Abstract
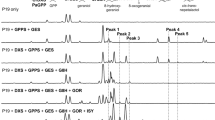
Plant genomes encode isopentenyl phosphate kinases (IPKs) that reactivate isopentenyl phosphate (IP) via ATP-dependent phosphorylation, forming the primary metabolite isopentenyl diphosphate (IPP) used generally for isoprenoid/terpenoid biosynthesis. Therefore, the existence of IPKs in plants raises unanswered questions concerning the origin and regulatory roles of IP in plant terpenoid metabolism. Here, we provide genetic and biochemical evidence showing that IP forms during specific dephosphorylation of IPP catalysed by a subset of Nudix superfamily hydrolases. Increasing metabolically available IP by overexpression of a bacterial phosphomevalonate decarboxylase (MPD) in Nicotiana tabacum resulted in significant enhancement in both monoterpene and sesquiterpene production. These results indicate that perturbing IP metabolism results in measurable changes in terpene products derived from both the methylerythritol phosphate (MEP) and mevalonate (MVA) pathways. Moreover, the unpredicted peroxisomal localization of bacterial MPD led us to discover that the step catalysed by phosphomevalonate kinase (PMK) imposes a hidden constraint on flux through the classical MVA pathway. These complementary findings fundamentally alter conventional views of metabolic regulation of terpenoid metabolism in plants and provide new metabolic engineering targets for the production of high-value terpenes in plants.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Hemmerlin, A., Harwood, J. L. & Bach, T. J. A raison d’être for two distinct pathways in the early steps of plant isoprenoid biosynthesis? Prog. Lipid Res. 51, 95–148 (2012).
Tholl, D. Biosynthesis and biological functions of terpenoids in plants. Adv. Biochem. Eng. Biotechnol. 148, 63–106 (2015).
Gershenzon, J. & Dudareva, N. The function of terpene natural products in the natural world. Nat. Chem. Biol. 3, 408–414 (2007).
Tippmann, S., Chen, Y., Siewers, V. & Nielsen, J. From flavors and pharmaceuticals to advanced biofuels: Production of isoprenoids in Saccharomyces cerevisiae. Biotechnol. J. 8, 1435–1444 (2013).
Zerbe, P. & Bohlmann, J. Plant diterpene synthases: Exploring modularity and metabolic diversity for bioengineering. Trends Biotechnol. 33, 419–428 (2015).
Vranová, E., Coman, D. & Gruissem, W. Network analysis of the MVA and MEP pathways for isoprenoid synthesis. Annu. Rev. Plant Biol. 64, 665–700 (2013).
Hemmerlin, A. Post-translational events and modifications regulating plant enzymes involved in isoprenoid precursor biosynthesis. Plant Sci. 203–204, 41–54 (2013).
Vickers, C. E., Bongers, M., Liu, Q., Delatte, T. & Bouwmeester, H. Metabolic engineering of volatile isoprenoids in plants and microbes. Plant Cell Environ. 37, 1753–1775 (2014).
Wu, S. et al. Redirection of cytosolic or plastidic isoprenoid precursors elevates terpene production in plants. Nat. Biotechnol. 24, 1441–1447 (2006).
Farhi, M. et al. Generation of the potent anti-malarial drug artemisinin in tobacco. Nat. Biotechnol. 29, 1072–1074 (2011).
Muñoz-Bertomeu, J., Sales, E., Ros, R., Arrillaga, I. & Segura, J. Up-regulation of an N-terminal truncated 3-hydroxy-3-methylglutaryl CoA reductase enhances production of essential oils and sterols in transgenic Lavandula latifolia. Plant Biotechnol. J. 5, 746–758 (2007).
Liao, P., Hemmerlin, A., Bach, T. J. & Chye, M.-L. The potential of the mevalonate pathway for enhanced isoprenoid production. Biotechnol. Adv. 34, 697–713 (2016).
Dellas, N., Thomas, S. T., Manning, G. & Noel, J. P. Discovery of a metabolic alternative to the classical mevalonate pathway. eLife 2, e00672 (2013).
Henry, L. K., Gutensohn, M., Thomas, S. T., Noel, J. P. & Dudareva, N. Orthologs of the archaeal isopentenyl phosphate kinase regulate terpenoid production in plants. Proc. Natl Acad. Sci. USA 112, 10050–10055 (2015).
VanNice, J. C. et al. Identification in Haloferax volcanii of phosphomevalonate decarboxylase and isopentenyl phosphate kinase as catalysts of the terminal enzyme reactions in an archaeal alternate mevalonate pathway. J. Bacteriol. 196, 1055–1063 (2014).
Karačić, Z. et al. A novel plant enzyme with dual activity: an atypical Nudix hydrolase and a dipeptidyl peptidase III. Biol. Chem. 398, 101–112 (2017).
Kraszewska, E. The plant Nudix hydrolase family. Acta Biochim. Pol. 55, 663–671 (2008).
McLennan, A. G. Substrate ambiguity among the nudix hydrolases: biologically significant, evolutionary remnant, or both? Cell. Mol. Life Sci. 70, 373–385 (2013).
Ogawa, T. et al. Molecular characterization of organelle-type Nudix hydrolases in Arabidopsis. Plant Physiol. 148, 1412–1424 (2008).
Ogawa, T., Ueda, Y., Yoshimura, K. & Shigeoka, S. Comprehensive analysis of cytosolic nudix hydrolases in Arabidopsis thaliana. J. Biol. Chem. 280, 25277–25283 (2005).
Yoshimura, K., Ogawa, T., Ueda, Y. & Shigeoka, S. AtNUDX1, an 8-oxo-7,8-dihydro-2′-deoxyguanosine 5′-triphosphate pyrophosphohydrolase, is responsible for eliminating oxidized nucleotides in Arabidopsis. Plant Cell Physiol. 48, 1438–1449 (2007).
Liu, J. et al. Structural insights into the substrate recognition mechanism of Arabidopsis GPP-bound NUDX1 for noncanonical monoterpene biosynthesis. Molecular Plant 11, 218–221 (2018).
Svensson, L. M. et al. Crystal structure of human MTH1 and the 8-oxo-dGMP product complex. FEBS Lett. 585, 2617–2621 (2011).
Nakamura, T. et al. Structural and dynamic features of the MutT protein in the recognition of nucleotides with the mutagenic 8-oxoguanine base. J. Biol. Chem. 285, 444–452 (2010).
Magnard, J.-L. et al. Biosynthesis of monoterpene scent compounds in roses. Science 349, 81–83 (2015).
Jones, C. G. et al. Sandalwood fragrance biosynthesis involves sesquiterpene synthases of both the terpene synthase (TPS)-a and TPS-b subfamilies, including santalene synthases. J. Biol. Chem. 286, 17445–17454 (2011).
Simkin, A. J. et al. Peroxisomal localisation of the final steps of the mevalonic acid pathway in planta. Planta 234, 903–914 (2011).
Reumann, S., Chowdhary, G. & Lingner, T. Characterization, prediction and evolution of plant peroxisomal targeting signals type 1 (PTS1s). Biochim. Biophys. Acta Mol. Cell Res. 1863, 790–803 (2016).
Curtis, I. S., Davey, M. R. & Power, J. B. Leaf disk transformation. Methods Mol. Biol. 44, 59–70 (1995).
Sparkes, I. A., Runions, J., Kearns, A. & Hawes, C. Rapid, transient expression of fluorescent fusion proteins in tobacco plants and generation of stably transformed plants. Nat. Protoc. 1, 2019–2025 (2006).
Nelson, B. K., Cai, X. & Nebenführ, A. A multicolored set of in vivo organelle markers for co-localization studies in Arabidopsis and other plants. Plant J. 51, 1126–1136 (2007).
Yoo, H. et al. An alternative pathway contributes to phenylalanine biosynthesis in plants via a cytosolic tyrosine:phenylpyruvate aminotransferase. Nat. Commun. 4, 2833 (2013).
Powell, H. R., Battye, T. G. G., Kontogiannis, L., Johnson, O. & Leslie, A. G. W. Integrating macromolecular X-ray diffraction data with the graphical user interface iMosflm. Nat. Protoc. 12, 1310–1325 (2017).
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution? Acta Crystallogr. D 69, 1204–1214 (2013).
Vagin, A. & Teplyakov, A. MOLREP: an automated program for molecular replacement. J. Appl. Crystallogr. 30, 1022–1025 (1997).
Stein, N. et al. CHAINSAW: a program for mutating pdb files used as templates in molecular replacement. J. Appl. Crystallogr. 41, 641–643 (2008).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Terwilliger, T. C. et al. Iterative model building, structure refinement and density modification with the PHENIX AutoBuildwizard. Acta Crystallogr. D 64, 61–69 (2008).
Curtis, M. D. & Grossniklaus, U. A Gateway cloning vector set for high-throughput functional analysis of genes in plants. Breakthr. Technol. 133, 462–469 (2003).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Acknowledgements
This work was supported by the USDA National Institute of Food and Agriculture Predoctoral Grant 2017-67011-26076 to L.K.H. and start-up funds from Purdue University to J.R.W. and S.A.K. This work was also supported by the USDA National Institute of Food and Agriculture Hatch Project number 177845.
Author information
Authors and Affiliations
Contributions
J.P.N. and N.D. conceived the project; L.K.H., S.T.T., J.R.W., S.A.K., J.B., J.P.N. and N.D. designed the experiments; L.K.H., S.T.T., J.R.W., J.H.L. and T.C.D. performed the experiments; L.K.H., S.T.T., J.R.W., J.H.L., T.C.D., S.A.K., J.B., J.P.N. and N.D. analysed the data; L.K.H., S.T.T., J.R.W., J.P.N. and N.D. wrote the paper. All authors read and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–10 and Supplementary Tables 1–2.
Supplementary Data 1
Primers used in this work.
Rights and permissions
About this article
Cite this article
Henry, L.K., Thomas, S.T., Widhalm, J.R. et al. Contribution of isopentenyl phosphate to plant terpenoid metabolism. Nature Plants 4, 721–729 (2018). https://doi.org/10.1038/s41477-018-0220-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-018-0220-z
This article is cited by
-
Effects of salt stress on soil enzyme activities and rhizosphere microbial structure in salt-tolerant and -sensitive soybean
Scientific Reports (2023)
-
Biosynthesis, natural distribution, and biological activities of acyclic monoterpenes and their derivatives
Phytochemistry Reviews (2023)
-
Dynamic changes in terpenoids metabolisms of mountain-cultivated ginseng harvested at different months and ages
Plant Growth Regulation (2023)
-
Suppression of GhGLU19 encoding β-1,3-glucanase promotes seed germination in cotton
BMC Plant Biology (2022)
-
Farnesyl pyrophosphate compartmentalization in the green microalga Chlamydomonas reinhardtii during heterologous (E)-α-bisabolene production
Microbial Cell Factories (2022)