Abstract
Fire blight disease, caused by the bacterium Erwinia amylovora (E. amylovora), is responsible for substantial losses in cultivated apples worldwide. An important mechanism of plant immunity is based on the recognition of conserved microbial molecules, named pathogen-associated or microbe-associated molecular patterns (PAMPs or MAMPs), through pattern recognition receptors (PRRs), leading to pattern-triggered immunity (PTI). The interspecies transfer of PRRs represents a promising strategy to engineer broad-spectrum and durable disease resistance in crops. EFR, the Arabidopsis thaliana PRR for the PAMP elf18 derived from the elongation factor thermal unstable (EF-Tu) proved to be effective in improving bacterial resistance when expressed into Solanaceae and other plant species. In this study, we tested whether EFR can affect the interaction of apple with E. amylovora by its ectopic expression in the susceptible apple rootstock M.26. Stable EFR expression led to the activation of PAMP-triggered immune response in apple leaves upon treatment with supernatant of E. amylovora, as measured by the production of reactive oxygen species and the induction of known defense genes. The amount of tissue necrosis associated with E. amylovora infection was significantly reduced in the EFR transgenic rootstock compared to the wild-type. Our results show that the expression of EFR in apple rootstock may be a valuable biotechnology strategy to improve the resistance of apple to fire blight.
Similar content being viewed by others
Introduction
Despite being constantly exposed to a wide range of pathogens in their immediate environment, plants are resistant to most microbes. Each plant cell can trigger an immune response autonomously by employing pattern recognition receptors (PRRs) for sensitive and rapid detection of potential threats caused by pests or pathogens1. This mechanism is based on the recognition of conserved microbial molecules (pathogen-associated or microbial-associated molecular patterns, PAMPs or MAMPs) and is known as pattern-triggered immunity (PTI). PAMPs are important for microbes (both pathogenic and non-pathogenic) and are widely conserved across microorganisms. When these molecules are recognized by plants, a series of defense reactions are induced such as reactive oxygen species (ROS) production, callose deposition, expression of defense genes, and activation of mitogen-activated protein kinase (MAPK) cascades2,3. Several PAMPs have been described and their recognition investigated in plants3,4,5 such as beta-glucan6, chitin7, flagellin8, and the elongation factor thermal unstable (EF-Tu)9.
EF-Tu is one of the most abundant bacterial proteins and acts in Arabidopsis thaliana (hereafter, Arabidopsis) as a very potent PAMP9. Arabidopsis recognizes the N-terminus of the protein and specifically the N-acetylated peptide comprising the first 18 amino acids of EF-Tu, called elf18. EF-Tu is recognized by EF-TU RECEPTOR (EFR), a Brassicaceae-specific10. EFR is related to FLAGELLIN SENSING 2 (FLS2), the receptor for the flagellin-derived PAMP flg22, and belongs to the leucine-rich repeat receptor kinase (LRR-RK) XII sub-family. Elf18 perception by EFR is specific to the Brassicaceae family. However, expression of the Arabidopsis EFR gene in Nicotiana benthamiana and in tomato conferred elf18 responsiveness and increased anti-bacterial resistance to these species, indicating that the downstream elements of PRR activation are conserved between Arabidopsis and other plants11,12. Moreover, expression of EFR has been also reported to increase resistance to bacterial diseases in other plants such as Medicago truncatula13, potato14, rice2,15, and wheat16.
Fire blight, caused by the Gram-negative bacterium Erwinia amylovora (E. amylovora), is a destructive disease that strongly affects the Rosaceae family, including the economically important species apple and pear17. E. amylovora is classified as a member of the Enterobacteriaceae family, and, as such, is closely related to many important human and animal pathogens, such as Escherichia coli, Salmonella spp., Shigella spp. and Yersinia spp.
Control of the fire blight pathogen is difficult and is typically achieved through the eradication of entire trees displaying symptoms or via preventive spraying of copper compounds or antibiotics. However, these methods are not completely effective when used in isolation17, and the application of antibiotics is not authorized in some countries due to the evolution of antibiotic resistance in populations of plant pathogens18. Within this framework, classical breeding and genetic engineering have been widely applied for the production of resistant cultivars and for targeted bacterial disease management19,20,21,22,23,24,25,26,27,28,29. However, on the one hand, classical breeding is time-consuming and the quality of the obtained new cultivars is often inferior to that of commercially available apples, thus hindering consumer acceptance. On the other hand, the use of genetically modified plants is still forbidden in many countries worldwide. Due to its destructive character and to the lack of effective control mechanisms, E. amylovora is capable of dispersing rapidly within susceptible plants and between trees in orchards, which could result in great economic losses.
In this study, we generated transgenic apple plants expressing the AtEFR gene under the control of the AtFLS2 promoter in the susceptible apple rootstock M.26. Transgenic plants were characterized by molecular and phenotypic analyses to evaluate EFR functionality and its ability to confer increased resistance to E. amylovora.
Results
Evaluation of E. amylovora EF-Tu eliciting activity
Based on the EF-Tu sequence from Escherichia coli (E. coli)9, the sequence of elf18 was retrieved from different Erwinia species (Fig. 1A, B). Alignment of the Erwina sequences with 14 selected elf18 sequences revealed that all of the elf18 sequences derived from Erwinia species are grouping with the E. coli elf18 sequence (Fig. 1B). The strong conservation of the consensus in the elf18 peptides can be observed (Fig. 1A). Indeed, 9 of the 18 amino acids of elf18 conserved domains are preserved in all the bacteria species (Fig. 1A), while specific amino acid substitution allows defining specific clades of bacteria (Fig. 1B).
A Alignment of elf18 regions from selected bacteria: XANAC: Xanthomonas Axonopodis pv. citrus, XANOM: Xanthomonas oryzea pv. oryzea; XANC8: Xanthomonas Campestris pv. Campestris 8004; XANCB: Xanthomonas Campestris pv. Campestris B100; AGRT5: Agrobacterium tumefaciens C58; RHIRD: Agrobacterium tumefaciens; AGRVS: Agrobacterium vitis S4; EaB661: Erwinia billingeae 661; EaT1/99: Erwinia tosmaniensis 1/99; Ea1430: Erwinia amylovora 1430; Ea273: Erwinia amylovora 273; EP1/79: Erwinia pyrifoliae Ep1/79; ERWCT: Erwinia carotovora ssp. Atroseptical/P. Atrosepticum; EcK12: Echerichia coli K12; RALSO: Ralsonia solanacearum GMI1000; PSE14: Pseudomonas syringae pv. Phaseolicola 1448; PSESM: Pseudomonas syringae pv. tomato DC3000; PSEU2: Pseudomonas syringae pv. syringae B728a; PSEPF: Pseudomonas fluorescence pfo-1. B Phylogenetic tree of the elf18 sequences from the selected bacteria mentioned above. C Reactive oxygen species generation in Arabidopsis thaliana wild-type and efr mutant in response to mock or Erwinia amylovora (Ea1430 or Ea273) supernatant treatments measured in relative light units (RLU). Data are average ± SE (n = 10). This experiment was repeated 2 times with similar results
To verify the activity of E. amylovora EF-Tu elicitor, heat-killed E. amylovora extracts were applied to wild-type and efr mutant A. thaliana leaf discs. Extracts from E. amylovora strains were able to elicit ROS production in wild-type Arabidopsis leaves; in contrast, this response was abolished in efr mutant leaves (Fig. 1C). These results indicate that the main elicitor present in the extract derives from EF-Tu.
Generation of transgenic apple, molecular evaluation of the transgenic events, and ploidy level
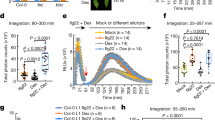
The susceptible apple rootstock M.26 was transformed to express EFR (see “Methods” section for details). A total of 6 transgenic lines were obtained from one transformation experiment. All the transgenic lines were confirmed by PCR for the presence of the EFR gene and then characterized for the T-DNA integration copy number into the genome. The lines TM-2017, TM2108, and TM2114 had one T-DNA insertion, while the lines TM2111, TM2113, and TM2112 had 3, 5, and 8 copies of inserted T-DNA, respectively. Moreover, the ploidy level determined by flow cytometry showed that all of the transformed lines were diploid, like the non-transformed M.26. Thus, the expression of EFR was evaluated in plants grown at greenhouse conditions. RT-qPCR analysis showed that EFR was expressed in leaves of all the transgenic lines before E. amylovora infection and significantly increased 10-fold to 20-fold 24 h post-infiltration with E. amylovora bacterial suspension or supernatant, respectively (Fig. 2).
This experiment was repeated 3 times with similar results. Bars indicate the mean values ± SE. Considering mock-treatments and pathogen-treatments separately, asterisks indicate statistically significant differences of datasets from the corresponding dataset at time zero (0 h), according to one-way ANOVA followed by post hoc Dunnett’s test (α = 0.05)
EFR expression in apple induces oxidative burst and defense gene upon Erwinia inoculation
To test the effect of the expression of EFR on the production of ROS in the transgenic lines (untreated and treated with E. amylovora or supernatant of E. amylovora), the production of superoxide anions was examined using NBT staining in those lines showing only one insertion copy. As shown in Fig. 3A, 48 h after infiltration with bacteria or supernatant of E. amylovora (strain Ea273), a significant accumulation of superoxide anions can be observed in the two M.26 transgenic lines (TM-2107 and TM-2108) tested. In the control M26, a small superoxide anion staining can be noticed after treatment with E. amylovara supernatant compared with infection with E amylovora. In the M.26 EFR transgenic lines, a low superoxide anion staining can be observed after water treatment, demonstrating that ROS burst is effectively due to the induction of EFR after treatments.
The histogram represent the merge data of 3 biological experiments. Bars indicate the mean values ± SE. Considering mock- and pathogen-treatments separately, asterisks indicate statistically significant differences of datasets from the corresponding dataset at time zero (T0), according to one-way ANOVA followed by post hoc Dunnett’s test (α = 0.05)
PAMPs elicit the rapid and strong expression of MPK3, MPK6, and other defense genes in Arabidopsis30. This observation led us to examine whether ROS marker genes were induced in our transgenic lines after bacterial infection. Transcript levels of MdMPK3-2 and MdWRKY25 increased in the transgenic lines after infiltration with bacteria or supernatant of E. amylovora (strain Ea273), whereas a small increase can be observed in M.26 only after inoculation with the bacteria suspension of E. amylovora (Fig. 3B). In contrast, no significant increase of expression can be observed for the MdNPR1 gene in all the tested samples.
Evaluation of transgenic lines on their resistance to fire blight
To characterize the response of the transformed lines to fire blight, 20 acclimated biological replicates per line were inoculated with E. amylovora Ea273 at 5 × 107 cfu mL–1 in three independent experiments. The response was similar in the replicated experiments, as indicated by low confidence intervals of means. The control M.26 showed average necrosis of 70%, indicating its high susceptibility to E. amylovora, while all the EFR transgenic lines showed significantly lower necrosis (Fig. 4A, B) compared to the wild-type plants. Indeed, all EFR transgenic lines exhibited a reduction in the necrosis of 70% to 80% compared to the control, one month post-inoculation with E. amylovora strain Ea273. Notably, this level of resistance was comparable to what was observed in the resistant genotype M.7 (Malnoy et al. 2007). In the transgenic lines, the necrosis due to E. amylovora was restricted in an area of few centimeters from the inoculation point, allowing the growth of new healthy shoots under the necrotic tissue (Fig. 4A). For the control, almost all the shoots were necrotic 15 days after inoculation, without new growing shoots. Moreover, the reduction of necrosis in the transgenic lines (T-2107, T-2108, and T2112) 5 days after infection correlated with a 100-fold reduction in the amount of E. amylovora that could be recovered from the transgenic plants compared to control plants. Indeed, 120 h after inoculation the bacterial population, evaluated by a population dynamic study, had reached 5 × 108 cfu g–1 (+/−0,5 cfu g–1) in the inoculated wild type leaves and 105 cfu g–1 (+/−0,7 cfu g–1) in the inoculated transgenic leaves. Furthermore, no E. amylovora bacteria was detected in the healthy transgenic tissues compared to the control. The line T-2107, T-2108, and T2112 were future inoculated with the virulent strain of E.a 3050, which contour the resistance gene of Malus Robusta 5) and the same results of reduction in the necrosis were observed as for the strain E.a 273 (data not shown).
A Percentage of necrosis (shown with 95% confidence interval bar). Susceptibility of the EFR transgenic rootstock lines to E. amylovora in comparison with the wild-type M.7 apple rootstock (resistant to fire blight) and M.26 (susceptible to fire blight) apple rootstock. Lengths of the shoot and the portions that appeared necrotic were measured 1 month after inoculation with E. amylovora. Transgenic rootstock susceptibility is significantly different from M.26 at P < 0,001 (three asterisks), according to Kruskall and Wallis test. B Pictures, taken 1 month after inoculation, showing the fire blight-induced necrotic phenotype in the wild-type M.26 and in selected transgenic lines. Data are average ± SD (n = 20). This experiment was repeated 3times with similar results
Discussion
Apple is one of the major fruit crops in the world with over 80 million tons produced each year (FAO, 2017). To maintain a high and good level of production, a considerable amount of chemical treatments is necessary (15–30 treatments/year) to protect apples against their wide range of pathogens. Major management strategies for bacterial diseases in Malus include biological control, cultivation practices, and breeding for disease-resistant cultivars. Although chemical control has proved to be very successful for pests and fungal diseases in apples, in the case of fire blight, no truly effective bactericide is commercially available17. Therefore, breeding for disease resistance has been considered the most effective and environmentally friendly approach to control bacterial diseases in apples. However, valuable resistant germplasms within domesticated Malus are limited for bacterial diseases such as fire blight31,32. Transfer of resistance genes is an alternative way to improve plant immunity and that has been already done in apple and pear to increase resistance to fire blight12,19,33,34. In the case of the resistance conveyed by the Rin4 gene, it has been overcome by the pathogen over a period of time35.In order to develop novel forms of disease control, understanding the key determinants of the relevant bacterium–host interaction is crucial18. One of these key determinants is the EF-Tu protein. Sequence alignment revealed that the key residues in the elf18 epitope of E. amylovora are conserved, thus suggesting that EF-Tu-derived from E. amylovora is recognized by EFR (Fig. 1). In addition, we showed that supernatant of two strains of E. amylovora can induce ROS production in A. thaliana but not in the efr mutant.
In this study, we generated a transgenic rootstock of apples that express AtEFR to confer recognition of EF-Tu from E. amylovora. The transgenic rootstock M.26 plants recognize the EF-Tu-derived peptide elf18, leading to the production of ROS burst in leaves as previously reported in other plants expressing EFR14. These results indicate that components required for EFR-triggered downstream immune signaling are conserved not only in Solanaceae but also in apple, the 50-million year divergence between Maloideae species and Arabidopsis thaliana.
We demonstrated that all the generated apple transgenic lines present a high level of resistance to fire blight similar to the resistant control (M.7)33. Furthermore, transgenic rootstock plants expressing EFR did show constitutive activation of defense responses, as measured by ROS production, and expression of defense genes. They also did not exhibit any developmental or growth defects over several generations (data not shown). After treatment with supernatant of E. amylovora or inoculation with E. amylovora, a strong ROS production and increase of defense gene expression could however be observed, documenting the inducibility of EFR activation in the transgenic lines.
The use of this transgenic rootstock can be interesting for protecting apple cultivars against fire blight or other bacterial pathogens such as Agrobacterium tumefaciens. Indeed, rootstocks have been used in agriculture for over 2000 years in Asia to improve production, reduce disease susceptibility, and increase sustainable agriculture by reducing inputs36. The first contribution of rootstock for food cultivation was by domestication and propagation of woody species difficult to root from cuttings such as apple, pear, and plum37. Nevertheless, the best example of rootstock contribution to food security was the rescue of the European grape and wine industry from the devastating effects of the soil-born insect phyloxera in the 19th century38.
Our transgenic rootstock can be used as resistant rootstock against fire blight in countries that already allow the cultivation of GM organisms, and might be a further motivation for the deregulation of GM crops to improve disease resistance in countries that do not do so yet. It may also improve the resistance of the scion to the disease; indeed, it was proven that grafting can be useful for the transport of signaling molecules from the rootstock to the scion to gain virus resistance39,40.
Materials and methods
Evaluation of elf18 peptides
Nineteen elf18 peptide sequences were retrieved from Lacombe et al.11 and from the E. amylovora genome. Phylogenetic trees (maximum likelihood model) for these sequences were constructed using MEGA7 software41 after sequence alignment via clustalW42.
Plant materials and growth conditions
For apple, wild-type and transgenic plantlets of the rootstock M.26 were grown in vitro following the procedure described by Malnoy et al.33 and acclimated to the soil according to Bolar et al.43. In vitro and glasshouse plant material was cultivated with a 16/8-h light/dark period at 20–24 °C.
Arabidopsis thaliana plantlets of ecotype Columbia-0 (Col-0) and efr mutant lines were grown in soil as one plant per pot or in plates containing half MS salts medium, 1% sucrose, and 1% plant agar, at 20–22 °C with a 16/8-h light/dark period.
Plant transformation and evaluation of ploidy level
The binary vector pBin19 used for plant transformation carries the AtEFR open reading frame flanked by the Arabidopsis FLS2 promoter and the octopine synthase terminator (pBin19/FLS2p:EFR-FLAG). This vector was transferred into Agrobacterium tumefaciens strain EHA105 by electroporation and used to transform the apple rootstock M.26.
Plant transformation, regeneration, propagation, and acclimatization were carried out as described by Borejsza-Wysocka et al.44. Regenerated transgenic plants were confirmed by PCR with primers efrF (5′-CTTGAATTTATTGGGGCTGTGGCG-3′) and efrR (5′-CCTGCAAGTTCAAAAGCTTCCCGA-3′) for the presence of the transgene. Following transformation, the ploidy level of the transgenic and untransformed clones was also estimated by flow cytometry as described by Malnoy et al.34.
Preparation of the bacterial supernatant
Overnight culture of E. amylovora (strains Ea273 and Ea1430) was centrifuged at 4000×g for 5 min. The obtained pellet was suspended in 10 mM phosphate buffer (pH 7,8) and boiled at 100 °C for 15 min. The bacterial suspension was then centrifuged at 4000 × g for 5 min and the supernatant was collected for further experiments.
Reactive oxygen species measurement
Leaves of Arabidopsis thaliana Col-0 plants or efr mutants were treated with the supernatant of two strains of E. amylovora (strains Ea273 and Ea1430). ROS measurement was performed as previously described by Zipfel et al.10.
The detection of superoxide anions in apple leaves was performed by staining the leaves with nitroblue tetrazolium (NBT). The youngest leaves of actively growing plants (control and transgenic lines) were transferred to an NBT staining solution containing 5 mM NBT in 20 mM phosphate buffer, pH 6,545,46. The leaves were incubated in staining solution under a constant vacuum until dark color appeared on the surfaces of the stained leaves. Stained leaves were washed with 100% ethanol for better visualization of the staining.
Quantification of nptII copy number by Taqman real-time PCR
The quantification of the nptII copy number was used to calculate the number of T-DNA insertion events in the transformed apple plants (Supplemental Data 1) according to the Taqman real-time PCR method developed by Dalla Costa et al.47. Primers and probes used in the analysis are listed in Table 1.
Determination of fire blight resistance
The evaluation for fire blight symptoms was carried out in three independent experiments, as described by Malnoy et al.33. In brief, 20 vigorously growing shoots of M.26 transgenic plants and untransformed control plants (M.26 susceptible and M7 resistant cultivars) were inoculated by cutting the two youngest expanded leaves with scissors previously dipped in an E. amylovora suspension (strain Ea273, 5 107 cfu mL−1). Only actively growing plants that showed a shoot length of at least 13.0 cm were considered for the experiments. Disease severity was rated 1 month after inoculation as a percentage of the length of the necrosis/total length of the shoot growth, as described by Campa et al.48. Data interpretation and statistical analysis were performed according to Pompili et al.49.
Inoculated shoot tips became orange-brown and eventually dark brown, often with the production of cloudy ooze droplets which darkened with time. These symptoms proceeded basipetally on the inoculated shoot and eventually terminated depending on the susceptibility of the shoot resulting from its genetics and the vigor of its growth. Three to fifteen biological replicates for each plant line were inoculated with E. amylovora strain Ea273 or Ea3050 (107 cfu mL−1) in each of the three independent experiments performed. Only actively growing plants that showed a shoot length of at least 13 cm were considered for the experiments. Data collection was performed according to Campa et al.48,50,51,52. Statistical analysis was performed using the DellTM StatisticaTM Software version 13.1. As the three experiments showed the same trend, measures of each plant line were merged and analyzed as a single experiment. Nonparametric Kruskal–Wallis test was used to compare groups as data did not show a normal distribution. Subsequently, all groups were compared simultaneously by multiple comparisons of mean rank. Statistical analysis was performed with probability level (α = 0.05).
Bacterial concentration in plants
Fifteen days post-inoculation, three sections of an inoculated leaf or healthy tissue from EFR transgenic plants and control M.26 were disrupted with two metal balls (5 mm) in 500 mL King B media in a GenoGrinder (2 9 20 s, 1250 spm) to release bacteria from the leaf apoplast. A 10−1–10−6 dilution series was made in LB medium and 100 μL from each dilution were spotted on Kado media and incubated at 28 °C for 24 h. The number of colonies was counted to estimate the number of colony-forming units (cfus) per inoculation site. Four independent experiments were performed.
Total RNA isolation and gene expression analysis
Total RNA was extracted from apple leaves with the Spectrum Plant Total RNA Kit (Sigma-Aldrich, St. Louis, MO). Two micrograms of total RNA were treated with DNAse (Ambion) and then reverse-transcribed with SuperScript VILO cDNA synthesis kit (Invitrogen, Carlsbad, CA), according to manufacturer instructions. Real-time quantitative PCR was conducted using Biorad CFX96 System (Bio-Rad Laboratories, Hercules, CA) as described in Campa et al. (2019). Each reaction was performed in triplicate. Gene-specific primers are reported in Table 1. Apple Actin (actin2), Ubiquitin (Ubi), and Elongation Factor 1 (EF1) were used as internal reference genes for normalization (Table 1). The Biorad CFX manager software version 3.0 was used to analyze the relative expression level. Statistical analysis was performed using Statistica software version 9.1 (StatSoft Inc., 2010). Gene expression variance was analyzed by one-way ANOVA followed by a post hoc Tukey’s test (alpha 0,05). All analyses were performed using three biological replicates.
References
Zipfel, C. Plant pattern recognition receptors. Trends Immunol. 35, 345–351 (2014).
Lu, F. et al. Enhancement of innate immune system in monocot rice by transferring the dicotyledonous elongation factor Tu receptor EFR. J. Integrat. Plant Biol. https://doi.org/10.1111/jipb.12306 (2014).
Wu, Y. & Zhou, J. M. Receptor like kinases in plant innate immunity. J. Integr. Plant Biol. 55, 1271–1286 (2013).
Kawano, Y. & Shimamoto, K. Early signaling network in rice PRR-mediated and R-mediated immunity. Curr. Opin. Plant Biol. 16, 496–504 (2013).
Schwessinger, B. & Ronald, P. C. Plant innate immunity: perception of conserved microbial signatures. Annu. Rev. Plant Biol. 63, 451–482 (2012).
Klarzynski, O. et al. Linear beta-1,3 glucans are elicitors of defense responses in tobacco. Plant Physiol. 124, 1027–1038 (2000).
Kaku, H. et al. Plant cells recognize chitin fragments for defense signaling through a plasma membrane receptor. Proc. Natl Acad. Sci. USA 103, 11086–11091 (2006).
Felix, G., Duran, J. D., Volko, S. & Boller, T. Plants have a sensitive perception system for the most conserved domain of bacterial flagellin. Plant J. 18, 265–276 (1999).
Kunze, G. et al. The N terminus of bacterial elongation factor Tu elicits innate immunity in Arabidopsis plants. Plant Cell. 16, 3496–3507 (2004).
Zipfel, C. et al. Perception of the bacterial PAMP EF-Tu by the receptor EFR restricts Agrobacterium-mediated transformation. Cell 125, 749–760 (2006).
Lacombe, S. et al. Interfamily transfer of a plant pattern recognition receptor confers broad-spectrum bacterial resistance. Nat. Biotechnol. 28, 365–369 (2009).
Malnoy, M., Venisse, J. S., Faize, M., Geider, K. & Chevreau, E. Expression of viral EPS-depolymearse reduces fire blight susceptibility in transgenic pear. Plant Cell Rep. 23, 632–638 (2005).
Pfeilmeier, S. et al. Expression of the Arabidopsis thaliana immune receptor EFR in Medicago truncatula reduces infection by a root pathogenic bacterium, but not nitrogen-fixing rhizobial symbiosis. Plant Biotechnol. J. 17, 569–579 (2019).
Boschi, F. et al. Enhanced bacterial wilt resistance in potato through expression of Arabidopsis EFR and introgression of quantitative resistance from Solanum commersonii. Front Plant Sci. 25, 1642 (2017). 8.
Schwessinger, B. et al. Transgenic expression of the dicotyledonous pattern recognition receptor EFR in rice leads to ligand dependent activation of defense responses. PLoS Pathogen 11, e1004809 (2015).
Schoonbeek, H. J. et al. Arabidopsis EF-TU receptor enhances bacterial disease resistance in transgenic wheat. N. Phytologist 206, 606–613 (2015).
Malnoy, M. et al. Fire Blight: applied genomic insights of the pathogen and host. Annu. Rev. Phytopathol. 8, 475–494 (2012).
Sundin, G. W., Castiblanco, L. F., Yuan, X., Zeng, Q. & Yang, C. H. Bacterial disease management: challenges, experience, innovation and future prospects. Mol. Plant Pathol. 17, 1506–1518 (2016).
Broggini, G. A. et al. Engineering fire blight resistance into the apple cultivar ‘Gala’ using the FB_MR5 CC-NBS-LRR resistance gene of Malus x robusta 5. Plant Biotechnol. J. 12, 728–733 (2014).
Durel, C. E., Denancé, C. & Brisset, M. N. Two distinct major QTL for resistance to fire blight co-localize on linkage group 12 in apple genotypes ‘Evereste’ and Malus floribunda clone 821. Genome 52, 139–147 (2009).
Emeriewen, O. et al. Identification of a major quantitative trait locus for resistance to fire blight in the wild apple species Malus fusca. Mol. Breed. 34, 407–419 (2014).
Emeriewen, O. F., Richter, K., Hanke, M. V., Malnoy, M. & Peil, A. The fire blight resistance QTL of Malus fusca (Mfu10) is affected but not broken down by the highly virulent Canadian Erwinia amylovora strain E2002A. Eur. J. Plant Pathol. 141, 631–635 (2015).
Emeriewen, O. F., Richter, K., Hanke, M. V., Malnoy, M. & Peil, A. Further insights into Malus fusca fire blight resistance. J. Plant Pathol. 99, 45–49 (2017a).
Emeriewen, O. F. et al. Fire blight resistance of Malus ×arnoldiana is controlled by a quantitative trait locus located at the distal end of linkage group 12. Eur. J. Plant Pathol. 148, 1011–1018 (2017b).
Emeriewen, O. F. et al. Towards map-based cloning of FB_Mfu10: identification of a receptor-like kinase candidate gene underlying the Malus fusca fire blight resistance locus on linkage group 10. Mol. Breed. 38, 106 (2018).
Gardiner, S. E. et al. Putative resistance gene markers associated with quantitative trait loci for fire blight resistance in Malus ‘Robusta 5’ accessions. BMC Genet. 13, 1–20 (2012).
Kost, T. D. et al. Development of the first cisgenic apple with increased resistance to fire blight. PLoS ONE 10, e0143980 (2015).
Malnoy, M., Xu, M., Borejsza-Wysocka, E. E., Korban, S. S. & Aldwinckle, H. S. Two receptor-like genes, Vfa1 and Vfa2, confer resistance to the fungal pathogen Venturia inaequalis inciting apple scab disease. Mol. Plant-Microbe Interact. 21, 448–458 (2008).
Malnoy, M., Reynoird, J. P., Borejsza-Wysocka, E. E. & Aldwinckle, H. S. Activation of the pathogen-inducible Gst1 promoter of potato after elicitation by Venturia ineaqualis and Erwinia amylovora in transgenic apple (Malus X domestica). Transgenic Res. 15, 83–93 (2006).
Pitzschke, A., Djamei, A., Bitton, F. & Hirt, H. A major role of the MEKK1-MKK1/2-MPK4 pathway in ROS signaling. Mol. Plant 2, 120–137 (2009).
Volk, G. M. et al. Ex situ conservation of vegetatively propagated species: development of a seed-based core collection for Malus sieversii. J. Am. Soc. Hort. Sci. 130, 203–210 (2005).
Volk, G. M. et al. Genetic diversity and disease resistance of wild Malus orientalis from Turkey and Southern Russia. J. Am. Soc. Hort. Sci. 133, 383–389 (2008).
Malnoy, M., Jin, Q., Borejsza-Wysocka, E., He, S. Y. & Aldwinckle, H. S. Over-expression of the Apple Gene MpNPR1 causes increased disease resistance in Malus X domestica. Mol. Plant-Microbe Interact. 20, 1568–1580 (2007).
Malnoy, M., Venisse, J. S. & Chevreau, E. Expression of a bacterial effector, Harpin NEa, causes increased resistance to fire blight in Pyrus communis. Tree Genet. Genomes 1, 41–49 (2005).
Vogt, I. et al. Gene-for-gene relationship in the host-pathogen system Malus x robusta 5-Erwinia amylovora. N. Phytol. 197, 1262–1275 (2013).
Haroldsen, V. M. et al. Mobility of transgenic nucleic acids and proteins within grafted rootstocks for agricultural improvement. Front. Plant Sci. 3, 39 (2012).
Mudge, K. et al. A history of grafting. Hortic. Rev. 35, 437–493 (2009).
Pouget, R. Histoire de la Lutte contre le Phylloxera de la Vigne en France (Institut National de la Recherche Agronomique, Paris, 1990).
Kasai, A., Bai, S., Li, T. & Harada, T. Graft-transmitted siRNA signal from the root induces visual manifestation of endogenouspost-transcriptional gene silencing in the scion. PLoS ONE 6, e16895 (2011).
Zhao, D. & Song, G. Rootstock-to-scion transfer of transgene derived small interfering RNAs and their effect on virus resistance in non transgenic sweet cherry. Plant Biotechnol. J. 12, 1319–1328 (2014).
Kumar, S. et al. Transgenic expression of EFR and Ds2 genes for field management of bacterial wilt and bacterial spot of tomato. Phytopathology 108, 1042–1411 (2018).
Larkin, M. et al. Clustal W and clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Bolar, J. P., Norelli, J. L., Aldwinckle, H. S. & Hanke, V. An efficient method for rooting and acclimation of micropropagated apple cultivar. HortScience 33, 1251–1252 (1998).
Borejsza-Wysocka, E., Norelli, J. L., Aldwinckle, H. S. & Malnoy, M. Long-tern stability of attacin E expression in transformed apple after 12 years in the field and effect of the expression of this gene in the fruit characteristics. BMC Biotechnol. 10, 41 (2010).
Müller, K. et al. In vivo cell wall loosening by hydroxyl radicals during cress seed germination and elongation growth. Plant Physiol. 150, 1855–1865 (2009).
Venisse, J. S., Gullner, G. & Brisset, M. N. Evidence for the involvement of an oxidative stress in the initiation of infection of pear by Erwinia amylovora. Plant Physiol. 125, 2164–2172 (2001).
Dalla Costa, L. et al. Development of a Taqman Real-time PCR method to quantify nptII in apple lines obtained with ‘established’ or ‘new breeding’ techniques of genetic modification. Food Anal. Methods 245, 643–652 (2019).
Campa, M. et al. HIPMis a susceptibility gene of Malus: reduced expression reduces susceptibility to Erwinia amylovora. Mol. Plant-Microbe Interact. 32, 167–175 (2019).
Pompili, V., Dalla Costa, L., Piazza, S., Pindo, M. & Malnoy, M. Reduced fire blight susceptibility in apple cultivars using a high-efficiency CRISPR/Cas9-FLP/FRT-based gene editing system. Plant Biotechnol. J. 18, 845–858 (2020).
Beranger, R. et al. Agricultural and domestic pesticides in house dust from different agricultural areas in France. Environ. Sci. Pollut. Res. 26, 19632–19645 (2019).
Holtappels, M., Noben, J. P. & Valcke, R. Virulence of Erwinia amylovora a prevalent apple pathogen; Outer membrane proteins and type III secreted effectors increase fitness and compromise plant defenses. Proteomics 16, 2377–2390 (2016).
Steinberg, E. M. & Beer, S. V. Creation and complementation of pathogenicity mutants of Erwinia amylovora. Mol. Plant Microbe Interact. 1, 135–144 (1988).
Acknowledgements
This work was funded by the Autonomous Province of Trento. Work in the Zipfel laboratory on the interfamily transfer of EFR is funded by the Gatsby Charitable Foundation and the 2Blades Foundation.
Author information
Authors and Affiliations
Contributions
S.P. conducted the majority of experiments and wrote the manuscript. M.C., V.P., U.S, and D.C.L. contributed to performing the experiments and revised the manuscript. C.Z. and M.M designed the project, contributed to designing the experiments, and revised the manuscript. All authors read and approved the manuscript. S.P. and M.C. conducted the experiments and wrote the manuscript. U.S., L.D.C., and V.Z. contributed to designing and performing the experiments, design and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Piazza, S., Campa, M., Pompili, V. et al. The Arabidopsis pattern recognition receptor EFR enhances fire blight resistance in apple. Hortic Res 8, 204 (2021). https://doi.org/10.1038/s41438-021-00639-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41438-021-00639-3
This article is cited by
-
This tree is on fire: a review on the ecology of Erwinia amylovora, the causal agent of fire blight disease
Journal of Plant Pathology (2023)
-
Tackling multiple bacterial diseases of Solanaceae with a handful of immune receptors
Horticulture, Environment, and Biotechnology (2022)