Abstract
We had previously reported the case of a male patient with schizophrenia, having de-novo balanced translocation. Here, we determined the exact breakpoints in chromosomes 4 and 13. The breakpoint within chromosome 4 was mapped to a region 32.6 kbp upstream of the LDB2 gene encoding Lim domain binding 2. Variant screening in LDB2 revealed a rare novel missense variant in patients with psychiatric disorder.
Similar content being viewed by others
Schizophrenia is a chronic and disabling brain disorder that affects approximately 1% of the population. Although the disease mechanism is still unknown, its genetic predisposition is clearly evidenced. Genome-wide association studies have identified over 100 independent loci defined by common single-nucleotide variants (SNVs)1. A number of rare variants have been identified till date, with far larger effects on individual risk; de-novo mutations have also been reported to confer substantial individual risk2,3. Increasing evidence has suggested an overlap of genetic susceptibility between schizophrenia and bipolar disorder4,5. Most notable association has been found with the Disrupted In Schizophrenia 1 (DISC1) gene, based upon chromosomal abnormality with a balanced chromosomal translocation (1;11)(q42;q14.3) in a large pedigree6,7.
We had previously reported a male patient with schizophrenia, carrying a de novo balanced translocation t(4;13)(p16.1; q21.31)8. However, the exact breakpoint had not been determined till date. Here, we report the exact breakpoints on chromosomes 4 and 13 using next-generation DNA-sequencing analysis, in combination with fluorescence in situ hybridization (FISH) experiments on the patient. The estimated breakpoints were confirmed by FISH using the BAC clone (RP11-141E13), which was selected from the UCSC genome browser (GRCh38/hg38) (Fig. 1b). According to the database, the breakpoint on chromosome (chr) 13 was within the so-called ‘gene desert’ interval, where no known gene has yet been mapped. The breakpoint on chr 4 was mapped to the upstream region of a gene encoding a putative transcription regulator lacking a DNA-binding domain, namely LDB2 (LIM domain-binding 2, also known as CLIM1) (Supplemental Fig. 1).
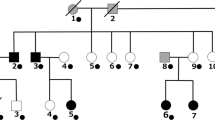
a Idiogram of the translocation karyotype. b FISH analysis, using the lymphoblastoid cell lines (LCL) from the proband. Metaphase spread of LCLs showing BAC (RP11-141E13) hybridization signal to chromosome 4, derivative 4, and derivative 13. Loss of green signal (a white arrow) indicated deletion of this region. c, d Determination of breakpoint sequences of der (4) and der (13).
To determine the exact breakpoint, we conducted whole-genome sequencing using peripheral blood-derived DNA from the proband and the HiSeq 2500 (Illumina, CA, USA) as per the manufacturer’s recommended protocol. Confirmation of the translocation breakpoint and flanking sequence, and resequencing of LDB2 were conducted by Sanger sequencing. Primer details are listed in Supplementary Table S2.
Fine mapping of breakpoints on chr 4 revealed chr 4:16,933,034 and chr13:55,324,705, which was 32.6 kbp upstream of LDB2 (Fig. 1c). The breakpoint on chr 13 was located at chr 4:16,933,035 and chr13:55,324,711 (Fig. 1d). There was no nucleotide deletion or duplication at the breakpoint of chr 4. However, 5 base pairs (ttaaa) were lost from the chr13 breakpoint.
We further performed resequencing of LDB2 to detect a rare variant (MAF < 0.005) having a major alteration in function of the gene. The subjects comprised of 520 unrelated Japanese patients with schizophrenia (SZ) (281 males and 239 females; mean age ± SD, 49.95 ± 12.00 years and 52.54 ± 13.29 years) and 423 with bipolar disorder (BP) (210 males and 213 females; mean age ± SD, 51.19 ± 13.36 years and 49.61 ± 13.94 years), diagnosed according to the Diagnostic and Statistical Manual of Mental Disorders, Fourth Edition (DSM-IV) with consensus from at least 2 experienced psychiatrists. Four hundred control subjects (183 males and 217 females; mean age ± SD, 40.48 ± 12.18 years and 40.63 ± 12.64 years) were included, whose second-degree relatives were free of psychosis, as reported by the subjects. All the participants provided written informed consent. This study was approved by the Ethics Committees of the Tokyo Metropolitan Institute of Medical Science and RIKEN. By resequencing of the coding region of LDB2, six rare variants (four synonymous and two nonsynonymous) were detected (Table 1). Of these, two variants (p.Thr83Asn and p.Ala287Ala) have not been reported in any database, including dbSNP (https://www.ncbi.nlm.nih.gov/snp/), gnomAD (https://gnomad.broadinstitute.org/), and jMorp (https://jmorp.megabank.tohoku.ac.jp/202001/variants) (Table 1). The two nonsynonymous variants (p.Thr83Asn and p.Pro170Leu) that were only found in BP were classified as probably damaging by PolyPhen-29.
Previous reports had suggested 4p16.1 region to be associated with schizophrenia and bipolar disorder10,11,12; however, there was no association between SZ, BP, and control, in this study (Supplemental Table 1). Recent studies indicated that clinical significance of balanced chromosomal abnormalities was due to disruption of the topologically associated domains (TADs)13. Chromosomal breakpoint was located on the same TAD region as LDB214, (Ohnishi et al. submitted), hence implying alteration of the gene expression of LDB2. Unfortunately, we could not collect RNA sample from the proband, due to which, we could not confirm the expression level of LBD2. Information regarding the function of LDB2 protein is limited, and several reports have shown LIM-domain proteins to regulate cell proliferation and cell fate in many regions of the CNS15,16. In support of our observation, study on the Ldb2 KO mouse had suggested Ldb2 deficiency to result in various behavioral and functional impairments relevant to mental disorders (Ohnishi et al. submitted).
In conclusion, we identified the breakpoint of balanced translocation t(4;13)(p16.1; q21.31), and proposed the LDB2 gene to possibly be linked to psychiatric disorder; however, the correlation between phenotype and genotype regarding this disorder would require further studies.
HGV database
The relevant data from this Data Report are hosted at the Human Genome Variation Database at https://doi.org/10.6084/m9.figshare.hgv.2909; https://doi.org/10.6084/m9.figshare.hgv.2912.
References
Schizophrenia Working Group of the Psychiatric Genomics C. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
MacIntyre, D. J., Blackwood, D. H., Porteous, D. J., Pickard, B. S. & Muir, W. J. Chromosomal abnormalities and mental illness. Mol. Psychiatry 8, 275–287 (2003).
Singh, T. et al. The contribution of rare variants to risk of schizophrenia in individuals with and without intellectual disability. Nat. Genet. 49, 1167–1173 (2017).
Bipolar, D., Schizophrenia Working Group of the Psychiatric Genomics Consortium. Electronic address drve, Bipolar D, Schizophrenia Working Group of the Psychiatric Genomics C. Genomic dissection of bipolar disorder and schizophrenia, including 28 subphenotypes. Cell 173, 1705–1715.e16 (2018).
Andlauer T. F. M. et al. Bipolar multiplex families have an increased burden of common risk variants for psychiatric disorders. Mol. Psychiatry. https://doi.org/10.1038/s41380-019-0558-2 (2019).
Millar, J. K. et al. Disruption of two novel genes by a translocation co-segregating with schizophrenia. Hum. Mol. Genet. 9, 1415–1423 (2000).
Blackwood, D. H. et al. Schizophrenia and affective disorders-cosegregation with a translocation at chromosome 1q42 that directly disrupts brain-expressed genes: clinical and P300 findings in a family. Am. J. Hum. Genet. 69, 428–433 (2001).
Itokawa, M., Kasuga, T., Yoshikawa, T. & Matsushita, M. Identification of a male schizophrenic patient carrying a de novo balanced translocation, t(4; 13)(p16.1; q21.31). Psychiatry Clin. Neurosci. 58, 333–337 (2004).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Als, T. D. et al. Possible evidence for a common risk locus for bipolar affective disorder and schizophrenia on chromosome 4p16 in patients from the Faroe Islands. Mol. Psychiatry 9, 93–98 (2004).
Suarez, B. K. et al. Genomewide linkage scan of 409 European-ancestry and African American families with schizophrenia: suggestive evidence of linkage at 8p23.3-p21.2 and 11p13.1-q14.1 in the combined sample. Am. J. Hum. Genet. 78, 315–33 (2006).
Xu, W. et al. Genome-wide association study of bipolar disorder in Canadian and UK populations corroborates disease loci including SYNE1 and CSMD1. BMC Med. Genet. 15, 2 (2014).
Redin, C. et al. The genomic landscape of balanced cytogenetic abnormalities associated with human congenital anomalies. Nat. Genet. 49, 36–45 (2017).
Wang, D. et al. Comprehensive functional genomic resource and integrative model for the human brain. Science 362, eaat8464 (2018).
Matthews, J. M. et al. Competition between LIM-binding domains. Biochemical Soc. Trans. 36, 1393–1397 (2008).
Gueta, K. et al. The stage-dependent roles of Ldb1 and functional redundancy with Ldb2 in mammalian retinogenesis. Development 143, 4182–4192 (2016).
Acknowledgements
We would like to express our sincere gratitude to the patient and his family for their participation in this study. We are grateful to Mr. Naohiko Maeda for technical support. This work was supported by JSPS KAKENHI [grant numbers 19H03589, 22129007, 20023039, 20249054, 18390323 (to M.I.), 20K07934, 16K07017, 24591737 (to T.O)], and SENSHIN Medical Research Foundation (to M.A. and T.I.). We thank the Support Unit for Bio-Material Analysis at RIKEN CBS for NGS.
Funding
This work was supported by JSPS KAKENHI [grant numbers 19H03589, 22129007, 20023039, 20249054, 18390323 (to M.I.), 20K07934, 16K07017, 24591737 (to T.O.)], and SENSHIN Medical Research Foundation (to M.A. and T.I.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Horiuchi, Y., Ichikawa, T., Ohnishi, T. et al. LDB2 locus disruption on 4p16.1 as a risk factor for schizophrenia and bipolar disorder. Hum Genome Var 7, 31 (2020). https://doi.org/10.1038/s41439-020-00117-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41439-020-00117-7