Abstract
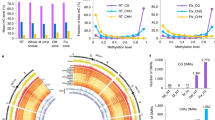
Advances in deciphering the functional architecture of eukaryotic genomes have been facilitated by recent breakthroughs in sequencing technologies, enabling a more comprehensive representation of genes and repeat elements in genome sequence assemblies, as well as more sensitive and tissue-specific analyses of gene expression. Here we show that PacBio sequencing has led to a substantially improved genome assembly of Medicago truncatula A17, a legume model species notable for endosymbiosis studies1, and has enabled the identification of genome rearrangements between genotypes at a near-base-pair resolution. Annotation of the new M. truncatula genome sequence has allowed for a thorough analysis of transposable elements and their dynamics, as well as the identification of new players involved in symbiotic nodule development, in particular 1,037 upregulated long non-coding RNAs (lncRNAs). We have also discovered that a substantial proportion (~35% and 38%, respectively) of the genes upregulated in nodules or expressed in the nodule differentiation zone colocalize in genomic clusters (270 and 211, respectively), here termed symbiotic islands. These islands contain numerous expressed lncRNA genes and display differentially both DNA methylation and histone marks. Epigenetic regulations and lncRNAs are therefore attractive candidate elements for the orchestration of symbiotic gene expression in the M. truncatula genome.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
This Whole Genome Shotgun project has been deposited at DDBJ/ENA/GenBank under the accession PSQE00000000. The version described in this paper is version PSQE01000000. Raw reads from PacBio, ChIP-seq and small RNAseq experiments have been deposited at the Sequence Read Archive (SRA) (project accession number: SRP131849). Data related to gene annotation, transposable element annotation and ChIP-seq analyses, as well as Supplementary Table 6, are available at the web portal: https://medicago.toulouse.inra.fr/MtrunA17r5.0-ANR/; downloads section.
References
Martin, F.M., Uroz, S. & Barker, D.G. Ancestral alliances: plant mutualistic symbioses with fungi and bacteria. Science 356, eaad4501 (2017).
Young, N. D. & Udvardi, M. Translating Medicago truncatula genomics to crop legumes. Curr. Opin. Plant Biol. 12, 193–201 (2009).
Young, N. D. et al. The Medicago genome provides insight into the evolution of rhizobial symbioses. Nature 480, 520–524 (2011).
Tang, H. et al. An improved genome release (version Mt4.0) for the model legume Medicago truncatula. BMC Genomics 15, 312 (2014).
Moll, K. M. et al. Strategies for optimizing BioNano and Dovetail explored through a second reference quality assembly for the legume model, Medicago truncatula. BMC Genomics 18, 578 (2017).
Kamphuis, L. G. et al. The Medicago truncatula reference accession A17 has an aberrant chromosomal configuration. New Phytol. 174, 299–303 (2007).
de Bang, T. et al. Genome-wide identification of Medicago peptides involved in macronutrient responses and nodulation. Plant Physiol. 175, 1669–1689 (2017).
Miller, J. R. et al. Hybrid assembly with long and short reads improves discovery of gene family expansions. BMC Genomics 18, 541 (2017).
Roux, B. et al. An integrated analysis of plant and bacterial gene expression in symbiotic root nodules using laser-capture microdissection coupled to RNA sequencing. Plant J. 77, 817–837 (2014).
Jardinaud, M. F. et al. A laser dissection-RNAseq analysis highlights the activation of cytokinin pathways by nod factors in the Medicago truncatula root epidermis. Plant Physiol. 171, 2256–2276 (2016).
Stanton-Geddes, J. et al. Candidate genes and genetic architecture of symbiotic and agronomic traits revealed by whole-genome, sequence-based association genetics in Medicago truncatula. PLoS ONE 8, e65688 (2013).
Ariel, F. et al. Noncoding transcription by alternative RNA polymerases dynamically regulates an auxin-driven chromatin loop. Mol. Cell 55, 383–396 (2014).
Krzyczmonik, K., Wroblewska-Swiniarska, A. & Swiezewski, S. Developmental transitions in Arabidopsis are regulated by antisense RNAs resulting from bidirectionally transcribed genes. RNA Biol. 14, 838–842 (2017).
Swiezewski, S., Liu, F., Magusin, A. & Dean, C. Cold-induced silencing by long antisense transcripts of an Arabidopsis polycomb target. Nature 462, 799–802 (2009).
Fedak, H. et al. Control of seed dormancy in Arabidopsis by a cis-acting noncoding antisense transcript. Proc. Natl Acad. Sci. USA 113, E7846–E7855 (2016).
Henriques, R. et al. The antiphasic regulatory module comprising CDF5 and its antisense RNA FLORE links the circadian clock to photoperiodic flowering. New Phytol. 216, 854–867 (2017).
Vernié, T. et al. EFD is an ERF transcription factor involved in the control of nodule number and differentiation in Medicago truncatula. Plant Cell 20, 2696–2713 (2008).
Satgé, C. et al. Reprogramming of DNA methylation is critical for nodule development in Medicago truncatula. Nat. Plants 2, 16166 (2016).
Kalo, P. et al. Nodulation signaling in legumes requires NSP2, a member of the GRAS family of transcriptional regulators. Science 308, 1786–1789 (2005).
Sinharoy, S. et al. The C2H2 transcription factor regulator of symbiosome differentiation represses transcription of the secretory pathway gene VAMP721a and promotes symbiosome development in Medicago truncatula. Plant Cell 25, 3584–3601 (2013).
Marsh, J. F. et al. Medicago truncatula NIN is essential for rhizobial-independent nodule organogenesis induced by autoactive calcium/calmodulin-dependent protein kinase. Plant Physiol. 144, 324–335 (2007).
Ovchinnikova, E. et al. IPD3 controls the formation of nitrogen-fixing symbiosomes in pea and Medicago Spp. Mol. Plant Microbe Interact. 24, 1333–1344 (2011).
Lefebvre, B. et al. A remorin protein interacts with symbiotic receptors and regulates bacterial infection. Proc. Natl Acad. Sci. USA 107, 2343–2348 (2010).
Berrabah, F. et al. A nonRD receptor-like kinase prevents nodule early senescence and defense-like reactions during symbiosis. New Phytol. 203, 1305–1314 (2014).
Alunni, B. et al. Genomic organization and evolutionary insights on GRP and NCR genes, two large nodule-specific gene families in Medicago truncatula. Mol. Plant Microbe Interact. 20, 1138–1148 (2007).
Graham, M. A., Silverstein, K. A., Cannon, S. B. & VandenBosch, K. A. Computational identification and characterization of novel genes from legumes. Plant Physiol. 135, 1179–1197 (2004).
Pan, H. & Wang, D. Nodule cysteine-rich peptides maintain a working balance during nitrogen-fixing symbiosis. Nat. Plants 3, 17048 (2017).
Liu, J. et al. Recruitment of novel calcium-binding proteins for root nodule symbiosis in Medicago truncatula. Plant Physiol. 141, 167–177 (2006).
Alunni, B. & Gourion, B. Terminal bacteroid differentiation in the legume-rhizobium symbiosis: nodule-specific cysteine-rich peptides and beyond. New Phytol. 211, 411–417 (2016).
Matzke, M. A. & Mosher, R. A. RNA-directed DNA methylation: an epigenetic pathway of increasing complexity. Nat. Rev. Genet. 15, 394–408 (2014).
Hurst, L. D., Pal, C. & Lercher, M. J. The evolutionary dynamics of eukaryotic gene order. Nat. Rev. Genet. 5, 299–310 (2004).
Nutzmann, H. W., Huang, A. & Osbourn, A. Plant metabolic clusters – from genetics to genomics. New Phytol. 211, 771–789 (2016).
Reimegard, J. et al. Genome-wide identification of physically clustered genes suggests chromatin-level co-regulation in male reproductive development in Arabidopsis thaliana. Nucleic Acids Res. 45, 3253–3265 (2017).
Plaza, S., Menschaert, G. & Payre, F. In search of lost small peptides. Annu. Rev. Cell Dev. Biol. 33, 391–416 (2017).
Hnisz, D. & Young, R. A. New insights into genome structure: genes of a feather stick together. Mol. Cell 67, 730–731 (2017).
Rowley, M. J. et al. Evolutionarily conserved principles predict 3D chromatin organization. Mol. Cell 67, 837–852 e7 (2017).
Mele, M. & Rinn, J. L. ‘Cat’s cradling’ the 3D genome by the act of LncRNA transcription. Mol. Cell 62, 657–664 (2016).
Mercer, T. R. & Mattick, J. S. Structure and function of long noncoding RNAs in epigenetic regulation. Nat. Struct. Mol. Biol. 20, 300–307 (2013).
Heo, J. B. & Sung, S. Vernalization-mediated epigenetic silencing by a long intronic noncoding RNA. Science 331, 76–79 (2011).
Mayjonade, B. et al. Extraction of high-molecular-weight genomic DNA for long-read sequencing of single molecules. Biotechniques 61, 203–205 (2016).
Berlin, K. et al. Corrigendum: assembling large genomes with single-molecule sequencing and locality-sensitive hashing. Nat. Biotechnol. 33, 1109 (2015).
Berlin, K. et al. Assembling large genomes with single-molecule sequencing and locality-sensitive hashing. Nat. Biotechnol. 33, 623–630 (2015).
Badouin, H. et al. The sunflower genome provides insights into oil metabolism, flowering and Asterid evolution. Nature 546, 148–152 (2017).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Chin, C. S. et al. Phased diploid genome assembly with single-molecule real-time sequencing. Nat. Methods 13, 1050–1054 (2016).
Raymond, O. et al. The Rosa genome provides new insights into the domestication of modern roses. Nat. Genet. 50, 772–777 (2018).
Tayeh, N. et al. A tandem array of CBF/DREB1 genes is located in a major freezing tolerance QTL region on Medicago truncatula chromosome 6. BMC Genomics 14, 814 (2013).
Kulikova, O. et al. Satellite repeats in the functional centromere and pericentromeric heterochromatin of Medicago truncatula. Chromosoma 113, 276–283 (2004).
Chin, C. S. et al. Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat. Methods 10, 563–569 (2013).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963 (2014).
Foissac, S. et al. Genome annotation in plants and fungi: EuGene as a model platform. Current Bioinformatics 3, 87–97 (2008).
Bao, W., Kojima, K. K. & Kohany, O. Repbase Update, a database of repetitive elements in eukaryotic genomes. Mob. DNA 6, 11 (2015).
Zerbino, D. R. Using the Velvet de novo assembler for short-read sequencing technologies. Curr. Protoc. Bioinformatics 11, Unit11 5 (2010).
Wu, T. D. & Watanabe, C. K. GMAP: a genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics 21, 1859–1875 (2005).
Tephra: A Tool for Discovering Transposable Elements and Describing Patterns of Genome Evolution v.0.12.2 (Staton, S., 2017); https://github.com/sestaton/tephra
Generic Feature Format Version 3 (GFF3) v.1.23 (Stein, L., 2013); https://github.com/The-Sequence-Ontology/Specifications/blob/master/gff3.md
Staton, S. E. & Burke, J. M. Transposome: a toolkit for annotation of transposable element families from unassembled sequence reads. Bioinformatics 31, 1827–1829 (2015).
Kurtz, S., Narechania, A., Stein, J. C. & Ware, D. A new method to compute K-mer frequencies and its application to annotate large repetitive plant genomes. BMC Genomics 9, 517 (2008).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Guizard, S., Piegu, B. & Bigot, Y. DensityMap: a genome viewer for illustrating the densities of features. BMC Bioinformatics 17, 204 (2016).
Veluchamy, A. et al. LHP1 regulates H3K27me3 spreading and shapes the three-dimensional conformation of the Arabidopsis genome. PLoS ONE 11, e0158936 (2016).
Acknowledgements
We thank C. Ben and L. Gentzbittel (EcoLab, Université de Toulouse, CNRS, Toulouse INP, UPS, France), G. Aubert, R. Thompson and K. Gallardo (INRA, UMR 1347, Agroécologie, Dijon, France) and B. Gronenborn (I2BC, CNRS, Paris Sud, CEA, University of Paris Saclay, Gif sur Yvette, France) for providing small RNA data on disease responses, seeds and viroid-infected plants, respectively, as well as N. Peeters (LIPM, Toulouse) for mRNA data used for genome annotation. We thank M.C. Le Paslier for her help in Illumina sequencing. This work was supported by the ANR grants EPISYM (grant no. ANR-15-CE20-0002), NODCCAAT (no. ANR-15-CE20-0012), REGULEG (no. ANR-15-CE20-0001), the ‘Laboratoire d’Excellence (LABEX)’ TULIP (no. ANR-10-LABX-41), the LABEX Saclay Plant Sciences (SPS; no. ANR-10-LABX-40) and the European Research Council (no. ERC-SEXYPARTH), and we made use of data previously generated in the ANR SYMbiMICS (ANR-08-GENO-106) and the INRA SPE EPINOD projects. The sequencing platform was supported by France Génomique National infrastructure (grant no. ANR-10-INBS-09) and by the GET-PACBIO programme (Programme opérationnel FEDER-FSE MIDI-PYRENEES ET GARONNE 2014-2020). We are grateful to the Genotoul bioinformatics platform Toulouse Midi-Pyrenees (Bioinfo Genotoul) for providing computing and storage resources. C. Satgé was supported by a doctoral grant from the French Ministry of Education and Research.
Author information
Authors and Affiliations
Contributions
S.Mo., B.M., C.L-R. and O.B. prepared DNA samples and performed PacBio sequencing. S.Cau., C.C-D., W.M. and H.B. built the Bionano optical maps. B.M., J.G., W.M., S.Mu. and A.Ber. designed and performed Illumina seq of BAC end sequencing (EcoR1 library). F.D. prepared DNA samples and managed Illumina sequencing. J.G. assembled the genome. E.S., S.Car. and J.G. annotated protein-coding genes and miRNAs. S.E.S. annotated and analysed repeats and transposable elements. S.Car. developed the Medicago bioinformatics portal. J.K. positioned HapMap data on the new reference genome. C.S. and C.L.-B. prepared samples for the sRNA analyses. C.L.-B. conducted the miRNA analyses. T.B., C.L.-B. and Y.P. conducted the siRNA analyses. S.Mo. and M.P. prepared the histone mark samples. D.L. and M.P. performed the ChIP experiments. M.B., D.L. and A.Ben. performed ChIP-seq. M.Za., M.Zo., M.B., S.Car., Y.P. and P.G. performed the analysis of the ChIP-seq data. Y.P. and P.G. conducted the lncRNA analyses. Y.P., S.Car. and P.G. performed the gene family analyses. J.G. and T.B. performed the sRNA and mRNA expression analyses. M.-F.J. performed the gene and siRNA differential expression analyses. Y.P. and P.G. performed the integrated analyses of the symbiotic islands. P.G., J.G., M.C., A.N. and J.B. contributed to the project set-up. P.G., J.G., S.E.S. and C.L.-B. wrote the manuscript, with contributions from M.C., F.F., J.B., B.M., Y.P., F.D., A.N., M.Zo., E.S. and S.Mu. P.G., J.G. and M.C. coordinated the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–6, Supplementary Tables 1 and 2, Supplementary Notes on genome sequencing and assembly; genome annotation; transposable elements and repeats; transcriptome analysis; analysis of symbiosis-related islands, and Supplementary References. Supplementary Table 6 (M. truncatula gene annotation, RNAseq data, MtV4 ID and affymetrix probe correspondence) can be found at https://medicago.toulouse.inra.fr/MtrunA17r5.0-ANR/; downloads section.
Supplementary Table 3
Transduplicate analyses
Supplementary Table 4
miRNA analyses
Supplementary Table 5
siRNA analyses
Supplemental Table 7
Expression correlation analyses
Supplementary Table 8
Genes expressed in symbiosis-related islands
Supplementary Table 9
Conservation of symbiosis-related island genes in M. truncatula R108 genome
Rights and permissions
About this article
Cite this article
Pecrix, Y., Staton, S.E., Sallet, E. et al. Whole-genome landscape of Medicago truncatula symbiotic genes. Nature Plants 4, 1017–1025 (2018). https://doi.org/10.1038/s41477-018-0286-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-018-0286-7
This article is cited by
-
Genome assembly of Medicago truncatula accession SA27063 provides insight into spring black stem and leaf spot disease resistance
BMC Genomics (2024)
-
Spatial co-transcriptomics reveals discrete stages of the arbuscular mycorrhizal symbiosis
Nature Plants (2024)
-
A cluster of putative resistance genes is associated with a dominant resistance to sunflower broomrape
Theoretical and Applied Genetics (2024)
-
A rare non-canonical splice site in Trema orientalis SYMRK does not affect its dual symbiotic functioning in endomycorrhiza and rhizobium nodulation
BMC Plant Biology (2023)
-
Systematic approaches to C-lignin engineering in Medicago truncatula
Biotechnology for Biofuels and Bioproducts (2023)