Abstract
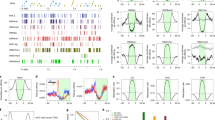
The non-random three-dimensional organization of genomes is critical for many cellular processes. Recently, analyses of genome-wide chromatin packing in the model dicot plant Arabidopsis thaliana have been reported1,2,3,4. At a kilobase scale, the A. thaliana chromatin interaction network is highly correlated with a range of genomic and epigenomic features1,2,3,4. Surprisingly, topologically associated domains (TADs), which appear to be a prevalent structural feature of genome packing in many animal species, are not prominent in the A. thaliana genome1,2,4,5,6. Using a genome-wide chromatin conformation capture approach, Hi-C (ref. 7), we report high-resolution chromatin packing patterns of another model plant, rice. We unveil new structural features of chromatin organization at both chromosomal and local levels compared to A. thaliana, with thousands of distinct TADs that cover about a quarter of the rice genome. The rice TAD boundaries are associated with euchromatic epigenetic marks and active gene expression, and enriched with a sequence motif that can be recognized by plant-specific TCP proteins. In addition, we report chromosome decondensation in rice seedlings undergoing cold stress, despite local chromatin packing patterns remaining largely unchanged. The substantial variation found already in a comparison of two plant species suggests that chromatin organization in plants might be more diverse than in multicellular animals.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Feng, S. et al. Genome-wide Hi-C analyses in wild-type and mutants reveal high-resolution chromatin interactions in Arabidopsis. Mol. Cell 55, 694–707 (2014).
Grob, S., Schmid, M. W. & Grossniklaus, U. Hi-C analysis in Arabidopsis identifies the KNOT, a structure with similarities to the flamenco locus of Drosophila. Mol. Cell 55, 678–693 (2014).
Liu, C. et al. Genome-wide analysis of chromatin packing in Arabidopsis thaliana at single-gene resolution. Genome Res. 26, 1057–1068 (2016).
Wang, C. et al. Genome-wide analysis of local chromatin packing in Arabidopsis thaliana. Genome Res. 25, 246–256 (2015).
Dekker, J., Marti-Renom, M. A. & Mirny, L. A. Exploring the three-dimensional organization of genomes: interpreting chromatin interaction data. Nat. Rev. Genet. 14, 390–403 (2013).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Nagano, T. et al. Comparison of Hi-C results using in-solution versus in-nucleus ligation. Genome Biol. 16, 175 (2015).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Fransz, P., De Jong, J. H., Lysak, M., Castiglione, M. R. & Schubert, I. Interphase chromosomes in Arabidopsis are organized as well defined chromocenters from which euchromatin loops emanate. Proc. Natl Acad. Sci. USA 99, 14584–14589 (2002).
Wu, Y. et al. Euchromatic subdomains in rice centromeres are associated with genes and transcription. Plant Cell 23, 4054–4064 (2011).
Di Pierro, M., Zhang, B., Aiden, E. L., Wolynes, P. G. & Onuchic, J. N. Transferable model for chromosome architecture. Proc. Natl Acad. Sci. USA 113, 12168–12173 (2016).
Santos, A. P. & Shaw, P. Interphase chromosomes and the Rabl configuration: does genome size matter? J. Microsc. 214, 201–206 (2004).
Prieto, P., Santos, A. P., Moore, G. & Shaw, P. Chromosomes associate premeiotically and in xylem vessel cells via their telomeres and centromeres in diploid rice (Oryza sativa). Chromosoma 112, 300–307 (2004).
Dong, F. & Jiang, J. Non-Rabl patterns of centromere and telomere distribution in the interphase nuclei of plant cells. Chromosome Res. 6, 551–558 (1998).
Hou, C., Li, L., Qin, Z. S. & Corces, V. G. Gene density, transcription, and insulators contribute to the partition of the Drosophila genome into physical domains. Mol. Cell 48, 471–484 (2012).
Sexton, T. et al. Three-dimensional folding and functional organization principles of the Drosophila genome. Cell 148, 458–472 (2012).
Sexton, T. & Cavalli, G. The role of chromosome domains in shaping the functional genome. Cell 160, 1049–1059 (2015).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Mizuguchi, T. et al. Cohesin-dependent globules and heterochromatin shape 3D genome architecture in S. pombe. Nature 516, 432–435 (2014).
Le, T. B., Imakaev, M. V., Mirny, L. A. & Laub, M. T. High-resolution mapping of the spatial organization of a bacterial chromosome. Science 342, 731–734 (2013).
Crane, E. et al. Condensin-driven remodelling of X chromosome topology during dosage compensation. Nature 523, 240–244 (2015).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl Acad. Sci. USA 112, E6456–E6465 (2015).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Ulianov, S. V. et al. Active chromatin and transcription play a key role in chromosome partitioning into topologically associating domains. Genome Res. 26, 70–84 (2016).
Liu, C. & Weigel, D. Chromatin in 3D: progress and prospects for plants. Genome Biol. 16, 170 (2015).
Danisman, S. et al. Arabidopsis class I and class II TCP transcription factors regulate jasmonic acid metabolism and leaf development antagonistically. Plant Physiol. 159, 1511–1523 (2012).
Kosugi, S. & Ohashi, Y. DNA binding and dimerization specificity and potential targets for the TCP protein family. Plant J. 30, 337–348 (2002).
O’Malley, R. C. et al. Cistrome and epicistrome features shape the regulatory DNA landscape. Cell 165, 1280–1292 (2016).
Ma, J. et al. Comprehensive analysis of TCP transcription factors and their expression during cotton (Gossypium arboreum) fiber early development. Sci. Rep. 6, 21535 (2016).
Martin-Trillo, M. & Cubas, P. TCP genes: a family snapshot ten years later. Trends Plant Sci. 15, 31–39 (2010).
Yao, X., Ma, H., Wang, J. & Zhang, D. Genome-wide comparative analysis and expression pattern of TCP gene families in Arabidopsis thaliana and Oryza sativa. J. Integr. Plant Biol. 49, 885–897 (2007).
Li, S. The Arabidopsis thaliana TCP transcription factors: a broadening horizon beyond development. Plant Signal. Behav. 10, e1044192 (2015).
Probst, A. V. & Mittelsten Scheid, O. Stress-induced structural changes in plant chromatin. Curr. Opin. Plant Biol. 27, 8–16 (2015).
Rosa, S. & Shaw, P. Insights into chromatin structure and dynamics in plants. Biology 2, 1378–1410 (2013).
Yaffe, E. & Tanay, A. Probabilistic modeling of Hi-C contact maps eliminates systematic biases to characterize global chromosomal architecture. Nat. Genet. 43, 1059–1065 (2011).
Imakaev, M. et al. Iterative correction of Hi-C data reveals hallmarks of chromosome organization. Nat. Methods 9, 999–1003 (2012).
Servant, N. et al. HiTC: exploration of high-throughput ‘C’ experiments. Bioinformatics 28, 2843–2844 (2012).
Heathcote, A. Fitting wald and ex-Wald distributions to response time data: an example using functions for the S-PLUS package. Behav. Res. Methods Instrum. Comput. 36, 678–694 (2004).
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Zhang, W. et al. High-resolution mapping of open chromatin in the rice genome. Genome Res. 22, 151–162 (2012).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5, 621–628 (2008).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
He, Y. et al. MEIOTIC F-BOX is essential for male meiotic DNA double-strand break repair in rice. Plant Cell 28, 1879–1893 (2016).
Feng, C. M., Qiu, Y., Van Buskirk, E. K., Yang, E. J. & Chen, M. Light-regulated gene repositioning in Arabidopsis. Nat. Commun. 5, 3027 (2014).
Wegel, E., Koumproglou, R., Shaw, P. & Osbourn, A. Cell type-specific chromatin decondensation of a metabolic gene cluster in oats. Plant Cell 21, 3926–3936 (2009).
He, G. et al. Global epigenetic and transcriptional trends among two rice subspecies and their reciprocal hybrids. Plant Cell 22, 17–33 (2010).
Fang, Y. et al. Functional characterization of open chromatin in bidirectional promoters of rice. Sci. Rep. 6, 32088 (2016).
Zhang, K. et al. Differential deposition of H2A.Z in combination with histone modifications within related genes in Oryza sativa callus and seedling. Plant J. 89, 264–277 (2017).
Stroud, H. et al. Plants regenerated from tissue culture contain stable epigenome changes in rice. eL ife 2, e00354 (2013).
Acknowledgements
We thank C. Lanz and K. Fritschi for assistance with sequencing. We thank X. Gao and Y. He for their assistance in microscopy. We thank members of the Liu and Weigel laboratories for critical review and comments on the manuscript. This work was supported by Marie Curie Fellowship PIIF-GA-2012-327608 (C.L.), Deutsche Forschungsgemeinschaft LI 2862/1 (C.L.), a grant from the DFG Collaborative Research Center SFB1101 (D.W.) and the Max Planck Society (D.W.).
Author information
Authors and Affiliations
Contributions
C.L. and D.W. conceived and designed the experiments. Y.C. and J.W. performed FISH experiments. CL performed the Hi-C and RNA-seq experiments. C.L. and D.W. analysed and interpreted the data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Information
Supplementary Figures 1–11, Supplementary Tables 3 and 5, Supplementary References.
Supplementary Table 1
Statistics of Hi-C reads.
Supplementary Table 2
TADs identified in the rice genome.
Supplementary Table 4
RNA-seq analyses.
Supplementary Table 6
DI and HMM-state of rice chromatin.
Rights and permissions
About this article
Cite this article
Liu, C., Cheng, YJ., Wang, JW. et al. Prominent topologically associated domains differentiate global chromatin packing in rice from Arabidopsis . Nature Plants 3, 742–748 (2017). https://doi.org/10.1038/s41477-017-0005-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-017-0005-9
This article is cited by
-
A fine-scale Arabidopsis chromatin landscape reveals chromatin conformation-associated transcriptional dynamics
Nature Communications (2024)
-
Mapping nucleosome-resolution chromatin organization and enhancer-promoter loops in plants using Micro-C-XL
Nature Communications (2024)
-
High-resolution Hi-C maps highlight multiscale chromatin architecture reorganization during cold stress in Brachypodium distachyon
BMC Plant Biology (2023)
-
Remodeling of the 3D chromatin architecture in the marine microalga Nannochloropsis oceanica during lipid accumulation
Biotechnology for Biofuels and Bioproducts (2023)
-
3D organization of regulatory elements for transcriptional regulation in Arabidopsis
Genome Biology (2023)