Abstract
RBM20 is a disease-causing gene associated with dilated cardiomyopathy (DCM). The proband presented with the dilated phase of hypertrophic cardiomyopathy (HCM), and the mother also suffered from HCM. A missense variant of RBM20, p.Arg636His, previously reported as pathogenic in several families with DCM, was found in both the proband and the mother. Therefore, RBM20 p.Arg636His could be the causative variant for this familial HCM, and RBM20 might be a novel causative gene for HCM.
Similar content being viewed by others
Hypertrophic cardiomyopathy (HCM) is characterized by unexplained left ventricular (LV) hypertrophy and diastolic dysfunction. On the other hand, dilated cardiomyopathy (DCM) is characterized by a dilated LV and decreased LV contractility. Most HCM patients show preserved LV contraction; however, some patients transition to a morphology similar to that of DCM with dilated ventricles, called the dilated phase of HCM (d-HCM)1,2,3. Furthermore, it is often difficult to differentiate between DCM and d-HCM owing to their similar morphologies. Moreover, the disease-causing genes of HCM, including those encoding sarcomeres, have also been registered as disease-causing genes of DCM, suggesting that there is some etiological overlap between HCM and DCM4.
A 41-year-old female proband patient (Fig. 1A, III-3) presented to our hospital with exertional dyspnea. We performed cardiac magnetic resonance (CMR) imaging using a Magnetom Skyra 3 T system (Siemens Healthineers, Erlangen, Germany)5 and found that the left ventricular end-diastolic diameter (LVDd) was dilated to 64 mm. In addition, the left ventricular end-systolic diameter (LVDs) was 48 mm, the diameter of the intraventricular septum (IVS) was 8.9 mm, the lateral wall diameter (LAD) was 6.5 mm, the left ventricular ejection fraction (LVEF) was reduced to 48%, and there was no coronary artery stenosis; therefore, we initially diagnosed the patient with DCM (Fig. 1B). In contrast, her 68-year-old mother (II-2, Fig. 1A) was asymptomatic and did not have hypertension, diabetes, or coronary stenosis. The mother’s CMR imaging revealed asymmetric septal thickening of the left ventricle, with a maximum IVS diameter of 13.2 mm and an LAD of 7.0 mm. The LVDd was 51 mm, and the LVEF was 54% (Fig. 1B). Consequently, we diagnosed the mother with HCM. With myocardial remodeling and deterioration due to fibrosis, the extracellular volume (ECV) increases, which can be quantitatively estimated by T1 mapping from CMR6,7. T1 mapping on a midventricular short-axis slice of myocardium was performed using the modified look-locker inversion recovery sequence before and after the intravenous administration of 0.1 mmol/kg gadobutrol (Gadovist, Bayer Healthcare, Leverkusen, Germany) as the MRI contrast agent, gadolinium. ECV was calculated using the following formula: ECV = (1-hematocrit) × (ΔR1myocardium/ΔR1blood), where R1 = 1/T16,7. Hematocrit was measured before CMR imaging. These images were analyzed using Ziostation 2 (Ziosoft, Tokyo, Japan). The ECV fraction is usually higher when myocardial damage is more severe. ECV fraction mapping of the mother’s CMR image revealed an elevated ECV fraction, indicating myocardial remodeling and fibrosis centered in the IVS (Fig. 1B). The same procedure was performed on the proband’s CMR image, and the same ECV as that of her mother diagnosed with HCM was observed (Fig. 1B). Owing to the similarity of the increased ECV localization in both the mother and the daughter, we diagnosed the proband with d-HCM rather than DCM.
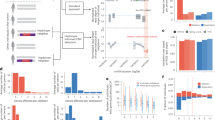
A Family pedigree showing the inheritance of cardiomyopathy. Squares denote males and circles denote females. Black symbols indicate affected individuals and open symbols indicate unaffected individuals. Gray symbols indicate individuals showing left ventricular hypertrophy on an electrocardiogram. The arrow indicates patients diagnosed with cardiomyopathy. P proband, d death, E genetic evaluation, + presence of RBM20 variant, − absence of RBM20 variant. B Short-axis images of cardiac magnetic resonance (CMR) for the proband (III-3) and her mother (II-2). The top and middle rows present end-diastolic and end-systolic cine images, respectively, and the bottom row shows extracellular volume (ECV) fraction mapping. Both III-3 and II-2 demonstrated elevated ECV (light green colored, arrowheads), primarily in the ventricular septum. RV right ventricle, LV left ventricle.
We conducted variant searches for 67 reported cardiomyopathy-causing genes in the peripheral blood genomic DNA of the proband patient who was diagnosed with d-HCM using an Ion Torrent PGM system (Thermo Fischer Scientific, CA, USA)8. This study was approved by the ethics committees of the local institutional review boards. A heterozygous missense variant, NM_001134363.3: c.1907 G > A, p.Arg636His, in the RBM20 gene, which encodes the cardiac splicing factor RNA-binding motif protein 20, was identified and confirmed via Sanger sequencing (Fig. 2A). This variant was registered as rs267607004 in the dbSNP variant database. In the Genome Aggregation Database (gnomAD), which contains variants from the general population and multiple ethnic groups (https://gnomad.broadinstitute.org), the minor allele frequency of this variant is 0.000014, which is very rare; moreover, this variant is not registered in the Human Genetic Variation Database (HGVD), which is specific to the Japanese population (https://www.hgvd.genome.med.kyoto-u.ac.jp). This variant is located in an arginine/serine (RS)-rich region, specifically, the arginine-serine-arginine-serine-proline residues called the RSRSP stretch domain (Fig. 2B). The phosphorylation of the RSRSP stretch domain, which is a critical region, is required for the nuclear localization of RBM209. Additionally, there have been previous reports of multiple families with DCM with the same variant, RBM20 p.Arg636His, and the rare variant is registered as pathogenic/likely pathogenic in the ClinVar database (https://www.ncbi.nlm.nih.gov/clinvar/)10,11. Therefore, we concluded that RBM20 p.Arg636His was the disease-causing variant in the proband. A family pedigree analysis of this variant revealed that the proband’s father (Fig. 1A, II-1), who had no heart disease, did not have the variant, whereas her mother (Fig. 1A, II-2), who was diagnosed with HCM, had the same variant. The proband’s daughter (Fig. 1A, IV-1) showed left ventricular hypertrophy on an electrocardiogram but normal echocardiography findings, and she had the same RBM20 variant as her mother.
A Sanger sequencing results of the RBM20 (NM_001134363.3) gene in healthy controls (left panel) and patients (right panel) are shown. The patients carry a heterozygous missense variant, NM_001134363.3 (RBM20_v001): c.1907 G > A, p.Arg636His. B In the RS region (marked in black) located in codons 613 to 673 of RBM20, all variants (upper row) of RBM20 that were registered as “pathogenic” or “likely pathogenic” in the ClinVar database are located in the RSRSP stretch domain (codons 634 to 638), shown in red. These variants were all found in patients with DCM. The variant (RBM20 p.R636H) found in this family with HCM (bottom row) is also located in the RSRSP stretch domain. DCM dilated cardiomyopathy, HCM hypertrophic cardiomyopathy.
RBM20 is one of the major disease-causing genes in DCM4,12. The RBM20 protein is involved in splicing, particularly in regulating the splicing of Titin, which is a major causative gene of DCM9. Genetic variants of RBM20 cause abnormal splicing of the Titin gene, which results in DCM. However, RBM20 is involved in the splicing of more than 30 genes, including sarcomere protein-coding genes such as MYH7 and TNNT2, which are the major causative genes of HCM9. Additionally, some RBM20 variants reportedly increase calcium sensitivity in the myocardium13. A number of HCM-causing variants in sarcomere genes increase calcium sensitivity in the myocardium14,15. These findings are consistent with the common functional alterations observed in HCM. Collectively, these results suggest that the RBM20 p.Arg636His variant, which has been reported in families with DCM, could be the disease-causing variant in this family with HCM. The penetrance of the RBM20 p.Arg636His variant has been reported to be 96–100% in patients over 30 years of age10,11. Therefore, the proband’s daughter (Fig. 1A, IV-1), carrying the same RBM20 variant as the proband, has a high possibility of developing cardiomyopathy in the future and thus should be carefully monitored. In addition, it might be necessary to investigate whether the proband’s mother has any protective factor for myocardial dilatation. Moreover, subclinical cases of HCM in the reported families with DCM with the RBM20 p.Arg636His variant may be identified in further family studies. Further studies are required, but the RBM20 gene could be a potential cause of HCM.
HGV database
The relevant data from this Data Report are hosted at the Human Genome Variation Database at https://doi.org/10.6084/m9.figshare.hgv.3122.
References
Maron, B. J., Rowin, E. J., Casey, S. A. & Maron, M. S. How hypertrophic cardiomyopathy became a contemporary treatable genetic disease with low mortality: shaped by 50 years of clinical research and practice. JAMA Cardiol. 1, 98–105 (2016).
McKenna, W. J., Maron, B. J. & Thiene, G. Classification, epidemiology, and global burden of cardiomyopathies. Circ. Res. 121, 722–730 (2017).
Kitaoka, H. et al. JCS/JHFS 2018 guideline on the diagnosis and treatment of cardiomyopathies. Circ. J. 85, 1590–1689 (2021).
Hershberger, R. E. et al. Genetic evaluation of cardiomyopathy: a clinical practice resource of the American College of Medical Genetics and Genomics (ACMG). Genet. Med. 20, 899–909 (2018).
Tezuka, D. et al. Myocardial perfusion reserve quantified by cardiac magnetic resonance imaging is associated with late gadolinium enhancement in hypertrophic cardiomyopathy. Heart Vessels 33, 513–520 (2018).
Martinez-Naharro, A. et al. Native T1 and extracellular volume in transthyretin amyloidosis. JACC Cardiovasc. Imaging 12, 810–819 (2019).
Robinson, A. A., Chow, K. & Salerno, M. Myocardial T1 and ECV measurement: underlying concepts and technical considerations. JACC Cardiovasc. Imaging 12, 2332–2344 (2019).
Hayashi, T. et al. Genetic background of Japanese patients with pediatric hypertrophic and restrictive cardiomyopathy. J. Hum. Genet. 63, 989–996 (2018).
Lennermann, D., Backs, J. & van den Hoogenhof, M. M. G. New insights in RBM20 cardiomyopathy. Curr. Heart Fail Rep. 17, 234–246 (2020).
Brauch, K. M. et al. Mutations in ribonucleic acid binding protein gene cause familial dilated cardiomyopathy. J. Am. Coll. Cardiol. 54, 930–941 (2009).
Wells, Q. S. et al. Whole exome sequencing identifies a causal RBM20 mutation in a large pedigree with familial dilated cardiomyopathy. Circ. Cardiovasc. Genet. 6, 317–326 (2013).
Haas, J. et al. Atlas of the clinical genetics of human dilated cardiomyopathy. Eur. Heart J. 36, 1123–1135a (2015).
Beqqali, A. et al. A mutation in the glutamate-rich region of RNA-binding motif protein 20 causes dilated cardiomyopathy through missplicing of titin and impaired Frank-Starling mechanism. Cardiovasc. Res. 112, 452–463 (2016).
Marian, A. J. & Braunwald, E. Hypertrophic cardiomyopathy: genetics, pathogenesis, clinical manifestations, diagnosis, and therapy. Circ. Res. 121, 749–770 (2017).
Morimoto, S., Yanaga, F., Minakami, R. & Ohtsuki, I. Ca2+-sensitizing effects of the mutations at Ile-79 and Arg-92 of troponin T in hypertrophic cardiomyopathy. Am. J. Physiol. 275, C200–C207 (1998).
Acknowledgements
We would like to thank Yukiko Ueda and Chinami Sonobe for their technical assistance with respect to sequencing using the Ion Torrent PGM system and Sanger sequencing. This work was supported by the Japan Society for the Promotion of Science KAKENHI grant numbers 26460407 (T.H.), 17K08684 (T.H.), 15K15095 (A.K.), 16H05296 (A.K.), and 21K08089 (N.I.), by Tokai University School of Medicine, Project Research, and by Nanken-Kyoten, Tokyo Medical and Dental University.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Inagaki, N., Hayashi, T., Takei, Y. et al. Pathogenic variant of RBM20 in a multiplex family with hypertrophic cardiomyopathy. Hum Genome Var 9, 6 (2022). https://doi.org/10.1038/s41439-022-00183-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41439-022-00183-z