Abstract
We report a patient with developmental delay, extremely short stature, small hands, dysmorphic facial features, hearing loss, and epilepsy carrying a de novo 2.76-Mb deletion of 2q36.3q37.1, including TRIP12 and NPPC. TRIP12 haploinsufficiency causes developmental delay with isolated dysmorphic facial features, whereas NPPC haploinsufficiency causes short stature and small hands. This is the first report of a unique phenotype, which is secondary to a microdeletion encompassing TRIP12 and NPPC.
Similar content being viewed by others
Thyroid hormone receptor interacting protein 12 (TRIP12) is a human protein homologous to the E6-AP carboxyl terminus (HECT) domain E3 ubiquitin ligase and is involved in cell cycle progression and the maintenance of chromosome integrity1. TRIP12 (MIM604506) haploinsufficiency has been reported to cause developmental delay, autism spectrum disorder (ASD), and facial dysmorphisms, which are collectively named Clark–Baraitser syndrome (MIM #617752)2,3,4.
C-type natriuretic peptide (CNP) plays an important role in cartilage homeostasis and endochondral bone formation and is encoded by the gene natriuretic peptide precursor-C (NPPC; MIM600296)5. NPPC variants have been reported to cause autosomal dominant idiopathic short stature with small hands6. Here, we describe a case of a 4-year-old boy with a de novo 2q36.3q37.1 deletion encompassing TRIP12 and NPPC who presented with severe developmental delay, extremely short stature, small hands, dysmorphic facial features, hearing loss, and epilepsy.
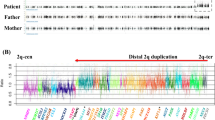
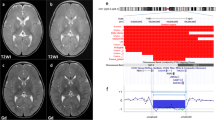
The boy was born to healthy nonconsanguineous Japanese parents at 40 gestational weeks. His parents had no medical history. At birth, his weight, length, and occipitofrontal circumference (OFC) were 2710 g (−0.7 standard deviation (SD)), 46.6 cm (−1.1 SD), and 33 cm (−0.4 SD), respectively. Automated auditory brainstem response detection in a neonatal hearing screening test revealed a refer outcome. Urinal cytomegalovirus DNA was not detected in the neonatal period. Hearing threshold levels were bilateral sensorineural hearing loss of 70 dB at the age of 6 months and 30–40 dB at 4 years. He showed severe developmental delay and could hold up his head at 9 months, sit alone at 14 months, and stand with support at ~2 years. At 4 years, when this report was written, he was unable to speak or stand alone. He displayed behavioral abnormalities, such as aggression, repeatedly biting his intravenous drip line during hospitalization, and frequently striking his head against the wall. His short stature became apparent by 6 months of age (Fig. 1a). His height, weight, and OFC were 76.5 cm (−4.0 SD), 8.5 kg (−3.3 SD), and 50.5 cm (+0.9 SD) at 3 years, respectively, indicating relative macrocephaly (OFC more than +1 SD compared to height). We performed several examinations to evaluate his severe short stature. Complete blood count, blood chemistry, and thyroid function tests showed normal results. Insulin-like growth factor-1 was 46 mg/mL (−1.1 SD) at 3 years. His karyotype was 46, XY. A growth hormone stimulation test was not performed because his parents did not provide consent. He experienced four febrile seizures and two afebrile seizures at 2 years. His electroencephalogram showed frequent spike or spike-and-wave discharges in the left parietal region during drowsy and sleep states. He was diagnosed with epilepsy, and levetiracetam was started at 3 years. Head magnetic resonance imaging showed normal results. His facial features included a prominent forehead, mild hypertelorism, epicanthal folds, short nose, low hanging columella, short philtrum, everted vermilion of the upper lip, strabismus, and low-set ears (Fig. 1b, c), and his hands were small (Fig. 1d). After obtaining written informed consent, we performed array comparative genomic hybridization analysis and identified a de novo 2.76 Mb deletion (arr[GRCh37] 2q36.3q37.1(230377789_233136164)x1) encompassing 25 protein-coding genes; of these genes, TRIP12 and NPPC are known to affect phenotypes attributable to autosomal dominant inheritance and cause developmental delay and short stature with small hands, respectively. Among the 25 genes, TRIP12, NCL, and PSMD1 each have a “probability of being loss-of-function intolerant” (pLI) score of 1.0, indicating that these three genes are probably intolerant to a loss-of-function variation7. NPPC is classified as tolerant to loss-of-function variation because the phenotype caused by NPPC haploinsufficiency is not thought to confer significant survival or reproductive disadvantages. In addition, although abnormalities in SP110, ARMC9, PDE6D, and DIS3L2, which reside in the deleted region, have been reported to cause autosomal recessive inheritance diseases, our patient displayed different clinical features than those observed with these diseases8.
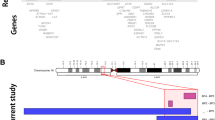
a The patient’s growth curve. b, c The patient exhibited prominent forehead, mild hypertelorism, epicanthal folds, short nose, low hanging columella, short philtrum, everted vermilion of the upper lip, strabismus, low-set ears, and relative macrocephaly at 4 years of age. We obtained written informed consent from the patient’s family for publication of this clinical report and accompanying images. d The patient presented with small hands. e Twenty-five genes in the 2.76-Mb 2q36.3q37.1 deletion (230377789_233136164) and deleted region of our case and previously reported copy number variant (CNV) deletion cases involving TRIP12 or NPPC in Table 1.
Genetic findings related to TRIP12 mainly include rare de novo variants, which have been identified from several large ASD or ID proband cohorts by whole-exome or target sequencing9,10,11,12,13,14. However, these reports did not describe detailed patient clinical information. Regarding clinical features, Bramswig et al. and Zang et al. reported 11 patients, including four previously reported cases9,10,15 and nine patients with inactivating single nucleotide variants or copy number variant (CNV) deletions in TRIP12, respectively, in 20172,3.
CNP encoded by NPPC plays an important role in skeletal development. Additionally, NPPC alteration in humans was reported by Tassano et al. for the first time in a boy with short stature and several dysmorphic features; he had a 1.91 Mb 2q37 deletion, including NPPC (case-N1 in Fig. 1e and Table 1)16. Two NPPC variants were identified in six patients with short stature and small hands from only two families by Hisado-Oliva et al.6. They also showed that COP-7 cells transfected with NPPC variants detected in the families showed significantly reduced CNP-dependent cGMP synthesis even in the heterozygous state.
PSMD1 (proteasome 26S subunit, non-ATPase, 1, MIM617842) encodes a regulatory subunit of the 26S proteasome, which is a major ATP-dependent intracellular proteinase17. NCL (nucleolin, MIM164035) encodes a eukaryotic nucleolar phosphoprotein and is associated with the synthesis and maturation of ribosomes18. No specific phenotypes associated with PSMD1 or NCL haploinsufficiency have been reported, whereas only one patient carrying a de novo NCL variant was identified by exome sequencing of 4293 families with developmental disorders19. Given that the deletion included four genes associated with autosomal recessive inheritance diseases, the major limitation of our study is that we did not analyze a hemizygous variation of the undeleted allele.
Eight CNV deletions smaller than 6 Mb involving TRIP12 have been reported3,20,21. In addition, we identified five CNV deletions with phenotype information in the DECIPHER database v9.318. None of these 13 CNVs included NPPC, NCL, or PSMD1, and five CNVs were intragenic deletions within TRIP12. The remaining eight deletions involved adjacent genes and TRIP12 (case-T1–T8 in Fig. 1e and Table 1). For NPPC, case N1 was the only reported patient with a CNV deletion smaller than 6 Mb involving NPPC, and the deletion included NCL and PSMD1 (case N1 in Fig. 1e and Table 1).
The clinical features associated with TRIP12 alterations have been previously summarized2,3. Clinical features such as developmental delay, hearing loss, and epilepsy in our patients could be attributed to TRIP12 haploinsufficiency. Our patient showed epicanthic folds, hypertelorism, short nose, low hanging columella, and low-set ears, which partly overlapped with the dysmorphic facial features observed in patients with TRIP12 alterations2,3. However, the low-set ears, prominent forehead, strabismus, short philtrum, everted vermilion of the upper lip, and relative macrocephaly in our patient were considerably similar to those in case N1. Case N1 had 19 deleted genes, including NPPC, NCL, and PSMD1, and all of them were included within the deletion range of our case. Recognizable facial dysmorphisms in patients with NPPC alterations have not been reported. Therefore, the characteristic facies in Case N1 and in our case is not explained by NPPC deletion but by other deleted genes, including NCL and PSMD1. Given the considerable facial similarity between Case 1 and our case, we speculated that facial dysmorphism in our case would be affected more by the 19 deleted genes than by TRIP12 alteration.
Our patient showed developmental delay and short stature to a more severe extent than previously described for patients with TRIP12 alterations. He was not able to speak any words or walk independently at 49 months of age, although previously described patients with TRIP12 alterations such as case T1 and case T2 could speak their first words and walk independently at a mean age of 28 months (range 10–60, n = 15) and 20 months (range 13–29, n = 16), respectively2,3. Their heights were variable, and the mean height SD score was −0.3 SD (range −2.2 to +1.1, n = 16). Short stature and small hands in our patient were thought to be mainly affected by the NPPC deletion. However, NPPC alterations alone do not explain his extremely short stature. The mean height SD score of the six patients with NPPC heterozygous variants (−2.9 SD, range −4.3 to −2.3) was 0.9 SD lower than that of the five patients without NPPC variants (−2.0 SD, range −4.8 to −0.4) in the two reported families6. Importantly, case T4, with the largest deletion expanding in the centromeric direction, showed a normal height and moderate developmental delay (initial walk at 22 months). Thus, genes other than TRIP12 and NPPC located in the deletion region might have contributed to the exaggerated phenotypes in terms of developmental delay and short stature. Because of extreme intolerance to loss-of-function variations, NCL and PSMD1 could be good candidate genes for the exaggerated phenotypes.
In conclusion, we report a patient with a de novo 2.76-Mb deletion of 2q36.3q37.1 encompassing TRIP12 and NPPC. This is the first report of a patient with a unique phenotype of combined TRIP12 and NPPC haploinsufficiency. The severe developmental delay and extremely short stature of the patient imply that NCL and PSMD1 are potentially disease-modulating genes.
HGV database
The relevant data from this Data Report are hosted at the Human Genome Variation Database at https://doi.org/10.6084/m9.figshare.hgv.2864; https://doi.org/10.6084/m9.figshare.hgv.2867.
References
Larrieu, D. et al. The E3 ubiquitin ligase TRIP12 participates in cell cycle progression and chromosome stability. Sci. Rep. 10, 789 (2020).
Bramswig, N. C. et al. Identification of new TRIP12 variants and detailed clinical evaluation of individuals with non-syndromic intellectual disability with or without autism. Hum. Genet 136, 179–192 (2017).
Zhang, J. et al. Haploinsufficiency of the E3 ubiquitin-protein ligase gene TRIP12 causes intellectual disability with or without autism spectrum disorders, speech delay, and dysmorphic features. Hum. Genet 136, 377–386 (2017).
Louie, R. J. et al. Clark-Baraitser syndrome is associated with a nonsense alteration in the autosomal gene TRIP12. Am. J. Med. Genet A 182, 595–596 (2020).
Peake, N. J. et al. Role of C-type natriuretic peptide signalling in maintaining cartilage and bone function. Osteoarthr. Cartil. 22, 1800–1807 (2014).
Hisado-Oliva, A. et al. Mutations in C-natriuretic peptide (NPPC): a novel cause of autosomal dominant short stature. Genet Med. 20, 91–97 (2018).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Firth, H. V. et al. DECIPHER: database of chromosomal imbalance and phenotype in humans using ensembl resources. Am. J. Hum. Genet 84, 524–533 (2009).
O’Roak, B. J. et al. Recurrent de novo mutations implicate novel genes underlying simplex autism risk. Nat. Commun. 5, 5595 (2014).
Lelieveld, S. H. et al. Meta-analysis of 2,104 trios provides support for 10 new genes for intellectual disability. Nat. Neurosci. 19, 1194–1196 (2016).
Wang, T. et al. De novo genic mutations among a Chinese autism spectrum disorder cohort. Nat. Commun. 7, 13316 (2016).
Rossi, M. et al. Outcomes of diagnostic exome sequencing in patients with diagnosed or suspected autism spectrum disorders. Pediatr. Neurol. 70, 34–43 e32 (2017).
Takata, A. et al. Integrative analyses of de novo mutations provide deeper biological insights into autism spectrum disorder. Cell Rep. 22, 734–747 (2018).
Kahrizi, K. et al. Effect of inbreeding on intellectual disability revisited by trio sequencing. Clin. Genet 95, 151–159 (2019).
Iossifov, I. et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 515, 216–221 (2014).
Tassano, E. et al. Genotype-phenotype correlation of 2q37 deletions including NPPC gene associated with skeletal malformations. PLoS ONE 8, e66048 (2013).
Yokota, K. et al. CDNA cloning of p112, the largest regulatory subunit of the human 26s proteasome, and functional analysis of its yeast homologue, sen3p. Mol. Biol. Cell 7, 853–870 (1996).
Srivastava, M., Fleming, P. J., Pollard, H. B. & Burns, A. L. Cloning and sequencing of the human nucleolin cDNA. FEBS Lett. 250, 99–105 (1989).
Deciphering Developmental Disorders S. Prevalence and architecture of de novo mutations in developmental disorders. Nature 542, 433–438 (2017).
Doco-Fenzy, M. et al. Deletion 2q36.2q36.3 with multiple renal cysts and severe mental retardation. Eur. J. Med. Genet 51, 598–607 (2008).
Pinto, D. et al. Convergence of genes and cellular pathways dysregulated in autism spectrum disorders. Am. J. Hum. Genet 94, 677–694 (2014).
Acknowledgements
We are grateful to the patient and his family for participating in this study. Array comparative genomic hybridization was performed by TOKAI-IRUD. This study was partially supported by the Initiative on Rare and Undiagnosed Diseases in Pediatrics from the Japanese Agency for Medical Research and Development. This study makes use of data generated by the DECIPHER community. A full list of centers who contributed to the generation of the data is available from http://decipher.sanger.ac.uk and via email from decipher@sanger.ac.uk. Funding for the project was provided by Welcome. This study was partially supported by the Initiative on Rare and Undiagnosed Diseases in Pediatrics from the Japanese Agency for Medical Research and Development.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kondo, Y., Aoyama, K., Suzuki, H. et al. De novo 2q36.3q37.1 deletion encompassing TRIP12 and NPPC yields distinct phenotypes. Hum Genome Var 7, 19 (2020). https://doi.org/10.1038/s41439-020-0107-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41439-020-0107-1