Abstract
Pathogenic FLNA variants can be identified in patients with seizures accompanied by periventricular nodular heterotopia (PVNH). It is unusual to find FLNA aberrations in epileptic patients without PVNH on brain imaging. We report a boy with cryptogenic West syndrome followed by refractory seizures and psychomotor delay. We performed whole-exome sequencing and identified a de novo missense variant in FLNA. It is noteworthy that this patient showed no PVNH. As no other pathogenic variants were found in epilepsy-related genes, this FLNA variant likely caused West syndrome but with no PVNH.
Similar content being viewed by others
In epileptic encephalopathy, epileptic activity contributes to severe cognitive and behavioral impairments1. Genetic causes can be detected in patients with epileptic encephalopathy, including age-dependent epilepsy in infancy such as West syndrome2.
The FLNA gene at Xq28 encodes the filamin A protein, which is known to interact with more than 90 other proteins that could involve neuronal migration and other functions3,4,5.
Pathogenic FLNA variants are known to cause several human phenotypes6. Loss-of-function variants in FLNA cause periventricular nodular heterotopia (PVNH1) (MIM #300017) or/and congenital intestinal pseudo-obstruction. Gain-of-function mutations in FLNA cause otopalatodigital spectrum disorders. Cardiac valvular dystrophy is observed in patients with both loss-of-function and gain-of-function variants, suggesting a different mechanism involved in valvular dystrophy. FLNA pathogenic variants could also cause thrombocytopenia through aberrant activation of GPIbα and αIIbβ3 integrin, which are receptors of FLNa and essential to platelet adhesion and aggregation4.
Seizure is a common symptom in PVNH patients (more than 70%)7. Based on the X-linked dominant inheritance pattern of FLNA aberration, FLNA-related PVNH patients are usually females (more than 90%)7, and FLNA variants in males might be lethal in association with high miscarriage rates in mothers affected with PVNH1 as well as high infantile mortality in affected boys8,9. The reason for such high mortality remains unclear, but early deaths in males could arise from hemorrhage3 or cardiovascular malformation10 but not from brain malformation. However, approximately 30 male patients whose FLNA variants are predicted to be partially loss-of-function or mosaic have been reported to date11. The Flna-null mouse model showed that they died at midgestation with widespread hemorrhage from abnormal vessels and truncus arteriosus12.
Here, we report a boy with West syndrome arising from a de novo FLNA variant detected by whole-exome sequencing (WES), but no PVNH was seen by brain MRI.
All human studies were approved by the institutional review boards of Yokohama City University, Showa University and Fukushima Medical University. Written informed consent was obtained from the parents of the patient.
WES was performed on the patient’s DNA. Genomic DNA was captured by the SureSelect Human All Exon v6 system (Agilent Technologies, Santa Clara, CA, USA) and sequenced on the HiSeq 2500 platform (Illumina, San Diego, CA, USA) as described previously13. The mean WES coverage was 71.07x, and at least 89.8% coverage of the target regions with 20 or more reads was achieved. To identify causative variants of infantile spasms, we narrowed down variants in our patient based on the allele frequency (<0.001 for autosomal dominant model, <0.01 for autosomal recessive model, and <0.01 for X-linked model) using the Human Genetic Variation Database (HGVD) (http://www.hgvd.genome.med.kyoto-u.ac.jp/), the Exome Aggregation Consortium (ExAC) (http://exac.broadinstitute.org/), the Tohoku Medical Megabank Organization (ToMMo) (https://www.megabank.tohoku.ac.jp/english/) and the Genome Aggregation Database (gnomAD) (https://gnomad.broadinstitute.org/). We also used our in-house whole-exome database of 575 Japanese control individuals and excluded nonpathogenic variants by their allele frequency. After selecting the variants in the database according to frequency, we determined whether the remaining variants were deleterious using three prediction tools recommended in the American College of Medical Genetics and Genomics (ACMG) Standards and Guidelines: SIFT (https://sift.bii.a-star.edu.sg/), Polyphen-2 (http://genetics.bwh.harvard.edu/pph2/) and CADD (https://cadd.gs.washington.edu/).
Copy number variations (CNVs) were also detected from the WES data using the eXome-Hidden Markov Model as previously described14.
To determine whether the deleterious variant detected by WES was a de novo variant, we sequenced the DNA of the parents and patient using the Sanger method. For polymerase chain reaction (PCR) analysis, we used Takara Ex Taq HS polymerase (Takara Bio, Shiga, Japan) and two primers (forward 5′-CTT TTG GGC CAT AGC AGT TAA GA-3′; reverse 5′-CAG TGC ACT TGC TGG CGT CC-3′) with the following PCR conditions: denaturation at 94 °C for 30 s; annealing at 68 °C for 30 s; and extension at 72 °C for 30 s for 35 cycles.
The paternal and maternal DNA were examined by fragment analysis in twelve different regions that included STRs as previously described15.
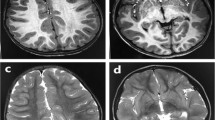
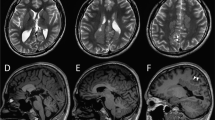
A 3-year-old boy was born to nonconsanguineous parents with no family history of seizures or other neurological disorders. His full-term birth height and weight were 49.5 cm and 3.225 g, respectively, which are within ±2.0 SD. The patient had an uneventful perinatal period. He had transient bilateral dystonic posture in the upper limbs at the age of 5 months. At 9 months, he developed infantile spasms. Hypsarrhythmia and spasms were observed on electroencephalography (EEG) (Fig. 1a, b). Thoracoabdominal X-ray photograph, echocardiogram, fundus examination and biochemical examination showed no abnormalities. Administration of adrenocorticotropic hormone (ACTH) improved the abnormal waveforms on interictal EEG (Fig. 1c) but could not control spasms. After ACTH therapy, vitamin B6, zonisamide, valproic acid, clonazepam, topiramate, vigabatrin, and lamotrigine were administered but were not effective against his spasms. Furthermore, a corpus callosotomy at the age of 21 months had no effect on his spasms. PVNH was not observed on brain MRI either before or after corpus callosotomy (Fig. 1d–k). The patient had a developmental delay (DQ = 28) and could not speak any meaningful words. He also exhibited autistic behavior. Skeletal abnormalities were not observed.
a Electroencephalography (EEG) results of the patient showing hypsarrhythmia during wakefulness at 9 months of age. b Ictal EEG results showing cluster spasms with head drop at 9 months of age. c EEG results at 11 months of age and after adrenocorticotropic hormone therapy. Sporadic slow waves were observed in the bitemporal regions. d, e Brain MRI T1-weighted and f, g T2-weighted images of the patient at the age of 15 months. Three pediatric neurologists independently confirmed the absence of periventricular nodular heterotopia. h, i Brain MRI T1-weighted and j, k T2-weighted images of the patient at the age of 29 months. l Filamin A protein with functional domains and FLNA variants found in males. Functional domains consist of the actin-binding domain containing two calponin homology (CH) domains and 24 Ig domains. Pathogenic variants detected in males are shown as triangles below Filamin A. FLNA variants in PVNH1 or in epilepsy without PVNH are shown as black or red triangles. Filled or open triangles indicate nonsense or missense/in-flame changes.
WES showed a rare hemizygous missense variant in FLNA (NM_001456.4: c.4804G>A: p.Gly1602Ser) (Fig. S1a, b). Sanger sequencing of the patient’s parents confirmed that the variant was de novo. Additionally, short tandem repeat analysis indicated that they were his biological parents.
This variant was not registered in the in-house database, HGVD, ExAC, ToMMo or gnomAD and was indicated as pathogenic based on the following in silico tools: SIFT: 0 (damaging), Polyphen-2: 0.996 (deleterious) and CADD: 27.0 (deleterious). No other rare variants in epilepsy-related genes registered in OMIM or deleterious CNVs were found in the patient. In addition, amino acid substitutions are highly evolutionarily conserved among different species (Fig. S1c).
On the basis of the ACMG Standards and Guidelines16, we concluded that this variant is likely pathogenic according to the following evidence of pathogenicity: strong: PS2, moderate: PM2, supporting: PP3.
Among the clinical consequences of FLNA variants, PVNH1 is the most common brain abnormality6. However, we could not detect PVNH or other brain MRI abnormalities in this patient who developed infantile spasms. We found at least three cases of epilepsy arising from possibly pathogenic FLNA variants that provided no description of PVNH or other abnormal MRI findings (but with no images presented) in the literature17,18,19 (Table 1). One reported male case had a p.Asp1527Asn variant within the Ig domain18 and our case variant, p.Gly1602Ser, was also located within the neighboring Ig domain (Fig. 1l). Two independent male cases could support that FLNA variants can cause epileptic encephalopathy with no PVNH. Interestingly, seizures associated with PVNH1 patients (females and males) are typically adolescent-onset, and early infantile onset is uncommon6,7,10,11,20. Our patient started spasms at the age of 9 months, and other epileptic patients without PVNH had seizures before the age of 1 year (Table 1).
While PVNH1 patients often suffer from seizures, it is unclear whether heterotopia is a direct cause of these seizures. The extent of PVNH on brain MRI is not associated with the age of onset of seizures or overall clinical severity as previously described7. Flna transcripts are highly expressed across the entire cerebral cortex in the late period of mouse embryogenesis (E14.5–E16.5), while filamin B, a homolog of filamin A, is localized near the ventricular and subventricular zone21. Interestingly, in the late period of embryogenesis (E14.5) of Flna-null mice, no neuronal accumulation in the ventricular zone or heterotopic neurons was recognized12. Filamin A is known to interact with the HCN1 channel and modulate neuronal excitability in the mature brain via endocytosis of the HCN1 channel22 encoded by HCN1. HCN1 is also expressed in the entire brain (especially in the cerebral cortex, hippocampus and cerebellum) of rats23. HCN1 variants cause early infantile epileptic encephalopathy (EIEE24, #615871). Considering these facts, FLNA abnormalities may cause subventricular zone abnormalities leading to PVNH1 and dysfunction of the entire cerebral cortex.
We report a boy who suffered from West syndrome without PVNH1 arising from a de novo missense variant in FLNA. Considering the wide expression of filamin A protein in the mature brain, FLNA1 variants may be considered one of rare causes of epileptic encephalopathy without PVNH1.
HGV database
The relevant data from this Data Report are hosted at the Human Genome Variation Database at https://doi.org/10.6084/m9.figshare.hgv.2945.
References
Berg, A. T. et al. Revised terminology and concepts for organization of seizures and epilepsies: report of the ILAE Commission on Classification and Terminology, 2005-2009. Epilepsia 51, 676–685 (2010).
Wang, J. et al. Epilepsy-associated genes. Seizure 44, 11–20 (2017).
Fox, J. W. et al. Mutations in filamin 1 prevent migration of cerebral cortical neurons in human periventricular heterotopia. Neuron 21, 1315–1325 (1998).
Rosa, J. P., Raslova, H. & Bryckaert, M. Filamin A: key actor in platelet biology. Blood 134, 1279–1288 (2019).
Nakamura, F., Stossel, T. P. & Hartwig, J. H. The filamins: organizers of cell structure and function. Cell Adh Migr. 5, 160–169 (2011).
Wade, E. M., Halliday, B. J., Jenkins, Z. A., O’Neill, A. C. & Robertson, S. P. The X-linked filaminopathies: synergistic insights from clinical and molecular analysis. Hum. Mutat. 41, 865–883 (2020).
Lange, M. et al. 47 patients with FLNA associated periventricular nodular heterotopia. Orphanet J. Rare Dis. 10, 134 (2015).
Ekşioğlu, Y. Z. et al. Periventricular heterotopia: an X-linked dominant epilepsy locus causing aberrant cerebral cortical development. Neuron 16, 77–87 (1996).
Moro, F. et al. Familial periventricular heterotopia: missense and distal truncating mutations of the FLN1 gene. Neurology 58, 916–921 (2002).
Reinstein, E. et al. Vascular and connective tissue anomalies associated with X-linked periventricular heterotopia due to mutations in Filamin A. Eur. J. Hum. Genet 21, 494–502 (2013).
Cannaerts, E. et al. FLNA mutations in surviving males presenting with connective tissue findings: two new case reports and review of the literature. BMC Med. Genet. 19, 140 (2018).
Feng, Y. et al. Filamin A (FLNA) is required for cell-cell contact in vascular development and cardiac morphogenesis. Proc. Natl Acad. Sci. USA 103, 19836–19841 (2006).
Nakashima, M. et al. Identification of de novo CSNK2A1 and CSNK2B variants in cases of global developmental delay with seizures. J. Hum. Genet. 64, 313–322 (2019).
Tsuchida, N. et al. Detection of copy number variations in epilepsy using exome data. Clin. Genet. 93, 577–587 (2018).
Okubo, M. et al. GGC Repeat expansion of NOTCH2NLC in adult patients with leukoencephalopathy. Ann. Neurol. 86, 962–968 (2019).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–424 (2015).
Allen, A. S. et al. De novo mutations in epileptic encephalopathies. Nature 501, 217–221 (2013).
Wei, C. M., Xia, G. Z. & Ren, R. N. [Gene mutations in unexplained infantile epileptic encephalopathy: an analysis of 47 cases]. Zhongguo Dang Dai Er Ke Za Zhi Chin. J. Contemp. Pediatrics 20, 125–129 (2018).
DiFrancesco, J. C. et al. HCN ion channels and accessory proteins in epilepsy: genetic analysis of a large cohort of patients and review of the literature. Epilepsy Res. 153, 49–58 (2019).
Guerrini, R. et al. Germline and mosaic mutations of FLN1 in men with periventricular heterotopia. Neurology 63, 51–56 (2004).
Sheen, V. L. et al. Filamin A and Filamin B are co-expressed within neurons during periods of neuronal migration and can physically interact. Hum. Mol. Genet. 11, 2845–2854 (2002).
Noam, Y. et al. Filamin A promotes dynamin-dependent internalization of hyperpolarization-activated cyclic nucleotide-gated type 1 (HCN1) channels and restricts Ih in hippocampal neurons. J. Biol. Chem. 289, 5889–5903 (2014).
Monteggia, L. M., Eisch, A. J., Tang, M. D., Kaczmarek, L. K. & Nestler, E. J. Cloning and localization of the hyperpolarization-activated cyclic nucleotide-gated channel family in rat brain. Brain Res. Mol. Brain Res. 81, 129–139 (2000).
Acknowledgements
This work was supported by AMED under grant numbers JP20ek0109486, JP20dm0107090, JP20ek0109301, JP20ek0109348, and JP20kk0205012 (N. Matsumoto); JSPS KAKENHI grant numbers JP17H01539 (N. Matsumoto), JP19H03621 (N. Miyake), JP20K07907 (S. Miyatake), JP20K08164 (T. Mizuguchi) and JP19K17307 (Y. Azuma); and intramural grants 30-6 and 30-7 from the Ministry of Health, Labour and Welfare (N. Matsumoto) and the Takeda Science Foundation (T. Mizuguchi, N. Miyake, and N. Matsumoto).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hiromoto, Y., Azuma, Y., Suzuki, Y. et al. Hemizygous FLNA variant in West syndrome without periventricular nodular heterotopia. Hum Genome Var 7, 43 (2020). https://doi.org/10.1038/s41439-020-00131-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41439-020-00131-9