. & Nowaczyk, M. Analysis of photosystem II biogenesis in cyanobacteria. Biochim. Biophys. Acta 1857, 274–287 (2016).
Nixon, P. J., Michoux, F., Yu, J., Boehm, M. & Komenda, J. Recent advances in understanding the assembly and repair of photosystem II. Ann. Bot. 106, 1–16 (2010).
Nickelsen, J. & Rengstl, B. Photosystem II assembly: from cyanobacteria to plants. Annu. Rev. Plant Biol. 64, 609–635 (2013).
Komenda, J., Sobotka, R. & Nixon, P. J. Assembling and maintaining the photosystem II complex in chloroplasts and cyanobacteria. Curr. Opin. Plant Biol. 15, 245–251 (2012).
Rast, A., Rengstl, B., Heinz, S., Klingl, A. & Nickelsen, J. The role of Slr0151, a tetratricopeptide repeat protein from Synechocystis sp. PCC 6803, during photosystem II assembly and repair. Front. Plant Sci. 7, 605 (2016).
Sacharz, J. et al. Sub-cellular location of FtsH proteases in the cyanobacterium Synechocystis sp. PCC 6803 suggests localised PSII repair zones in the thylakoid membranes. Mol. Microbiol. 96, 448–462 (2015).
Stengel, A. et al. Initial steps of photosystem II de novo assembly and preloading with manganese take place in biogenesis centers in Synechocystis. Plant Cell 24, 660–675 (2012).
Liberton, M., Howard Berg, R., Heuser, J., Roth, R. & Pakrasi, H. B. Ultrastructure of the membrane systems in the unicellular cyanobacterium Synechocystis sp. strain PCC 6803. Protoplasma 227, 129–138 (2006).
Nierzwicki-Bauer, S. A., Balkwill, D. L. & Stevens, S. E. Three-dimensional ultrastructure of a unicellular cyanobacterium. J. Cell Biol. 97, 713–722 (1983).
van de Meene, A. M., Hohmann-Marriott, M. F., Vermaas, W. F. & Roberson, R. W. The three-dimensional structure of the cyanobacterium Synechocystis sp. PCC 6803. Arch. Microbiol. 184, 259–270 (2006).
Armbruster, U. et al. Arabidopsis CURVATURE THYLAKOID1 proteins modify thylakoid architecture by inducing membrane curvature. Plant Cell 25, 2661–2678 (2013).
Heinz, S. et al. Thylakoid membrane architecture in Synechocystis depends on CurT, a homolog of the granal CURVATURE THYLAKOID1 proteins. Plant Cell 28, 2238–2260 (2016).
Kunkel, D. D. Thylakoid centers: structures associated with the cyanobacterial photosynthetic membrane system. Arch. Microbiol. 133, 97–99 (1982).
Engel, B. D. et al. Native architecture of the Chlamydomonas chloroplast revealed by in situ cryo-electron tomography. eLife 4, e04889 (2015).
Tauschel, H. D. & Drews, G. Thylakoidmorphogenese bei Rhodopseudomonas palustris. Archiv Mikrobiologie 59, 381–404 (1967).
Remsen, C. C., Watson, S. W., Waterbury, J. B. & Trüper, H. G. Fine structure of Ectothiorhodospira mobilis Pelsh. J. Bacteriol. 95, 2374–2392 (1968).
Noble, J. M. et al. Connectivity of centermost chromatophores in Rhodobacter sphaeroides bacteria. Mol. Microbiol. 109, 812–825 (2018).
Arteni, A. A., Ajlani, G. & Boekema, E. J. Structural organisation of phycobilisomes from Synechocystis sp. strain PCC6803 and their interaction with the membrane. Biochim. Biophys. Acta 1787, 272–279 (2009).
Chang, L. et al. Structural organization of an intact phycobilisome and its association with photosystem II. Cell Res. 25, 726–737 (2015).
Tang, K. et al. The terminal phycobilisome emitter, LCM: A light-harvesting pigment with a phytochrome chromophore. Proc. Natl Acad. Sci. USA 112, 15880–15885 (2015).
Harris, D., Bar-Zvi, S., Lahav, A., Goldshmid, I. & Adir, N. in Membrane Protein Complexes: Structure and Function Vol. 87 (eds Harris, J. R. & Boekema, E. J.) 57–82 (Springer Singapore, 2018).
Rigort, A. et al. Focused ion beam micromachining of eukaryotic cells for cryoelectron tomography. Proc. Natl Acad. Sci. USA 109, 4449–4454 (2012).
Schaffer, M. et al. Optimized cryo-focused ion beam sample preparation aimed at in situ structural studies of membrane proteins. J. Struct. Biol. 197, 73–82 (2017).
Asano, S., Engel, B. D. & Baumeister, W. In situ cryo-electron tomography: a post-reductionist approach to structural biology. J. Mol. Biol. 428, 332–343 (2016).
Kirchhoff, H. et al. Dynamic control of protein diffusion within the granal thylakoid lumen. Proc. Natl Acad. Sci. USA 108, 20248–20253 (2011).
Danev, R., Buijsse, B., Khoshouei, M., Plitzko, J. M. & Baumeister, W. Volta potential phase plate for in-focus phase contrast transmission electron microscopy. Proc. Natl Acad. Sci. USA 111, 15635–15640 (2014).
Brandt, F. et al. The native 3D organization of bacterial polysomes. Cell 136, 261–271 (2009).
Ortiz, J. O. et al. Structure of hibernating ribosomes studied by cryoelectron tomography in vitro and in situ. J. Cell Biol. 190, 613–621 (2010).
Beckert, B. et al. Structure of the Bacillus subtilis hibernating 100S ribosome reveals the basis for 70S dimerization. EMBO J. 36, 2061 (2017).
Beckert, B. et al. Structure of a hibernating 100S ribosome reveals an inactive conformation of the ribosomal protein S1. Nat. Microbiol. 3, 1115–1121 (2018).
Flygaard, R. K., Boegholm, N., Yusupov, M. & Jenner, L. B. Cryo-EM structure of the hibernating Thermus thermophilus 100S ribosome reveals a protein-mediated dimerization mechanism. Nat. Commun. 9, 4179 (2018).
Matzov, D. et al. The cryo-EM structure of hibernating 100S ribosome dimer from pathogenic Staphylococcus aureus. Nat. Commun. 8, 723 (2017).
Nevo, R. et al. Thylakoid membrane perforations and connectivity enable intracellular traffic in cyanobacteria. EMBO J. 26, 1467–1473 (2007).
Hinterstoisser, B., Cichna, M., Kuntner, O. & Peschek, G. A. Cooperation of plasma and thylakoid membranes for the biosynthesis of chlorophyll in cyanobacteria: the role of the thylakoid centers. J. Plant Physiol. 142, 407–413 (1993).
Frain, K. M., Gangl, D., Jones, A., Zedler, J. A. Z. & Robinson, C. Protein translocation and thylakoid biogenesis in cyanobacteria. Biochim. Biophys. Acta 1857, 266–273 (2016).
Zak, E. et al. The initial steps of biogenesis of cyanobacterial photosystems occur in plasma membranes. Proc. Natl Acad. Sci. USA 98, 13443–13448 (2001).
van de Meene, A. M. et al. Gross morphological changes in thylakoid membrane structure are associated with photosystem I deletion in Synechocystis sp. PCC 6803. Biochim. Biophys. Acta 1818, 1427–1434 (2012).
Junglas, B. & Schneider, D. What is Vipp1 good for? Mol. Microbiol. 108, 1–5 (2018).
Gutu, A., Chang, F. & O’Shea, E. K. Dynamical localization of a thylakoid membrane binding protein is required for acquisition of photosynthetic competency. Mol. Microbiol. 108, 16–31 (2018).
Hennig, R. et al. IM30 triggers membrane fusion in cyanobacteria and chloroplasts. Nat. Commun. 6, 7018 (2015).
Saur, M. et al. A Janus-faced IM30 ring involved in thylakoid membrane fusion is assembled from IM30 tetramers. Structure 25, 1380–1390 (2017).
Marx, A. & Adir, N. Allophycocyanin and phycocyanin crystal structures reveal facets of phycobilisome assembly. Biochim. Biophys. Acta 1827, 311–318 (2013).
Glazer, A. N. Light harvesting by phycobilisomes. Annu. Rev. Biophys. Biophys. Chem. 14, 47–77 (1985).
Zhang, J. et al. Structure of phycobilisome from the red alga Griffithsia pacifica. Nature 551, 57–63 (2017).
Olive, J., Ajlani, G., Astier, C., Recouvreur, M. & Vernotte, C. Ultrastructure and light adaptation of phycobilisome mutants of Synechocystis PCC 6803. Biochimi. Biophys. Acta 1319, 275–282 (1997).
Westermann, M., Neuschaefer-Rube, O., Morschel, E. & Wehrmeyer, W. Trimeric photosystem I complexes exist in vivo in thylakoid membranes of the Synechocystis strain B09201 and differ in absorption characteristics from monomeric photosystem I complexes. J. Plant Physiol. 155, 24–33 (1999).
Giddings, T. H., Wasmann, C. & Staehelin, L. A. Structure of the thylakoids and envelope membranes of the cyanelles of cyanophora paradoxa. Plant Physiol. 71, 409–419 (1983).
MacGregor-Chatwin, C. et al. Lateral segregation of photosystem I in cyanobacterial thylakoids. Plant Cell 29, 1119–1136 (2017).
Rippka, R., Deruelles, J., Waterbury, J. B., Herdman, M. & Stanier, R. Y. Generic assignments, strain histories and properties of pure cultures of cyanobacteria. Microbiology 111, 1–61 (1979).
Schaffer, M. et al. Cryo-focused ion beam sample preparation for imaging vitreous cells by cryo-electron tomography. Bio Protoc. 5, e1575 (2015).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Xiong, Q., Morphew, M. K., Schwartz, C. L., Hoenger, A. H. & Mastronarde, D. N. CTF determination and correction for low dose tomographic tilt series. J. Struct. Biol. 168, 378–387 (2009).
Grant, T. & Grigorieff, N. Measuring the optimal exposure for single particle cryo-EM using a 2.6 Å reconstruction of rotavirus VP6. eLife 4, e06980 (2015).
Schur, F. K. et al. An atomic model of HIV-1 capsid-SP1 reveals structures regulating assembly and maturation. Science 353, 506–508 (2016).
Hrabe, T. et al. PyTom: a python-based toolbox for localization of macromolecules in cryo-electron tomograms and subtomogram analysis. J. Struct. Biol. 178, 177–188 (2012).
Wan, W. et al. Structure and assembly of the Ebola virus nucleocapsid. Nature 551, 394 (2017).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013).
Villa, E. et al. Ribosome-induced changes in elongation factor Tu conformation control GTP hydrolysis. Proc. Natl Acad. Sci. USA 106, 1063–1068 (2009).
Goddard, T. D., Huang, C. C. & Ferrin, T. E. Visualizing density maps with UCSF Chimera. J. Struct. Biol. 157, 281–287 (2007).
Chen, Y., Pfeffer, S., Hrabe, T., Schuller, J. M. & Forster, F. Fast and accurate reference-free alignment of subtomograms. J. Struct. Biol. 182, 235–245 (2013).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Acknowledgements
The authors thank W. Baumeister for enabling this project by providing support and instrumentation, D. Tegunov for creating the membranogram software, R. Danev for help with the phase plate, as well as S. Heinz and G. Weiss for discussion. This work was supported by grants awarded to B.D.E. (EN 1194/1-1) and J.N. (NI 390/9-2) by the Deutsche Forschungsgemeinschaft in the context of the Research Unit FOR2092. Additional funding was provided by the Max Planck Society and by LMU Munich’s Institutional Strategy “LMU Excellent” within the framework of the German Excellence Initiative.
Author information
Authors and Affiliations
Contributions
A.R. and M.S. cultured and froze cells. M.S. performed the FIB milling. A.R., M.S., S.A. and B.D.E. acquired the tomograms. A.R. performed all the analyses in the paper, with assistance from the following people: W.W. guided subtomogram averaging using STOPGAP, producing the phycobilisome structure shown in Fig. 3; S.A. guided subtomogram averaging using PyTom, producing the phycobilisome structure shown in Fig. 4; S.P. helped with the averaging and interpretation of the ribosome structure shown in Fig. 5; and F.B. provided additional computational guidance. J.M.P. enabled instrumentation and provided funding. J.N. and B.D.E. provided funding and conceived and supervised the project. A.R., J.N. and B.D.E. wrote the paper, with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Journal peer review information Nature Plants thanks Matthew Johnson and other anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–4, Supplementary Table 1 and Supplementary Video Legends.
Supplementary Video 1
In situ cryo-ET of the Synechocystis thylakoid convergence zone (overview 1). Accompanies Fig. 1a and b. The movie slices back and forth through the tomographic volume from Fig. 1a, then reveals the 3D segmentation and mapped in macromolecules shown in Fig. 1b. First, the cytosolic ribosomes (grey) are hidden to show the biogenic regions of the thylakoid network (green), decorated with membrane-associated ribosomes (pink). Photosynthetic regions are decorated with phycobilisomes (yellow). Later, the outer membrane (light blue), plasma membrane (dark blue) and associated macromolecules are hidden for a close-up view of the convergence membrane and thylapse contact.
Supplementary Video 2
In situ cryo-ET of the Synechocystis thylakoid convergence zone (overview 2). Accompanies Fig. 1e and f. The movie slices back and forth through the tomographic volume from Fig. 1e, then reveals the 3D segmentation and mapped in macromolecules shown in Fig. 1f. First, the cytosolic ribosomes (grey) are hidden to show the biogenic regions of the thylakoid network (green), decorated with membrane-associated ribosomes (pink). Photosynthetic regions are decorated with phycobilisomes (yellow). Later, the outer membrane (light blue), plasma membrane (dark blue) and associated macromolecules are hidden for a close-up view of the convergence membrane and thylapse contact.
Supplementary Video 3
Close-up views of convergence zone and thylapse architecture. Accompanies Fig. 2. The movie slices back and forth through tomographic volumes corresponding to Fig. 2a,d,g–i,k and l. An extra volume is shown from the Fig. 2k tomogram in a region not displayed in the figure that contains an exemplary convergence membrane, but not a thylapse. For each volume, it is indicated whether defocus or the Volta phase plate was used to generate contrast.
Rights and permissions
About this article
Cite this article
Rast, A., Schaffer, M., Albert, S. et al. Biogenic regions of cyanobacterial thylakoids form contact sites with the plasma membrane. Nat. Plants 5, 436–446 (2019). https://doi.org/10.1038/s41477-019-0399-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-019-0399-7
This article is cited by
-
Unveiling metabolic pathways involved in the extreme desiccation tolerance of an Atacama cyanobacterium
Scientific Reports (2023)
-
The Ycf48 accessory factor occupies the site of the oxygen-evolving manganese cluster during photosystem II biogenesis
Nature Communications (2023)
-
Changes in intracellular energetic and metabolite states due to increased galactolipid levels in Synechococcus elongatus PCC 7942
Scientific Reports (2023)
-
Absolute quantification of cellular levels of photosynthesis-related proteins in Synechocystis sp. PCC 6803
Photosynthesis Research (2023)
-
Expanding the arsenal of bacterial spearguns
Nature Microbiology (2022)