Abstract
Asymptomatic hyper-CK-emia (ASCK) is characterized by persistent elevation of creatine kinase (CK) in serum without any neurological symptoms. We ascertained a two-generation family of ASCK patients without clear neurological abnormalities except for the high levels of serum CK (810.5 ± 522.4 U/L). We identified a novel 1-bp deletion variant in the DAG1 gene shared by the patients in the family (NM_001177639: exon 3: c.930delC:p.R311Gfs*70). The variant causes premature termination of translation at codon 477, resulting in a protein product completely devoid of the essential DAG1 domain. Since ASCK has been associated with DAG1 in only one case carrying compound heterozygous missense variants, our new finding of a novel 1-bp deletion revealed the previously unknown dominant effect of DAG1 on ASCK.
Similar content being viewed by others
Serum creatine kinase (CK) levels are clinically important for the diagnosis of patients with muscle weakness, myopathy, or rhabdomyolysis. However, persistent and abnormally high levels of serum CK are occasionally observed in normal persons without any symptoms, which is termed aymptomatic hyper-CK-emia (ASCK). Although most ASCK cases, including idiopathic hyper-CK-emia (IHCK), are known to be sporadic, more than 20 pedigrees have been reported to be familial1,2,3,4,5,6,7,8,9,10,11.
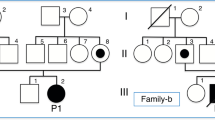
We studied a two-generation Japanese pedigree of ASCK from Hiroshima Prefecture, a western province of Japan. The mode of inheritance in the ASCK pedigree is suggested to be autosomal dominant by the presence of male to male transmission (Fig. 1). All family members had no inconvenience in their daily activities. Three children (II-1, II-2, II-3) were unable to do a back-hip circle. All four affected members (I-1, II-1, II-2, II-3) had poor long-distance running. They had no clear neurological abnormalities except for the elevated levels of serum CK (810.5 ± 522.4 U/L). The ages of three children (II-1, II-2, II-3) were 10, 11, and 14 years old at the time of blood collection, respectively.
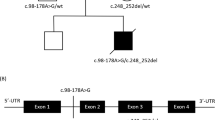
Whole-exome sequencing was carried out for two affected family members, II-2 and II-3, with sequencing depths of 69x and 50x, respectively (Fig. 1). Out of 124,432 single nucleotide variants (SNVs) identified in total, we selected 31,533 SNVs shared by both patients. We identified 315 SNVs located in 127 candidate genes of IHCK, malignant hyperthermia, and muscular dystrophy, including 151 functional SNVs (Supplementary Table 1). We excluded variants with frequencies larger than 0.002 in the 1000 G Project [http://www.1000genomes.org], Ensembl [http://asia.ensembl.org/index.html], HGVD [http://www.hgvd.genome.med.kyoto-u.ac.jp/], ToMMo [http://www.megabank.tohoku.ac.jp/], and ExAC [http://exac.broadinstitute.org/] databases, with the remaining five SNVs located in four genes (DAG1, RYR1, SYNE1, TTN). By Sanger sequencing using the primers shown in Supplementary Table 2, we identified only one SNV validated and cosegregated in the pedigree, which was a novel 1-bp deletion variant in exon 3 in the DAG1 gene, NM_001177639:exon3:c.930delC:p.R311Gfs*70. The SNV is expected to cause a frameshift and premature termination of translation at codon 477 (Fig. 2, Supplementary Figure 1). We confirmed the absence of the SNV in 506 unrelated Japanese controls, indicating a frequency < 0.099% in the Japanese normal population.
The DAG1 gene encodes dystroglycan protein (NP_004384, Fig. 3), which is a dystrophin-associated glycoprotein (DAG) that is known to be responsible for muscular dystrophy (MD) and muscle-eye-brain disease12,13. Although the elevation of serum CK is commonly observed in MD patients, the current pedigree shows no neurological abnormalities related to MD. The premature termination of the translation caused by the 1-bp deletion in DAG1 results in a protein product completely devoid of the DAG1 domain, which is known to be essential for DAG1 function (Fig. 3)14,15. Since ASCK has been associated with DAG1 in only one case carrying compound heterozygous missense variants in DAG1, our new finding of a novel 1-bp deletion shows the previously unknown dominant effect of DAG1 for ASCK16. Dong et al. (2015) also observed very mild MD diagnosed only by immunohistochemistry16. The current patients, therefore, may show a similar subclinical MD, although immunohistochemical examination is not applicable due to the lack of muscle tissue samples. According to the ACMG/AMP/CAP guidelines, the 1-bp deletion variant in DAG1 is classified as “pathogenic”, meeting the criteria of PVS1, PM1, PM2, PM4, PP1, and PP2. Therefore, we conclude that the novel 1-bp deletion in DAG1, NM_001177639:exon3:c.930delC:p. R311Gfs*70 is the causative dominant variant for the ASCK family.
Locations of the cadherin-like repeat domains of alpha dystroglycan (CADA), alpha-dystroglycan N-terminal domain 2 (a_DG1_N2), dystrophin-associated glycoprotein 1 (DAG1) are shown according to the Conserved Domains Database (CDD) (https://www.ncbi.nlm.nih.gov/Structure/cdd/cdd.shtml) and the Subfamily Protein Architecture Labeling Engine (SPARCLE) (https://www.ncbi.nlm.nih.gov/Structure/sparcle/docs/sparcle_about.html). Red and blue arrows indicate the locations of the variant and the premature stop codon, respectively.
HGV Database
The relevant data from this Data Report are hosted at the Human Genome Variation Database at https://doi.org/10.6084/m9.figshare.hgv.3125.
References
Rowland L. P. et al. Approaches to the membrane theory of Duchenne muscular dystrophy. Muscular dystrophy research: advances and new trends. Amsterdam: Excerpta Medica. 3–13 (1980).
Cagliani, R. et al. A CAV3 microdeletion differentially affects skeletal muscle and myocardium. Neurology 61, 1513–1519 (2003).
Sugie, K. et al. Two novel CAV3 gene mutations in Japanese families. Neuromuscul. Disord. 14, 810–814 (2004).
Capasso, M. et al. Familial idiopathic hyper-CK-emia: an underrecognized condition. Muscle Nerve. 33, 760–765 (2006).
Traverso, M. et al. Caveolin-3 T78M and T78K missense mutations lead to different phenotypes in vivo and in vitro. Lab Invest. 88, 275–283 (2008).
González-Pérez, P. et al. Phenotypic variability in a Spanish family with a Caveolin-3 mutation. J. Neurol. Sci. 276, 95–98 (2009).
Sano, K. et al. A novel missense mutation of RYR1 in familial idiopathic hyper CK-emia. J. Neurol. Sci. 356, 142–147 (2015).
Macias, A. et al. CAV3 mutation in a patient with transient hyperCKemia and myalgia. Neurol. Neurochir. Pol. 50, 468–473 (2016).
Zullo, A. et al. RYR1 sequence variants in myopathies: expression and functional studies in two families. Biomed. Res. Int. 2019, 7638946 (2019).
Rubegni, A. et al. Next-generation sequencing approach to hyperCKemia A 2-year cohort study. Neurol. Genet. 5, e352 (2019).
Bruno, G. et al. A novel missense mutation in CAV3 gene in an Italian family with persistent hyperCKemia, myalgia and hypercholesterolemia: Double-trouble. Clin. Neurol. Neurosurg. 191, 105687 (2020).
Hara, Y. et al. A dystroglycan mutation associated with limb-girdle muscular dystrophy. N. Engl. J. Med. 364, 939–946 (2011).
Geis, T. et al. Homozygous dystroglycan mutation associated with a novel muscle-eye-brain disease-like phenotype with multicystic leukodystrophy. Neurogenetics 14, 205–213 (2013).
Roberds, S. L., Anderson, R. D., Ibraghimov-Beskrovnaya, O. & Campbell, K. P. Primary structure and muscle-specific expression of the 50-kDa dystrophin-associated glycoprotein (adhalin). J. Biol. Chem. 268, 23739–23742 (1993).
Cohn, R. D. et al. Disruption of DAG1 in differentiated skeletal muscle reveals a role for dystroglycan in muscle regeneration. Cell 110, 639–648 (2002).
Dong, M. et al. DAG1 mutations associated with asymptomatic hyperCKemia and hypoglycosylation of α-dystroglycan. Neurology 84, 273–279 (2015).
Acknowledgements
We are grateful to the family for allowing us to report their cases. Written informed consent was obtained from all participating individuals. This study was approved by the Ethics Committees of Kurume University School of Medicine (#79) and of Kyushu University, Faculty of Medicine (#397-01). This work was supported by the Cooperative Research Project Program of the Medical Institute of Bioregulation, Kyushu University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fan, L., Miura, S., Shimojo, T. et al. A novel 1-bp deletion variant in DAG1 in Japanese familial asymptomatic hyper-CK-emia. Hum Genome Var 9, 4 (2022). https://doi.org/10.1038/s41439-022-00182-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41439-022-00182-0
This article is cited by
-
DAG1 haploinsufficiency is associated with sporadic and familial isolated or pauci-symptomatic hyperCKemia
European Journal of Human Genetics (2024)