Abstract
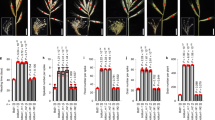
Transcriptional silencer and copy number variants (CNVs) are associated with gene expression. However, their roles in generating phenotypes have not been well studied. Here we identified a rice quantitative trait locus, SGDP7 (Small Grain and Dense Panicle 7). SGDP7 is identical to FZP (FRIZZY PANICLE), which represses the formation of axillary meristems. The causal mutation of SGDP7 is an 18-bp fragment, named CNV-18bp, which was inserted ~5.3 kb upstream of FZP and resulted in a tandem duplication in the cultivar Chuan 7. The CNV-18bp duplication repressed FZP expression, prolonged the panicle branching period and increased grain yield by more than 15% through substantially increasing the number of spikelets per panicle (SPP) and slightly decreasing the 1,000-grain weight (TGW). The transcription repressor OsBZR1 binds the CGTG motifs in CNV-18bp and thereby represses FZP expression, indicating that CNV-18bp is the upstream silencer of FZP. These findings showed that the silencer CNVs coordinate a trade-off between SPP and TGW by fine-tuning FZP expression, and balancing the trade-off could enhance yield potential.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ikeda, M. et al. Genes offering the potential for designing yield-related traits in rice. Curr. Opin. Plant Biol. 16, 213–220 (2013).
Wang, Y. H. & Li, J. Y. Branching in rice. Curr. Opin. Plant Biol. 14, 1–6 (2011).
Zhang, D. B. & Zheng, Y. Molecular control of grass inflorescence development. Annu. Rev. Plant Biol. 65, 553–578 (2014).
Bai, X. F. et al. Genome-wide association analysis reveals different genetic control in panicle architecture between indica and japonica rice. Plant Genome 9, 2 (2016).
Xing, Y. Z. & Zhang, Q. F. Genetic and molecular bases of rice yield. Annu. Rev. Plant Biol. 61, 421–442 (2010).
Zuo, J. R. & Li, J. Y. Molecular genetic dissection of quantitative trait loci regulating rice grain size. Annu. Rev. Genet. 48, 99–118 (2014).
Huang, X. Z. et al. Natural variation at the DEP1 locus enhances grain yield in rice. Nat. Genet. 41, 494–497 (2009).
Jiao, Y. Q. et al. Regulation of OsSPL14 by OsmiR156 defines ideal plant architecture in rice. Nat. Genet. 42, 541–544 (2010).
Li, M. et al. Mutations in the F-box gene LARGER PANICLE improve the panicle architecture and enhance the grain yield in rice. Plant Biotechnol. J. 9, 1002–1013 (2011).
Zha, X. J. et al. Over-expression of the rice LRK1 gene improves quantitative yield components. Plant Biotechnol. J. 7, 611–620 (2009).
Yoshida, A. et al. TAWAWA1, a regulator of rice inflorescence architecture, functions through the suppression of meristem phase transition. Proc. Natl Acad. Sci. USA 110, 767–772 (2013).
Fan, C. C. et al. GS3, a major QTL for grain length and weight and minor QTL for grain width and thickness in rice, encodes a putative transmembrane protein. Theor. Appl. Genet. 112, 1164–1171 (2006).
Qi, P. et al. The novel quantitative trait locus GL3.1 controls rice grain size and yield by regulating Cyclin-T1;3. Cell Res. 22, 1666–1680 (2012).
Zhang, X. J. et al. Rare allele of OsPPKL1 associated with grain length causes extra-large grain and a significant yield increase in rice. Proc. Natl Acad. Sci. USA 109, 21534–21539 (2012).
Wang, Y. X. et al. Copy number variation at the GL7 locus contributes to grain size diversity in rice. Nat. Genet. 47, 944–948 (2015).
Wang, S. K. et al. The OsSPL16-GW7 regulatory module determines grain shape and simultaneously improves rice yield and grain quality. Nat. Genet. 47, 949–954 (2015b).
Si, L. Z. et al. OsSPL13 controls grain size in cultivated rice. Nat. Genet. 48, 447–456 (2016).
Song, X. J. et al. A QTL for rice grain width and weight encodes a previously unknown RING-type E3 ubiquitin ligase. Nat. Genet. 39, 623–630 (2007).
Shomura, A. et al. Deletion in a gene associated with grain size increased yields during rice domestication. Nat. Genet. 40, 1023–1028 (2008).
Weng, J. F. et al. Isolation and initial characterization of GW5, a major QTL associated with rice grain width and weight. Cell Res. 18, 1199–1209 (2008).
Wang, S. K. et al. Control of grain size, shape and quality by OsSPL16 in rice. Nat. Genet. 44, 950–954 (2012).
Wang, E. T. et al. Control of rice grain-filling and yield by a gene with a potential signature of domestication. Nat. Genet. 40, 1370–1374 (2008).
Li, Y. B. et al. Natural variation in GS5 plays an important role in regulating grain size and yield in rice. Nat. Genet. 43, 1266–1269 (2011).
Miura, K. et al. OsSPL14 promotes panicle branching and higher grain productivity in rice. Nat. Genet. 42, 545–549 (2010).
Che, R. H. et al. Control of grain size and rice yield by GL2-mediated brassinosteroid responses. Nat. Plants 2, 15195 (2015).
Duan, P. G. et al. Regulation of OsGRF4 by OsmiR396 controls grain size and yield in rice. Nat. Plants 2, 15203 (2015).
Gao, F. et al. Blocking miR396 increases rice yield by shaping inflorescence architecture. Nat. Plants 2, 15196 (2015).
Sakabe, N. J. et al. Transcriptional enhancers in development and disease. Genome Biol. 13, 238 (2012).
Kolovos, P. et al. Enhancers and silencers: an integrated and simple model for their function. Epigenet. Chrom. 5, 1 (2012).
Ishii, T. et al. OsLG1 regulates a closed panicle trait in domesticated rice.Nat. Genet. 45, 462–465 (2013).
Zhu, Z. F. et al. Genetic control of inflorescence architecture during rice domestication. Nat. Commun. 4, 2200 (2013).
Mathelier, A. et al. JASPAR 2014: an extensively expanded and updated open-access database of transcription factor binding profiles. Nucleic Acids Res. 42, D142–D147 (2014).
Clark, R. M. et al. A distant upstream enhancer at the maize domestication gene t b1 has pleiotropic effects on plant and inflorescent architecture.Nat. Genet. 38, 594–597 (2006).
Liu, L. et al. Induced and natural variation of promoter length modulates the photoperiodic response of FLOWERING LOCUS T. Nat. Commun. 5, 4558 (2014).
Liu, J. F. et al. GW5 acts in the brassinosteroid signaling pathway to regulate grain width and weight in rice. Nat. Plants 3, 17043 (2017).
McEachern, L. & Lloyd, V. The maize b1 paramutation control region causes epigenetic silencing in Drosophila melanogaster. Mol. Genet. Genomics 287, 591–606 (2012).
Bai, X. F. et al. Genetic dissection of rice grain shape using a recombinant inbred line population derived from two contrasting parents and fine mapping a pleiotropic quantitative trait locus qGL7. BMC Genet. 11, 16 (2010).
Bañuelos, M. A. et al. Inventory and functional characterization of the HAK potassium transporters of rice. Plant Physiol. 130, 784–795 (2002).
Komatsu, M. et al. FRIZZY PANICLE is required to prevent the formation of axillary meristems and to establish floral meristem identity in rice spikelets. Development 130, 3841–3850 (2003).
Bai, X. F. et al. Regulatory role of FZP in the determination of panicle branching and spikelet formation in rice. Sci. Rep. 6, 19022 (2016).
He, J. X. et al. BZR1 is a transcriptional repressor with dual roles in brassinosteroid homeostasis and growth responses. Science 307,1634–1638 (2005).
Lou, S. L. et al. The far-upstream regulatory region of RFL is required for its precise spatial-temporal expression for floral development in rice.Plant Mol. Biol. 93, 185–195 (2017).
Oh, E. et al. TOPLESS mediates brassinosteroid-induced transcriptional repression through interaction with BZR1. Nat. Commun. 5, 4140 (2014).
Civáň, P. et al. Three geographically separate domestications of Asian rice. Nat. Plants 1, 15164 (2015).
Li, Y. B. et al. Chalk5 encodes a vacuolar H+-translocating pyrophosphatase influencing grain chalkiness in rice. Nat. Genet. 46, 398–404 (2014).
Tan, Y. F. et al. The three important traits for cooking and eating quality of rice grains are controlled by a single locus in an elite rice hybrid, Shanyou 63. Theor. Appl. Genet. 99, 642–648 (1999).
Qiao, S. L. et al. The RLA1/SMOS1 transcription factor functions with OsBZR1 to regulate brassinosteroid signaling and rice architecture. Plant Cell 29, 292–309 (2017).
McGinnis, K. et al. Transgene-induced RNA interference as a tool for plant functional genomics. Methods Enzymol. 392, 1–24 (2005).
Yuan, B. et al. Mitogen-activated protein kinase OsMPK6 negatively regulates rice disease resistance to bacterial pathogens. Planta 226, 953–960 (2007).
De, B. M. & Debrouwer, D. RNA–RNA in situ hybridization using digoxigenin-labelled probes: the use of high-molecular-weight polyvinyl alcohol in the alkaline phosphatase indoxyl–nitroblue tetrazolium reaction. Anal. Biochem. 215, 86–89 (1993).
Librado, P. & Rozas, J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452 (2009).
Lin, R. H. et al. Transposase-derived transcription factors regulate light signaling in Arabidopsis. Science 318, 1302–1305 (2007).
Smaczniak, C. et al. Characterization of MADS-domain transcription factor complexes in Arabidopsis flower development. Proc. Natl Acad. Sci. USA. 109, 1560–1565 (2012).
Zhu, X. L. et al. Brassinosteroids promote development of rice pollen grains and seeds by triggering expression of Carbon Starved Anther, a MYB domain protein. Plant J. 82, 570–581 (2015).
Ohta, M. et al. Repression domains of class II ERF transcriptional repressors share an essential motif for active repression. Plant Cell 13, 1959–1968 (2001).
Hao, Y. J. et al. Plant NAC-type transcription factor proteins contain a NARD domain for repression of transcriptional activation. Planta 232, 1033–1043 (2010).
Zong, W. et al. Feedback regulation of ABA signaling and biosynthesis by a bZIP transcription 4 factor targets drought resistance related genes. Plant Physiol. 171, 2810–2825 (2016).
Acknowledgements
We thank K. Kaufmann at Potsdam University for providing the pSPUTK expression vector. We thank D. Zhang at Shanghai Jiao Tong University for kindly providing the antiserum for OsBZR1, and S. Sun at Huazhong Agricultural University for kindly providing the OsBZR1-activation plants. This work was financially supported through grants from the National Natural Science Foundation of China (91535301), the National Key Research and Development Program of China (2016YFD0100403), the National Special Program for Research of Transgenic Plants of China (2014ZX0800944B) and Hubei Collaborative Innovation Center for Grain Industry (2015ZD003).
Author information
Authors and Affiliations
Contributions
X.B. performed fine mapping of the quantitative trait locus, in situ hybridization, qRT–PCR, RNA-seq analysis, ChIP analysis, phenotypic observation and data analysis. Y.Hua. conducted genetic transformation and SEM. Y.Hu performed yeast one-hybrid assays and transcriptional activity assays. H.L. performed electrophoretic mobility shift assays. B.Z. collected some phenotype data. C.S. constructed the vectors for expression of the OsBZR1 protein. G.H. and Z.H. performed the nucleotide diversity analysis. Y.X. designed the experiments and wrote the paper together with X.B.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Information
Supplementary Figures 1–15, Supplementary Tables 1–9
Rights and permissions
About this article
Cite this article
Bai, X., Huang, Y., Hu, Y. et al. Duplication of an upstream silencer of FZP increases grain yield in rice. Nature Plants 3, 885–893 (2017). https://doi.org/10.1038/s41477-017-0042-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-017-0042-4
This article is cited by
-
Fine mapping of the panicle length QTL qPL5 in rice
Molecular Breeding (2024)
-
Analysis of co-expression and gene regulatory networks associated with sterile lemma development in rice
BMC Plant Biology (2023)
-
FRIZZLE PANICLE (FZP) regulates rice spikelets development through modulating cytokinin metabolism
BMC Plant Biology (2023)
-
Genomic introgressions from African rice (Oryza glaberrima) in Asian rice (O. sativa) lead to the identification of key QTLs for panicle architecture
BMC Genomics (2023)
-
Transcriptome-wide association analyses reveal the impact of regulatory variants on rice panicle architecture and causal gene regulatory networks
Nature Communications (2023)