Abstract
Brassica species include many economically important crops that provide nutrition and health-promoting substances to humans worldwide. However, as with all crops, their production is constantly threatened by emerging viral, bacterial, and fungal diseases, whose incidence has increased in recent years. Traditional methods of control are often costly, present limited effectiveness, and cause environmental damage; instead, the ideal approach is to mine and utilize the resistance genes of the Brassica crop hosts themselves. Fortunately, the development of genomics, molecular genetics, and biological techniques enables us to rapidly discover and apply resistance (R) genes. Herein, the R genes identified in Brassica crops are summarized, including their mapping and cloning, possible molecular mechanisms, and application in resistance breeding. Future perspectives concerning how to accurately discover additional R gene resources and efficiently utilize these genes in the genomic era are also discussed.
Similar content being viewed by others
Introduction
The Brassica genus is a member of Brassicaceae (Cruciferae) and contains 39 species (http://www.theplantlist.org/)1. Among the Brassica species, six constitute U’s Triangle2: three diploid species, namely Brassica rapa (AA genome: 2n = 2× = 20), Brassica nigra (BB: 2n = 2× = 16), and Brassica oleracea (CC: 2n = 2× = 18), and three allotetraploid species, namely Brassica juncea (AABB: 2× = 4× = 36), Brassica napus (AACC: 2n = 4× = 38), and Brassica carinata (BBCC: 2n = 4× = 34). The triangle model provides the fundamental relationships among these Brassica species and is used as an important guideline for both evolutionary research and the improvement of Brassica crops via interspecies crossing to facilitate gene exchanges.
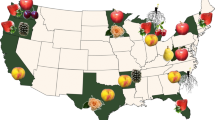
Many Brassica crops are of great economic significance, as they are cultivated as vegetables, oilseed sources, condiments, and forages3 (Table 1). Climate change, pathogen variation, and inappropriate farming methods, such as continuous and high-intensity cropping, contribute to disease outbreaks, which pose threats to current Brassica production. Various pathogens can infect Brassica crops and cause production losses, including viruses, bacteria, fungi, and oomycetes (Table 1). Among these diseases, Turnip mosaic virus (TuMV), black rot (BR), blackleg (BL), stem rot (SR), Fusarium wilt (FW), downy mildew (DM), and clubroot receive the most attention and are studied most extensively, according to a comprehensive literature search; thus, we will focus on these diseases in the following text (Fig. 1).
a Clubroot of B. napus. b Blackleg of B. napus. c Stem rot of B. napus. d TuMV-infected B. rapa. e Black rot of B. oleracea. f Downy mildew of B. rapa. g Fusarium wilt of B. oleracea. Arrows indicate part of the infected areas with typical symptoms. Images in a, e, f, and g were acquired by Honghao Lv in diseased fields in Alberta, Canada, and Hebei, Beijing, and Gansu, China, respectively. Images in b, c, and d were acquired in diseased fields in Hubei, Jiangsu, and Beijing, China, respectively, and were provided by Dr Xiaohui Cheng from the Oil Crops Research Institute, Chinese Academy of Agricultural Sciences (CAAS), Dr Qi Peng from Jiangsu Academy of Agricultural Sciences, and Dr Guoliang Li from the Institute of Vegetables and Flowers, CAAS, respectively
Traditional approaches for disease prevention include agricultural, physical, chemical, and biological controls, and integrated pest management (IPM) strategies. Physical approaches, such as high-temperature treatment and light trapping, chemicals, such as fungicides and bactericides, and biological agents, such as Bacillus subtilis and arbuscular mycorrhizae, are frequently used. IPM has been extensively studied and can achieve some effect for certain diseases. However, the approaches are often complicated, costly, and/or environmentally damaging. In contrast, natural resistance in Brassica hosts is the most desirable strategy and could be integrated with other approaches for high-efficiency disease control. Two types of plant immunity have been identified to date: pathogen/microbe-associated molecular pattern (PAMP/MAMP)-triggered immunity, which is activated by cell surface-localized pattern recognition receptors by the recognition of PAMPs/MAMPs, and effector-triggered immunity activated by host resistance (R) genes through the recognition of pathogen-specific effector molecules, which is in accord with the gene-for-gene theory4,5. Most R genes identified to date encode nucleotide-binding leucine-rich repeats (NB-LRRs), including coiled-coil NB-LRRs (CC-NB-LRRs) and Toll interleukin 1 receptor NB-LRRs (TIR-NB-LRRs). Moreover, some R genes encode receptor-like kinases (RLKs), transmembrane receptor-like proteins (RLPs), cytoplasmic kinases, and proteins with atypical molecular motifs6,7,8,9. Various R genes with flexible molecular mechanisms provide powerful weapons that protect the plant host from pathogens.
In recent years, many R genes have been identified and successfully applied to improve Brassica crop resistance against various diseases, which not only ensures Brassica production but also facilitates the discovery of host–pathogen interactions. Moreover, the genomic era characterized by massive genome and omic data has made fast and accurate R gene studies possible. The release of the reference genome data of the six Brassica species in addition to B. carinata has provided vital information for determining the genetic and molecular basis of disease resistance10,11,12,13,14. Since the 2010s, researchers have performed extensive, high-quality genomic, postgenomic, and omic studies in Brassica species and have discovered a variety of R genes and closely related genes, which not only provide further insight into the resistance molecular mechanism and host–pathogen coevolutionary arms race but also facilitate accurate molecular breeding at the whole-genome level.
Turnip mosaic virus
TuMV is the most prevalent viral disease of Brassica crops and causes heavy production losses. In 1921, the disease was first reported in B. rapa in the United States15, followed by reports in B. oleracea in the UK16, and in B. napus in China17. TuMV is currently threatening worldwide Brassica crop production, especially in Europe, Asia, and North America, resulting in a production loss of over 30%18,19. TuMV exhibits a high level of variation and 12 pathotypes have been revealed thus far, among which 1, 3, and 4 are the most prevalent20. The disease is difficult to control due to its rapid variation and nonpersistent mode of transmission by more than 89 aphid species19,21. Fortunately, a series of R genes or quantitative trait loci (QTLs) have been characterized and applied in Brassica resistance breeding.
Resistance has mostly been uncovered from the A genomes of B. rapa and B. napus. More than ten TuMV R genes have been characterized in Brassica crops thus far (Table 2). TuRB01, a monodominant gene for pathotype 1, was first located by Walsh et al.22 to a 7.2 cM segment on chromosome N6 of B. napus. TuRB01b was delimited to a 2.9 Mb segment of A06 from B. rapa and comparative analysis showed that TuRB01 and TuRB01b might be similar to each other23. TuRB02, characterized in the B. napus C genome, determines the degree of susceptibility21. TuRB03, a monodominant gene controlling pathotype 4 resistance, was localized to a 7.9 cM region on N6 in B. napus24. retr01 represents the first mapped recessive gene in Brassica species25; other recessive genes from B. rapa include rnt1 and trs, which were mapped to R6 and A4, respectively26,27. Using bulked segregant analysis by sequencing (BSA-seq), Shopan et al.28 identified another recessive gene, retr03, in B. juncea. These mapping studies have facilitated the isolation of candidate genes. The dominant gene ConTR01 and the recessive genes retr01, retr02, and retr03 are all assumed to be eIF-encoding genes, whereas TuRB07, a monodominant gene from B. rapa, was shown to encode a CC-NB-LRR29.
Molecular markers located close to R loci have been successfully applied in the breeding process through marker-assisted selection (MAS). For example, considering that molecular marker types including amplified fragment length polymorphism (AFLP), random amplified polymorphic DNA (RAPD), and restriction fragment length polymorphism (RFLP) markers present low efficacy and accuracy, Li et al.30 designed two Kompetitive Allele-Specific PCR (KASP) markers according to a single-nucleotide polymorphism (SNP) of the TuMV R gene retr02, which might be applied in high-throughput MAS. In addition, some resistance genes have been directly applied in resistance breeding. For example, eIF(iso)4E variants have been transferred to B. rapa and the transgenic plants display broad-spectrum resistance31.
Black rot
BR, the causal agent of which is Xanthomonas campestris pv. campestris (Xcc), is one of the most prevalent bacterial diseases in Brassica crops. The disease was first described in the United States by Garman32 as a disease of cabbage. The disease has since been identified on all Brassica-growing continents, especially in Asia, Europe, and North America, bringing about considerable losses to Brassica production33,34. Xcc exhibits a high level of genetic diversity and 11 races distributed worldwide have been discovered to date, with 1 and 4 being the most prevalent and very virulent to many commercial cultivars35,36,37. In recent years, only a few resistance resources have been identified, greatly hindering the resistance breeding process.
Most BR resistance research conducted to date has focused on QTL analysis or preliminary mapping. The first mapping analysis of BR resistance in cabbage revealed two major QTLs38. Vicente et al.39 placed a major locus, Xca4, in B. napus in linkage group N5. Soengas et al.40 reported broad-spectrum resistance in B. rapa, with a cluster of major-effect QTLs being characterized on A06, each of which could explain 24%–64% of the observed phenotype variation. High-throughput markers allow improved mapping accuracy. Kifuji et al.41 applied expressed sequence tag-based SNP markers to map resistance genes in cabbage and three QTLs, including the major QTL-1, were detected. Tuno et al.42 analyzed BR resistance QTLs and the major QTL XccBo(Reiho)2 was detected on C8. Saha et al.43 mapped the Xcc race 1 resistance gene Xca1bo in the cauliflower line BR-161 within a 1.6 cM interval. Sharma et al.44 first developed a B. carinata F2 mapping population and mapped the BR race 1 resistance locus Xca1bc to a 6.6 cM interval. Lee et al.45 first developed genome-wide SNP markers and identified one major QTL on C3 in cabbage. In total, more than 20 QTLs have been placed on over eight Brassica chromosomes, indicating that the resistance to BR is highly complicated (Table 2).
Although resistance genes are far from being isolated, some molecular markers closely linked to them have been widely adopted in MAS. Kalia et al.46 converted the formerly developed RAPD and inter-simple sequence repeat (ISSR) markers to sequence-characterized amplified region (SCAR) markers, showing great potential for MAS in cauliflower breeding. Using a B. carinata-derived F2 population, Sharma et al.44,47 developed markers linked to BR resistance, which were further used in the selection of introgression lines (ILs) from B. carinata to cauliflower.
Blackleg
BL or stem canker is a disastrous fungal disease for Brassica crops caused by Leptosphaeria maculans (Lm). The first epidemic was reported on cabbage in Wisconsin48. However, BL has become a problem in terms of mass infection of oilseed rape only since the middle of the twentieth century, especially in Australia, North America, and Europe49,50,51. Lm exhibits a high level of diversification and has been assigned to different races/pathotypes52,53,54,55. Resistance gene mapping work has been conducted since the 1990s and some cultivars with improved resistance to BL are available.
Most BL resistance genes/QTLs originated from the B. napus A genome (Table 2). Ferreira et al.56 first applied a double haploid (DH) population from B. napus to localize the major locus LEM1 on N7. Using a similar method, Dion et al.57 identified another major gene, LmFr1. Mayerhofer et al.58 detected a major locus, LmR1, and cosegregating markers were developed59. Delourme et al.60 reported the mapping of resistance loci in two genomic regions and a cluster consisting of five R genes was proposed as the candidate. Fine mapping work was conducted extensively after 2010. Long et al.61 identified two resistance genes, BLMR1 and BLMR2, and fine mapping of BLMR1 resulted in the closest marker distance of 0.13 cM. Jestin et al.62 used an association mapping method to characterize the molecular diversity using 128 oilseed rape accessions and identified five novel alleles. Rayman et al.63 positioned a new major locus, Rlm4, and the deposited region was further analyzed, with several candidates being characterized64. In addition, BL resistance loci have been transferred from wild relatives of B. rapa and B. oleracea to B. napus65. Yu et al.66,67 mapped BL resistance derived from the wild relative and LepR1-LepR3 were identified. Larkan et al.68,69 employed map-based cloning to isolate LepR3, which encoded an RLP, representing the first cloned BL disease resistance gene; the authors further isolated the Rlm2 gene, which is an allelic variant of LepR370. More recently, the authors cloned another BL resistance gene, Rlm9, which encodes a wall-associated kinase-like protein, a newly discovered class of race-specific plant RLK resistance genes71. In addition to the major locus, some QTLs have also been characterized, including six and four that are stable under different environmental conditions72,73.
Currently, Brassica cultivars with improved resistance to BL are frequently cultivated due to extensive R gene mapping work. In addition, MAS is often integrated with other breeding methods to shorten the breeding period. For instance, Yu et al.65 described the successful introgression of BL resistance from wild B. rapa subsp. sylvestris to B. napus via interspecific hybridization and MAS, which generates a series of resistant cultivars. In addition, based on both the major genes and QTLs identified, the next breeding effort could involve a combination of qualitative and quantitative loci to provide more durable resistance74.
Stem rot
SR of Brassica crops is a fungal disease caused by Sclerotinia sclerotiorum (Ss). SR is a worldwide catastrophe for Brassica production, especially in oilseed rape, in which yield losses can range from 10% to 80%, with low oil quality75. Ss was reported as the pathogen of SR in 1837 and is now found worldwide76,77. Ss exhibits little host specificity and eight pathotypes have been identified78,79. Ss is able to persist for several years in the soil and the most desirable approach for its control to use resistant cultivars77,80. Unfortunately, to date, no highly resistant resource has been characterized in Brassica crops, making breeding work for SR resistance difficult.
Almost all the mapping work in this context has focused on B. napus; however, only partial resistance has been characterized in both the A and C genomes (Table 2). Zhao and Meng81 first identified three QTLs for leaf resistance and three other QTLs for stem resistance in the seedling and adult stages, respectively, but no common QTLs. Zhao et al.82 identified eight and one QTL involved in two segregating DH populations, with each explaining 6–22% of the observed variance, still with no common QTLs. Yin et al.83 detected ten, one, and ten QTLs in one DH population using three inoculation procedures, and only two common QTLs were detected. Wu et al.84 identified three QTLs at the seedling stage for leaf resistance and ten QTLs for stem resistance at the adult stage. Two major QTLs could be detected repeatedly and a candidate resistance gene, BnaC.IGMT5, was first identified. These studies revealed abundant QTLs but seldom common ones, indicating the complicated genetic structure of these plants. Recently, the release of the B. napus genome sequence has strongly facilitated mapping work. Fomeju et al.85 first adopted a genome-wide association study (GWAS) using 116 materials genotyped with 3228 SNPs and the results indicated that 64 genomic regions are involved in SR resistance. Wei et al.86 combined GWAS and SNP array analyses using 347 B. napus accessions and 17 significant regions were located on the A8 and C6 chromosomes. These SNPs on Chr. A8 were placed in a 409 kb segment, with candidate genes being suggested. Using a similar method, Wu et al.87 genotyped 448 accessions and 26 SNPs corresponding to three loci were associated with SR resistance. In total, 39 candidates were proposed. Gyawali et al.88 performed a GWAS using microsatellite markers in a global collection of 152 accessions and found that 34 loci were significantly associated. To date, many loci opposing SR have been characterized but none have been functionally characterized.
Considering that high resistance to SR in B. napus is not available, researchers tend to investigate wild Brassica relatives for novel germplasm, such as Berteroa incana and Brassica cretica. MAS combined with distant hybridization plays significant roles in resistance transfer. For example, Mei et al.89,90,91 successfully introgressed resistance from wild B. incana into B. napus through hexaploidy hybridization and MAS using newly developed simple sequence repeat (SSR) markers and phenotype evaluation.
Fusarium wilt
FW disease, caused by the fungus Fusarium oxysporum f. sp. conglutinans (Foc), is posing a threat to Brassica production worldwide, especially for cole crops92,93. FW was first observed on cabbage by Smith94 in the United States in 1895. Since 1910, FW has spread quickly from the United States to almost the whole world95,96. To date, two Foc races have been reported, but only race 1 is found worldwide97,98,99. FW is a soil-borne disease and Foc can survive for more than 10 years, even without a host100. Currently, type A resistance conferred by a dominant monogene has been identified and applied successfully. However, race 2 can overcome type A resistance, indicating that single resistance application is at high risk.
Most resistance resources have been identified in B. oleracea (Table 2). Specifically, two types of resistance have been characterized, i.e., A and B. Type A resistance is stable under high or low temperature and follows a single dominant inheritance pattern; type B polygenic resistance is unstable under high temperatures above 24 °C101,102. The type A single dominant resistance gene for Foc race 1 has been explored extensively in the last several years. The FW R gene FocBo1 was first mapped to linkage group seven using both BSA and QTL analysis by Pu et al.103. Lv et al.104,105 generated a genetic linkage map based on a cabbage DH population and mapped the R gene FOC1 to a 1.8 cM interval between two adjacent InDel markers. The authors further mapped the candidate gene FOC1 to a repredicted Bol037156, which encodes a TIR-NBS-LRR, using an enlarged population. Shimizu et al.106 also mapped the resistance locus FocBo1 by using 139 recombinant F2 plants and identified a candidate gene, Bra012688. The two mapped candidates are homologous with high identity. However, the functions of these genes remain to be identified.
Type A resistance to Foc race 1 conferred by a dominant single gene, FOC1, has been successfully mapped and molecular markers have been developed and applied to generate various resistant cultivars. In addition, MAS using these markers has been combined with other breeding methods to promote the breeding process. For example, Lv et al.107 reported the use of isolated microspore cultures with MAS to rapidly obtain target DH lines with FW resistance, which could be used directly in resistance breeding, thereby shortening the breeding period by 2–3 years.
Downy mildew
DM is a foliar disease of Brassica crops and the causal agent is the oomycete pathogen Hyaloperonospora brassicae (Hb)108,109. DM causes considerable yield losses to all Brassica crops worldwide, especially in Europe, Asia, and Australia110,111. Physiological races or pathotype variations have also been described in various studies; however, few studies have achieved clear race differentiation112,113,114. Notably, Coelho et al.115 summarized six pathotypes and suggested five major-effect R loci corresponding to the observed phenotypes. The ideal control approach for DM is to breed genetically resistant varieties. To date, several R loci have been mapped and applied in breeding.
Resistance to DM is thought to be distinct at the Brassica seedlings and adult stages. Resistance mapping work has identified several R genes/loci (Table 2). In B. oleracea, the first locus conferring resistance in the broccoli seedling stage was placed in a linkage group116 and was found to be located close to the glucosinolate-related gene BoGsl elong117. Another single dominant resistance gene expressed at the adult stage was identified in broccoli and was named Pp523118. The genomic region containing this gene was further analyzed using SCAR and cleaved amplified polymorphic sequence (CAPS) markers, as well as two bacterial artificial chromosome (BAC) libraries119,120. In B. rapa, QTLs conferring seedling-stage resistance were discovered. Using a genetic linkage map generated with a DH population, the major-effect locus BraDM was delimited to a region spanning 2.9 cM in the A08 linkage group121. For adult-stage resistance, a monodominant gene, BrRHP1, was localized to a 2.2 Mb interval on the A01 linkage group122. In recent years, mapping methods based on high-throughput resequencing have greatly promoted the identification of R genes. For example, using a high-density SNP-based map, a major locus, sBrDM8, was localized to a physical segment of ~228 kb, with one candidate kinase gene, Bra016457123.
The developed markers closely located with these R loci have been adopted for resistance breeding through MAS and have greatly contributed to resistance breeding. For example, Yu et al.124 converted the closely linked RAPD marker K14-1030 to a SCAR marker, which greatly improved selection efficiency in the progenies.
Clubroot
Clubroot (CR) caused by Plasmodiophora brassicae (Pb) is now threatening almost all Brassica crops worldwide. Pb is neither a fungus nor a slime mold and has been classified into the new taxon Rhizaria125. CR was first reported in Russia in 1878126 and rapidly expanded to Europe, Asia, and America, becoming one of the most serious problems for Brassica production around the world125,127. Pb exhibits complex pathotypes and two differentiation systems are used extensively: the Williams system and the European clubroot differential set128,129,130,131. The variation in this pathogen and its ability to survive in soil in the form of resting spores make it difficult to control132,133,134. Thus, breeding resistant cultivars represents an ideal control method. Currently, extensive studies addressing CR have generated the largest number of resistance loci among all Brassica diseases (Table 2) and MAS has been widely used for resistance improvement.
In B. rapa, several important CR genes conferring complete resistance in accessions against specific pathogen isolates have been identified. The mapping and cloning of the CRb/CRa loci took over 20 years. CRa was mapped and the candidate gene encodes a TIR-NBS-LRR135,136. Another locus, CRb, from the Chinese cabbage cultivar CR Shinki, was extensively mapped to a final 84 kb region137,138,139. Kato et al.140 identified another CR resistance locus, CRbKato, in Akiriso Chinese cabbage. Hatakeyama et al.141 further determined that CRbKato and CRa were the same TIR-NB-LRR gene, whereas CRb might be a different locus. Another example is the Crr1-4 genes from turnip, which were initially primarily mapped using different molecular markers and populations142,143,144,145. Through fine mapping, Hatakeyama et al.146 discovered that Crr1 consists of two genes: Crr1a and Crr1b. The former encodes a TIR-NB-LRR and was functionally confirmed. With the development of genomic and molecular genetics, several loci were further identified using newly developed marker techniques147. Yu et al.148 applied BSA-seq and identified a novel resistance gene, Rcr1, and two candidates encoding TIR-NB-LRRs. Huang et al.149 adopted KASP markers and BSR-seq strategies to finely map Rcr2 to a 0.4 cM interval, identifying two TIR-NBS-LRRs as candidates. Using BSA-seq, Pang et al.150 identified the new locus CRd in a 60 kb region on chromosome A03, which is located upstream of Crr3.
In B. oleracea, CR resistance appears to be determined by quantitative genes. Figdore et al.151 first identified three QTLs conferring resistance to Pb race 7 in broccoli. In the resistant kale line C10, Grandclément and Thomas152 performed QTL detection with RAPD markers and the results indicated at least two types of genetic mechanisms. Rocherieus et al.153 further found two to five QTLs depending on which of five pathotypes were used and Pb-Bo1 was uncovered for all Pb isolates, accounting for 20.7–80.7% of the phenotypic variation. In another resistant kale line, K269, Moriguchi et al.154 and Nomura et al.155 detected two and three loci, respectively, conferring resistance to different isolates. In cabbage, Voorrips et al.156 first reported two major QTLs, pb-3 and pb-4. Nagaoka et al.157 identified a major QTL, pbBo(Anju)1, from the cabbage accession Anju. Lee et al.158 employed the genotyping by sequenceing (GBS) technique and a QTL survey to reveal two and one major loci for races 2 and 9, respectively. These loci showed positions close to the previously identified resistance loci in B. oleracea but in distinct locations from those discovered in B. rapa, indicating divergence of R loci between the Brassica A and C genomes.
For B. napus, a few loci conferring resistance to various isolates have been characterized. Landry et al.159 identified two QTLs controlling CR resistance to race 2, which contributed 58% and 15% of the observed phenotypic variation. Manzanares-Dauleux et al.160 reported the mapping of R loci in Darmor-bzh and identified one major gene, Pb-Bn1. Using a DH population, Werner et al.161 detected 19 QTLs that conferred resistance to 7 different isolates, but none of them could confer resistance to all these isolates. Fredua-Agyeman and Rahman162 mapped canola CR resistance to a DNA segment that comprised 12 markers linked to the CRa locus, indicating its possible A genome origin. Hasan and Rahman163 used rutabaga-derived populations for resistance mapping and characterized a genomic segment on chromosome A8 conferring resistance to all five tested pathotypes. GWAS enables rapid detection of recombinants and variations using natural populations based on whole-genome SNP data. Li et al.164 first applied GWAS to 472 accessions to identify CR resistance with the 60 K Brassica Infinium SNP. A total of nine loci were characterized through integrative analysis, with seven of them being novel and six of them being in the C genome.
The closely linked markers and resistance genes have been widely used in Brassica CR resistance breeding, generating a series of resistant cultivars that successfully control CR in many areas. For example, considering that high resistance is found mostly in B. rapa, researchers have frequently applied interspecies crossing to facilitate R gene transfer combined with MAS and phenotype evaluation165,166.
Future perspectives
Creating novel germplasms via close or distant hybridization
For a certain Brassica species, the resistance resources for diseases such as BR, BL, SR, and CR are highly limited. In general, the A genome is rich in TuMV, BL, DM, and CR resistance, whereas the B genome possesses BR and BL resistance, and the C genome harbors SR, FW, and DM resistance. Fortunately, the six species in the Brassica genus and others, such as B. incana, B. cretica (C genome), and Brassica fruticulosa (B genome), as well as its close Brassicae relatives, could be used to facilitate resistance gene exchanges in breeding programs.
Interspecies crossing within the Brassica genus is widely adopted using embryo rescue, reciprocal crossing and MAS. For example, there are notably few BR-resistant resources in the C genome of B. oleracea, whereas high resistance is present in both the A and B genomes. Thus, interspecies hybridization has been used to transfer and utilize the resistance found in the A and B genomes. Tonguç and Griffiths167 developed interspecific hybrids between B. oleracea and B. juncea accession A19182, which show resistance to both Xcc races 1 and 4. The progenies displayed resistance to both races. Similarly, Sharma et al.47 successfully transferred BR resistance from B. carinata to cauliflower. In addition, distant hybridization has been used by breeders to generate novel resistant germplasms. For example, only partial resistance to BL has been characterized in B. napus thus far, whereas complete or highly resistant lines are not available. In contrast, high-level resistance was observed in a few wild species, such as Erucastrum cardaminoides, Diplotaxis tenuisiliqua, and Sinapis arvensis. Snowdon et al.168 performed a cross between B. napus and S. arvensis, and through resistance tests and molecular analyses, the ILs were successfully identified from the BC3 progenies, which exhibited high resistance at both the seedling and adult stages. Garg et al.169 first obtained high levels of SR resistance from crosses between B. napus/B. juncea and E. cardaminoides/D. tenuisiliqua/Erucastrum abyssinicum. The novel resistance germplasms generated in such studies provide valuable materials in future breeding programs for Brassica crops. However, there is still much work to be done for them to be practically applied in commercial cultivars. For example, there are rich resources with CR resistance in B. rapa but very few in B. oleracea, and breeders have spent ~17 years introgressing resistance from Chinese cabbage (B. rapa subsp. pekinensis) cv. Parkin to different B. oleracea cultivars, which are now widely used170.
MAS as an approach for high-efficiency integrated breeding in the genomic era
Molecular markers are inheritable and detectable genomic segments. The techniques for molecular markers and gene mapping have been significantly improved from the 1990s to the present in the genomic era. First, methods such as RAPD, AFLP, and RFLP, representing the first-generation markers with low efficiency, were constantly applied to map the resistance genes and QTLs for CR, BR, BL, and TuMV. Since the 2000s, convenient and highly efficient markers, including SSRs, microsatellites, and InDels, have gradually become mainstream and have been applied for the discovery of novel R loci for SR, CR, and BL. As the 2010s, mapping methods based on high-throughput sequencing data have developed rapidly, especially SNP-based methods, such as KASP markers and microarrays, BSA/BSR and GWAS. Based on whole-genome level mutations and their association with trait values, GWAS enables fast and accurate target trait gene characterization using natural populations. For example, Wei et al.86 applied combined GWAS and SNP array analyses to 347 B. napus accessions to detect resistance to SR and identified 17 significant associations on two chromosomes. These SNPs on chromosome A8 were localized to a segment of 409 kb, with candidate genes being proposed. In addition, KASP technology possesses high assay robustness and accuracy, and allows notable savings in terms of cost and time. Huang et al.149 adopted KASP markers and BSR-seq strategies to rapidly identify the Rcr2 locus in the CR-resistant Chinese cabbage cultivar Jazz and Rcr2 was delimited to a 0.4 cM region, where two TIR-NBS-LRRs were identified as candidates. In addition, based on the sequence variations of the TuMV resistance gene retr02, Li et al.30 designed a KASP marker that could be used to accurately genotype the allele in Chinese cabbage accessions.
In many cases, there is one main disease in a specific production region and application of cultivars with resistance to that main disease is appropriate to realize the balance between defense and growth. However, for some continuous or high-intensity cropping regions, various pathogens may accumulate; therefore, cultivars with multiple resistances to different diseases are in great need. Currently, MAS using abundant molecular markers enables us to realize the pyramiding of R alleles for different diseases and breed multiresistant cultivars. MAS combined with other methods, such as hybridization and microspore culture, has greatly shortened the breeding circle. These cultivars are now available on the market, such as cabbage cv. Zhonggan 628 (Institute of Vegetables and Flowers, Chinese Academy of Agricultural Sciences, Beijing, China), with resistance to TuMV and FW; Chinese cabbage cv. Jingchun CR1 (Beijing Academy of Agriculture and Forestry Sciences, Beijing, China), with resistance to TuMV, DM, and clubroot; and oilseed rape cv. Huashuang 5R (Huazhong Agricultural University, Wuhan, Hubei, China), with resistance to TuMV and clubroot, and tolerance to SR.
The genomic era is also characterized by high-efficiency integrated breeding (HIB), in which multiple methods are combined, including traditional ways, such as microspore culture, backcrossing, and distant introgression, and modern ways, such as MAS, gene editing, and genome design (Fig. 2). During HIB, genomic background analysis is helpful in eliminating undesirable linkage drags and rapidly identifying desirable individuals. For example, in a study by Liu et al.171, resistance-specific markers and genome background markers were used to breed cabbage with resistance to FW. By combining these methods with microspore culture and backcrossing, the authors presented a rapid and effective approach for generating FW-resistant ILs in the BC2 generation. Notably, the quickly emerging gene-editing technique helps realize accurate alteration of the target DNA sequence. Ma et al.172 applied CRISPR/Cas9-mediated multiple gene editing in cabbage, with the targets BoPDS, BoSRK, and BoMS1, and successfully generated albino, self-compatible, and male sterile lines, showing its great power in improving plant traits.
a Selfing using honeybees is one of the most traditional breeding methods. b A combined use of microspore culture and MAS helps promote the breeding cycle in B. oleracea107. c BR resistance introgression from B. carinata to B. oleracea using distant hybridizing and embryo rescue47. d SNP-based high-throughput KASP markers prove efficient and cost saving in genotyping during MAS in B. rapa30. e Whole-genome background analysis helps eliminate the undesired linkage drags during MAS in B. oleracea207. f Pyramiding both the qualitative and quantitative R loci generates durable BL resistance in B. napus74. g CRISPR/Cas9-based gene editing helps knockout multiple target genes in B. oleracea172. h Expressing the CP gene from TuMV confers high resistance in B. napus200
Pyramiding of qualitative and quantitative loci to acquire durable resistance
A single resistance gene is easily overcome because of pathogen variations and global climate changes. For example, a few B. rapa, B. oleracea, and B. napus varieties resistant to specific Pb races have been successfully cultivated. However, all these varieties lose resistance within a few years. At the same time, vast genetic variability in the clubroot pathogen Pb and infection by multiple races have been reported129,130,131. For the BR pathogen Xcc, pathogen variations are frequently discovered and at least 11 races have been reported thus far35,36,37.
More durable resistance is urgently needed to ensure Brassica crop production. Durable resistance was first proposed by Johnson173 as resistance that maintains effectiveness during long-term widespread application. Complete race-specific resistance genes are very effective in a short period but are easily overcome by the pathogens; polygene-inherited resistance is thought to be more durable, but its effects might be unstable owing to variable environmental conditions174. Thus, pyramiding qualitative genes with major quantitative loci in cultivars represents an ideal means to ensure the effectiveness and durability of resistance. An example is the utilization of BL resistance in B. napus. Brun et al.74 evaluated a cultivar with single race-specific Rlm6-mediated resistance and another cultivar with both Rlm6 and quantitative resistance in a 5-year field experiment. The single Rlm6 resistance became ineffective as soon as the third cropping season. When integrated with quantitative resistance, however, Rlm6-mediated resistance maintained effectiveness until the seventh year. Another cultivar, Jet Neuf, has been widely used for as long as 10–15 years, both as a cultivar and as a source of resistance. Jet Neuf was shown to harbor both polygenic resistance and the mono R gene Rlm450,175,176. This pyramiding model is also supported and used in resistance breeding against BR39, SR177, and CR127. Thus, combining quantitative resistance with single R genes is a promising strategy for resistance breeding. In addition, coevolution between the host and the pathogen has been extensively studied; e.g., the mapped seven BL resistance genes/alleles and their contrasting avirulence genes have indicated a rapid and fierce arms race between Brassica hosts and Lm, and the single use of one type of pyramiding could pose a high selection pressure on the pathogen and thus raise the risk of pathogen mutation and host resistance loss. Thus, the rationalized pyramiding of dissimilar sets of resistance genes/QTLs should be deployed in different cultivars to provide heterogeneity in the selection pressure on the pathogen population and result in more durable resistance178. In addition, pyramided resistance could be integrated with agricultural, physical, chemical, and biological controls to realize IPM, further maximize durability and guarantee stability.
Digging deeper into the resistance mechanism in prebreeding studies
To date, hundreds of R loci in Brassica crops have been characterized; however, candidates have only been found for approximately a dozen of them and their mechanisms are far from being revealed compared with those of the model plants Arabidopsis, tobacco, and rice. Current molecular and omics methods, including transcriptomics, proteomics, and metabolics, provide new opportunities for mining genes in the resistance-regulating network, which could be either directly used in resistance breeding or used in indirect prebreeding studies to promote our understanding of Brassica–pathogen interactions. For example, in SR, transcriptomic and proteomic studies have revealed a series of key genes associated with the response to pathogen infection, including RLKs, NBS-LRRs, calcium-binding proteins, PRs, TFs, and polygalacturonase inhibitor proteins, which are associated with plant–pathogen interactions, the mitogen-activated protein kinase signaling cascade, plant hormone biosynthesis and signaling, and oxalic acid (OA) metabolism179,180,181,182,183,184. These genes were subjected to functional clarification and prebreeding studies. For instance, overexpression of BnMPK4185, BnWRKY33186, chimeric chitinase187, OA188, and PGIP2189 can enhance host resistance. In addition, the sequenced Brassica accessions do not contain all R genes due to variations between individuals, whereas the establishment of the pangenomes could facilitate gene mining from a wider platform. Bayer et al.190 performed a comparative analysis of resistance gene analogs (RGAs) in the pangenome of B. oleracea and identified 59 RGAs linked to SR, CR, and FW resistance, some of which probably resulted from a recombination within R gene clusters.
Achieving expanded knowledge of the molecular host–pathogen interactions could provide evidence for disease control. Protein–protein interactions in the Brassica-TuMV system are a heavily researched topic. Previous studies have shown that the cytoplasmic inclusion protein determines the viral avirulence for TuRB01/01b/04, whereas P3 determines the avirulence of TuRB03/05191,192,193,194. Another example is the plant eukaryotic initiation factor 4E (eIF4E) family, which is well characterized as a key factor during the invasion of several potyviruses. The viral protein genome-linked (VPg) protein of potyviruses interacts directly with the host eIF4E/eIF(iso)4E and determines virulence195,196. This eIF4E-mediated mode of resistance is generally strong and broad spectrum197,198. In Brassica, the recessive R genes to TuMV, including retr01, retr02, and trs, have been identified to encode eIF(iso)4E25,27,199. Furthermore, researchers induced different key amino acid mutations in eIF(iso)4E by a systematic knowledge-based approach to interrupt the interaction between TuMV VPg and host eIF(iso)4E, and transgenic plants with eIF(iso)4E variants display high and broad-spectrum resistance31. This example shows the great potential of artificially designed R alleles/mutants in resistance breeding. In addition to the direct application of the identified resistance genes, genes from TuMV have also been used in resistance breeding as a method of host-induced gene silencing, especially the CP gene. The CP protein can accumulate in host cells and inhibit virus replication, thereby conferring resistance. Successful resistance enhancement via the CP gene strategy has been reported in Brassica crops, including oilseed rape and Chinese cabbage200,201. In Brassica crops, although over ten R genes have been characterized to date, most avirulence or interaction genes in the pathogens have not yet been thoroughly characterized (Fig. 3). Thus, the next emphasis should be on the establishment of global collections of pathogen isolates for the identification of important avirulence or interaction genes. In addition, for the host, the development of a series of ILs through recurrent backcrossing to “Mendalize” the quantitative loci would be beneficial. For example, to clarify the relationship between the Lm isolates and the mapped seven BL resistance genes/loci, Larken et al.202 introgressed each of the seven R genes/loci into a common susceptible B. napus DH line through reciprocal backcrossing, producing single R gene ILs that could provide for the accurate assessment of Avr-R gene interactions by avoiding non-Avr-dependent alterations.
ASMT, N-acetylserotonin O-methyltransferase; CC, coiled-coil domain; eIF(iso)4E, eukaryotic translation initiation factor isoform 4E; eIF2Bβ, eukaryotic translation initiation factor 2Bβ; LRR, leucine-rich repeat; NB, nucleotide-binding domain; PK, protein kinase; RLP, receptor-like protein. *Putative genes that have not been functionally validated. ???The avirulence or interaction genes in the pathogens that have not yet been characterized
To date, using the Arabidopsis–Brassica pathogen pathosystem, great progress has been made in the characterization of resistance genes and their molecular mechanisms, which can provide critical clues for Brassica resistance studies, as both Brassica and Arabidopsis belong to Cruciferae. For example, the interactions between Hb and Arabidopsis have been well established. To date, more than 10 RPP loci conferring resistance to Hb have been cloned in Arabidopsis, most of them being NLRs that regulate the activation of programmed cell death203,204. In addition, some important genes, such as EDS1, NDR1, PRs, NPRs, and WRKYs, have been shown to play important roles in DM resistance in Arabidopsis205. In addition, the pathogen effectors have been isolated and the host–pathogen interactions have also been well characterized206. As many R genes have been cloned in Arabidopsis, the orthologous genes in Brassica can be investigated, which will greatly facilitate the cloning of these genes and the clarification of their molecular functions.
Concluding remarks
Brassica species comprise many economically important crops, but their production is constantly threatened by emerging diseases, such as TuMV, BR, FW, DM, and clubroot. The most ideal measure is to mine and utilize the resistance genes of the Brassica crop hosts themselves. Fortunately, the development of genomics, molecular genetics, and biological techniques enables us to rapidly discover more than 100 R genes/loci. However, only a dozen of them have strong candidates and are still not well functionally validated. Moreover, only a small portion of them has been applied in resistance breeding. Thus, the next efforts should be more accurate identification of the R genes and clarification of their molecular mechanisms using emerging high-efficiency genomic, postgenomic, and omic methods, and more efficient application of the R resources through an integration of approaches, such as haploid culture, MAS, distant introgression, genome design, pyramiding, and transgenic breeding, to control the diseases and secure Brassica production.
References
Al–Shehbaz, I. A., Beilstein, M. A. & Kellogg, E. A. Systematics and phylogeny of the Brassicaceae (Cruciferae): an overview. Plant Syst. Evol. 259, 89–120 (2006).
Nagaharu, N. Genome analysis in Brassica with special reference to the experimental formation of B. napus and peculiar mode of fertilization. Jpn. J. Bot. 7, 389–452 (1935).
Dixon, G. R. in Vegetable Brassicas and Related Crucifers (eds Jeff, A. & Alun, R.) Ch. 1, 1–2 (CABI, Wallingford, 2007).
Flor, H. H. Current status of the gene–for–gene concept. Annu. Rev. Phytopathol. 9, 275–296 (1971).
Chisholm, S. T., Coaker, G., Day, B. & Staskawicz, B. J. Host–microbe interactions: shaping the evolution of the plant immune response. Cell 124, 803–814 (2006).
Jeffery, L. D. & Jonathan, D. G. Plant pathogens and integrated defense responses to infection. Nature 411, 826–833 (2001).
Jones, J. D. & Dangl, J. L. The plant immune system. Nature 444, 323–329 (2006).
Liu, J., Liu, X., Dai, L. & Wang, G. Recent progress in elucidating the structure, function and evolution of disease resistance genes in plants. J. Genet. Genomics 34, 765–776 (2007).
Neik, T. X., Barbetti, M. J. & Batley, J. Current status and challenges in identifying disease resistance genes in Brassica napus. Front. Plant Sci. 8, 1788 (2017).
Wang, X. et al. The genome of the mesopolyploid crop species Brassica rapa. Nat. Genet. 43, 1035–1039 (2011).
Parkin, I. A. et al. Transcriptome and methylome profiling reveals relics of genome dominance in the mesopolyploid Brassica oleracea. Genome Biol. 15, R77 (2014).
Liu, S. et al. The Brassica oleracea genome reveals the asymmetrical evolution of polyploid genomes. Nat. Commun. 5, 3930 (2014).
Chalhoub, B. et al. Early allopolyploid evolution in the post–neolithic Brassica napus oilseed genome. Science 345, 950–953 (2014).
Yang, J. et al. The genome sequence of allopolyploid Brassica juncea and analysis of differential homoeolog gene expression influencing selection. Nat. Genet. 48, 1225 (2016).
Gardner, M. W. & Kendrick, J. B. Turnip mosaic. J. Agr. Res. 22, 123–124 (1921).
Smith, K. M. A virus disease of cultivated Crucifers. Ann. Appl. Biol. 22, 239–242 (1935).
Ling, L. & Yang, J. A mosaic disease of rape and other cultivated Crucifers in China. Phytopathology 30, 338–342 (1940).
Shattuck, V. I. The Biology, Epidemiology, and Control of Turnip Mosaic Virus. Horticultural Reviews Vol. 14 (John Wiley and Sons, Inc., 1992).
Walsh, J. A. & Jenner, C. E. Turnip mosaic virus and the quest for durable resistance. Mol. Plant Pathol. 3, 289–300 (2002).
Jenner, C. E. & Walsh, J. A. Pathotypic variation in Turnip mosaic virus with special reference to European isolates. Plant Pathol. 45, 848–856 (1996).
Tomlinson, J. A. Epidemiology and control of virus disease of vegetables. Ann. Appl. Biol. 110, 661–681 (1987).
Walsh, J. A., Sharpe, A. G., Jenner, C. E. & Lydiate, D. J. Characterisation of resistance to Turnip mosaic virus in oilseed rapa (Brassica napus) and genetic mapping of TuRB01. Theor. Appl. Genet. 99, 1149–1154 (1999).
Lydiate, D. J., Pilcher, R. L., Higgins, E. E. & Walsh, J. A. Genetic control of immunity to Turnip mosaic virus (TuMV) pathotype 1 in Brassica rapa (Chinese cabbage). Genome 57, 419 (2014).
Hughes, S. L. et al. Genetic mapping of the novel Turnip mosaic virus resistance gene TuRB03 in Brassica napus. Theor. Appl. Genet. 107, 1169–1173 (2003).
Rusholme, R., Higgins, E., Walsh, J. & Lydiate, D. Genetic control of broad–spectrum resistance to Turnip mosaic virus in Brassica rapa (Chinese cabbage). J. Gen. Virol. 88, 3177–3186 (2007).
Fujiwara, A., Inukai, T., Bo, M. K. & Masuta, C. Combinations of a host resistance gene and the CI gene of Turnip mosaic virus differentially regulate symptom expression in Brassica rapa cultivars. Arch. Virol. 156, 1575 (2011).
Kim, J. et al. Identification of a broad–spectrum recessive gene in Brassica rapa and molecular analysis of the eIF4E gene family to develop molecular markers. Mol. Breed. 32, 385–398 (2013).
Shopan, J. et al. Eukaryotic translation initiation factor 2b‐beta (eIF2Bβ), a new class of plant virus resistance gene. Plant J. 90, 929–940 (2017).
Jin, M. et al. Identification and mapping of a novel dominant resistance gene, TuRB07, to Turnip mosaic virus in Brassica rapa. Theor. Appl. Genet. 127, 509–519 (2014).
Li, G. et al. Development of gene–based markers for the Turnip mosaic virus resistance gene retr02 in Brassica rapa. Plant Breed. 135, 466–470 (2016).
Kim, J. et al. Transgenic Brassica rapa plants over‐expressing eIF(iso)4E variants show broad‐spectrum Turnip mosaic virus (TuMV) resistance. Mol. Plant Pathol. 15, 615–626 (2014).
Garman, H. A bacterial disease of cabbage. Ky. Agr. Exp. Stn. Rep. 3, 43–46 (1894).
Williams, P. H. Black rot: continuing threat to world crucifers. Plant Dis. 64, 736–742 (1980).
Vicente, J. G. & Holub, E. B. Xanthomonas campestris pv. campestris (cause of black rot of crucifers) in the genomic era is still a worldwide threat to Brassica crops. Mol. Plant Pathol. 14, 2–18 (2013).
Vicente, J. G., Conway, J., Roberts, S. J. & Taylor, J. D. Identification and origin of Xanthomonas campestris pv. campestris races and related pathovars. Phytopathology 91, 492–499 (2001).
Fargier, E. & Manceau, C. Pathogenicity assays restrict the species Xanthomonas campestris into three pathovars and reveal nine races within X. campestris pv. campestris. Plant Pathol. 56, 805–818 (2007).
Cruz, J., Tenreiro, R. & Cruz, L. Assessment of diversity of Xanthomonas campestris pathovars affecting cruciferous plants in Portugal and disclosure of two novel X. campestris pv. campestris races. J. Plant Pathol. 99, 403–414 (2017).
Camargo, L. E. A., Williams, P. H. & Osborn, T. C. Mapping of quantitative trait loci controlling resistance of Brassica oleracea to Xanthomonas campestris pv. campestris in the field and green house. Genetics 85, 1296–1300 (1995).
Vicente, J. G. et al. Inheritance of race–specific resistance to Xanthomonas campestris pv. campestris in Brassica genomes. Phytopathology 92, 1134–1141 (2002).
Soengas, P., Hand, P., Vicente, J. G., Pole, J. M. & Pink, D. A. Identification of quantitative trait loci for resistance to Xanthomonas campestris pv. campestris in Brassica rapa. Theor. Appl. Genet. 114, 637–645 (2007).
Kifuji, Y., Hanzawa, H., Terasawa, Y., Ashutosh & Nishio, T. QTL analysis of black rot resistance in cabbage using newly developed EST–SNP markers. Euphytica 190, 289–295 (2013).
Tonu, N. N. et al. Comparison of positions of QTLs conferring resistance to Xanthomonas campestris pv. campestris in Brassica oleracea. Am. J. Plant Sci. 4, 11–20 (2013).
Saha, P., Kalia, P., Sonah, H. & Sharma, T. R. Molecular mapping of black rot resistance locus Xca1bo on chromosome 3 in Indian cauliflower (Brassica oleracea var. botrytis L.). Plant Breed. 133, 268–274 (2014).
Sharma, B. B., Pritam, K., Kumar, Y. D., Dinesh, S. & Raj, S. T. Genetics and molecular mapping of black rot resistance locus Xca1bc on chromosome B–7 in Ethiopian mustard (Brassica carinata Braun). PLoS ONE 11, e0152290 (2016).
Lee, J. et al. Genome–wide SNP identification and QTL mapping for black rot resistance in cabbage. BMC Plant Biol. 15, 32 (2015).
Kalia, P., Saha, P. & Ray, S. Development of RAPD and ISSR derived SCAR markers linked to Xca1bo, gene conferring resistance to black rot disease in cauliflower (Brassica oleracea var. botrytis L.). Euphytica 213, 232 (2017).
Sharma, B. B., Kalia, P., Singh, D. & Sharma, T. R. Introgression of black rot resistance from Brassica carinata to cauliflower (Brassica oleracea botrytis group) through embryo rescue. Front. Plant Sci. 8, 1255 (2017).
Henderson, M. P. The black–leg disease of cabbage caused by Phoma lingam (Tode) Desmaz. Phytopathology 8, 379–431 (1918).
Hall, R. Epidemiology of blackleg of oilseed rape. Can. J. Plant Pathol. 14, 46–55 (1992).
Sivasithamparam, K., Barbetti, M. J. & Li, H. Recurring challenges from a necrotrophic fungal plant pathogen: a case study with Leptosphaeria maculans (causal agent of blackleg disease in brassicas) in Western Australia. Ann. Bot. 96, 363–377 (2005).
Fitt, B. D. L., Brun, H., Barbetti, M. J. & Rimmer, S. R. World–wide importance of phoma stem canker (Leptosphaeria maculans, and L. biglobosa) on oilseed rape (Brassica napus). Eur. J. Plant Pathol. 114, 3–15 (2006).
Mengistu, A., Rimmer, S. R., Koch, E. & Williams, P. H. Pathogenicity grouping of isolates of Leptosphaeria maculans on Brassica napus cultivars and their disease reaction profiles on rapid–cycling brassicas. Plant Dis. 75, 1279 (1991).
Williams, R. H. & Fitt, B. D. L. Differentiating A and B groups of Leptosphaeria maculans, causal agent of stem canker (blackleg) of oilseed rape. Plant Pathol. 48, 161–175 (1999).
Howlett, B. J., Idnurm, A. & Pedras, M. S. Leptosphaeria maculans, the causal agent of blackleg disease of Brassicas. Fungal Genet. Biol. 33, 1–14 (2001).
Balesdent, M. H. et al. Analysis of Leptosphaeria maculans race structure in a worldwide collection of isolates. Phytopathology 95, 1061 (2005).
Ferreira, M. E., Rimmer, S. R., Williams, P. H. & Osborn, T. C. Mapping loci controlling Brassica napus resistance to Leptosphaeria maculans under different screening conditions. Phytopathology 85, 213–217 (1995).
Dion, Y., Gugel, R. K., Rakow, G. F., Seguinswartz, G. & Landry, B. S. RFLP mapping of resistance to the blackleg disease [causal agent, Leptosphaeria maculans (Eesm.) Ces. et De not.] in canola (Brassica napus L.). Theor. Appl. Genet. 91, 1190–1194 (1995).
Mayerhofer, R., Good, A. G., Bansal, V. K., Thiagarajah, M. R. & Stringam, G. R. Molecular mapping of resistance to Leptosphaeria maculans in Australian cultivars of Brassica napus. Genome 40, 294–301 (1997).
Mayerhofer, R. et al. Complexities of chromosome landing in a highly duplicated genome: toward map–based cloning of a gene controlling blackleg resistance in Brassica napus. Genetics 171, 1977–1988 (2005).
Delourme, R. et al. A cluster of major specific resistance genes to Leptosphaeria maculans in Brassica napus. Phytopathology 94, 578–583 (2004).
Long, Y. et al. Identification of two blackleg resistance genes and fine mapping of one of these two genes in a Brassica napus canola cultivar ‘surpass 400’. Theor. Appl. Genet. 122, 1223–1231 (2011).
Jestin, C. et al. Association mapping of quantitative resistance for Leptosphaeria maculans in oilseed rape (Brassica napus L.). Mol. Breed. 27, 271–287 (2011).
Raman, R. et al. Molecular mapping of qualitative and quantitative loci for resistance to Leptosphaeria maculans causing blackleg disease in canola (Brassica napus L.). Theor. Appl. Genet. 125, 405 (2012).
Tollenaere, R. et al. Identification and characterization of candidate Rlm4 blackleg resistance genes in Brassica napus using next-generation sequencing. Plant Biotechnol. J. 10, 709–715 (2012).
Yu, F., Gugel, R. K., Sharpe, A. G. & Rimmer, S. R. Introgression of Brassica rapa subsp. sylvestris blackleg resistance into B. napus. Mol. Breed. 30, 1495–1506 (2012).
Yu, F., Lydiate, D. J. & Rimmer, S. R. Identification of two novel genes for blackleg resistance in Brassica napus. Theor. Appl. Genet. 110, 969–979 (2005).
Yu, F., Lydiate, D. J. & Rimmer, S. R. Identification and mapping of a third blackleg resistance locus in Brassica napus derived from B. rapa subsp. sylvestris. Genome 51, 64–72 (2008).
Larkan, N. J. et al. The Brassica napus blackleg resistance gene LepR3 encodes a receptor–like protein triggered by the Leptosphaeria maculans effector AVRLM1. N. Phytol. 197, 595–605 (2013).
Larkan, N. J., Lydiate, D. J., Yu, F., Rimmer, S. R. & Borhan, M. H. Co–localisation of the blackleg resistance genes Rlm2 and LepR3 on Brassica napus chromosome A10. BMC Plant Biol. 14, 1–9 (2014).
Larkan, N. J., Ma, L. & Borhan, M. H. The Brassica napus receptor–like protein Rlm2 is encoded by a second allele of the LepR3/Rlm2 blackleg resistance locus. Plant Biotechnol. J. 13, 983–992 (2015).
Larkan, N. J. et al. The Brassica napus Wall-Associated Kinase-Like (WAKL) gene Rlm9 provides race-specific blackleg resistance. Preprint at https://doi.org/10.1101/815845 (2019).
Huang, Y. J. et al. Identification of environmentally stable QTL for resistance against Leptosphaeria maculans in oilseed rape (Brassica napus). Theor. Appl. Genet. 129, 169–180 (2016).
Larkan, N. J. et al. studies suggest a role for cysteine–rich protein kinase genes in quantitative resistance to blackleg disease in Brassica napus. BMC Plant Biol. 16, 183 (2016).
Brun, H. et al. Quantitative resistance increases the durability of qualitative resistance to Leptosphaeria maculans in Brassica napus. N. Phytol. 185, 285–299 (2010).
Purdy, L. H. Sclerotinia sclerotiorum: history, diseases and symptomatology, host range, geographic distribution, and impact. Phytopathology 69, 875–880 (1979).
Libert, M. A. Plante crytogamicae arduennae (Exsiccati) no. 326. Published by the author (1837).
Derbyshire, M. C. & Denton–Giles, M. The control of sclerotinia stem rot on oilseed rape (Brassica napus): current practices and future opportunities. Plant Pathol. 65, 859–877 (2016).
Bolton, M. D., Thomma, B. P. & Nelson, B. D. Sclerotinia sclerotiorum (Lib.) de Bary: biology and molecular traits of a cosmopolitan pathogen. Mol. Plant Pathol. 7, 1–16 (2006).
Ge, X. T. et al. Delineation of Sclerotinia sclerotiorum, pathotypes using differential resistance responses on Brassica napus, and B. juncea, genotypes enables identification of resistance to prevailing pathotypes. Field Crop Res. 127, 248–258 (2012).
Barbetti, M. J., Banga, S. S. & Salisbury, P. A. Challenges for crop production and management from pathogen biodiversity and diseases under current and future climate scenarios–case study with oilseed Brassicas. Field Crop. Res. 127, 225–240 (2012).
Zhao, J. & Meng, J. Genetic analysis of loci associated with partial resistance to Sclerotinia sclerotiorum in rapeseed (Brassica napus L.). Theor. Appl. Genet. 106, 759–764 (2003).
Zhao, J. et al. Quantitative trait loci for resistance to Sclerotinia sclerotiorum and its association with a homeologous non–reciprocal transposition in Brassica napus L. Theor. Appl. Genet. 112, 509–516 (2006).
Yin, X. et al. Mapping of QTLs detected in a Brassica napus DH population for resistance to Sclerotinia sclerotiorum in multiple environments. Euphytica 173, 25–35 (2010).
Wu, J. et al. Identification of QTLs for resistance to Sclerotinia stem rot and BnaC.IGMT5.a as a candidate gene of the major resistant QTL SRC6 in Brassica napus. PLoS ONE 8, e67740 (2013).
Fomeju, B. F., Falentin, C., Lassalle, G., Manzanares–Dauleux, M. J. & Delourme, R. Homoeologous duplicated regions are involved in quantitative resistance of Brassica napus to stem canker. BMC Genomics 15, 498 (2014).
Wei, L. et al. Genome-wide association analysis and differential expression analysis of resistance to sclerotinia stem rot in Brassica napus. Plant Biotechnol. J. 14, 1368 (2015).
Wu, J. et al. Genome-wide association study identifies new loci for resistance to sclerotinia stem rot in Brassica napus. Front. Plant Sci. 7, 1418 (2016).
Gyawali, S. et al. Microsatellite markers used for genome–wide association mapping of partial resistance to Sclerotinia sclerotiorum in a world collection of Brassica napus. Mol. Breed. 36, 1–13 (2016).
Mei, J. et al. Identification of resistant sources against Sclerotinia sclerotiorum in Brassica species with emphasis on B. oleracea. Euphytica 177, 393–399 (2011).
Mei, J. et al. Identification of genomic regions involved in resistance against Sclerotinia sclerotiorum from wild Brassica oleracea. Theor. Appl. Genet. 126, 549–556 (2013).
Mei, J. et al. Transfer of sclerotinia resistance from wild relative of Brassica oleracea, into Brassica napus, using a hexaploidy step. Theor. Appl. Genet. 128, 1–6 (2015).
Gaetán, S. A. Occurrence of Fusarium wilt on canola caused by Fusarium oxysporum f. sp. conglutinans in Argentina. Plant Dis. 89, 432–432 (2005).
Michiels, C. B. & Rep, M. Pathogen profile update: Fusarium oxysporum. Mol. Plant Pathol. 10, 311–324 (2009).
Smith, E. F. The fungus infection of agricultural soils in the United States. Sci. Am. Sup. 48, 19981–19982 (1899).
Bosland, P. W. & Williams, P. H. Pathogenicity of geographic isolates of Fusarium oxysporum from crucifers on a differential set of crucifer seedlings. Phytopathology 123, 63–68 (1988).
Liu, X. et al. Genetic diversity, virulence, race profiling and comparative genomic analysis of the Fusarium oxysporum f. sp. conglutinans strains infecting cabbages in China. Front. Microbiol. 10, 1373 (2019).
Armstrong, G. M. Formae Speciales and Races of Fusarium oxysporum Causing Wilt Diseases (Pennsylvania State University Press, University Park, PA, 1981).
Ramirez–Villupadua, J., Endo, R. M., Bosland, P. & Wiliams, P. H. A new race of Fusarium oxysporum f. sp. conglutinans that attacks cabbage with type A resistance. Plant Dis. 69, 612–613 (1985).
Morrison, R. H., Mengistu, A. & Williams, P. H. First report of race 2 of cabbage yellows caused by Fusarium oxysporum f. sp. conglutinans in Texas. Plant Dis. 78, 641 (1994).
Bosland, P. W., Williams, P. H. & Morrison, R. H. Influence of soil temperature on the expression of yellows and wilt of crucifers by Fusarium oxysporum. Plant Dis. 72, 777–780 (1988).
Walker, J. C. Inheritance of Fusarium resistance in cabbage. J. Agric. Res. 40, 721–745 (1930).
Blank, L. M. Fusarium resistance in Wisconsin All Seasons cabbage. J. Agric. Res. 55, 497–510 (1937).
Pu, Z. et al. Genetic mapping of a Fusarium wilt resistance gene in Brassica oleracea. Mol. Breed. 30, 809–818 (2012).
Lv, H. et al. Development of InDel markers linked to Fusarium wilt resistance in cabbage. Mol. Breed. 32, 961–967 (2013).
Lv, H. et al. Mapping and analysis of a novel candidate Fusarium wilt resistance gene FOC1 in Brassica oleracea. BMC Genomics 15, 1094 (2014).
Shimizu, M. et al. Map–based cloning of a candidate gene conferring Fusarium yellows resistance in Brassica oleracea. Theor. Appl. Genet. 128, 119–130 (2015).
Lv, H. et al. Breeding of cabbage (Brassica oleracea L. var. capitata) with Fusarium wilt resistance based on microspore culture and marker–assisted selection. Euphytica 200, 465–473 (2014).
Natti, J. J., Dickson, M. K., Atkin, D. & Resistance, D. of Brassica oleracea varieties to downy mildew. Phytopathology 51, 144–147 (1967).
Göker, M., Voglmayr, H., Riethmüller, A., Wei, M. & Oberwinkler, F. Taxonomic aspects of Peronosporaceae inferred from Bayesian molecular phylogenetics. Can. J. Bot. 81, 672–683 (2003).
Yuen, J. E. Resistance to Peronospora parasitica in Chinese cabbage. Plant Dis. 75, 10 (1991).
Vicente, J. G., Gunn, N. D., Bailey, L., Dac, P. & Holub, E. B. Genetics of resistance to downy mildew in Brassica oleracea and breeding towards durable disease control for UK vegetable production. Plant Pathol. 61, 600–609 (2012).
Jensen, B. D., Hockenhull, J. & Munk, L. Seedling and adult plant resistance to downy mildew (Peronospora parasitica) in cauliflower (Brassica oleracea var. botrytis). Plant Pathol. 48, 604–612 (1999).
Agnola, B. et al. Evidence that a leaf–disk test allows assessment of isolate–specific resistance in Brassica oleracea crops against downy mildew (Peronospora parasitica). Eur. J. Plant Pathol. 109, 471–478 (2003).
Göker, M., Voglmayr, H. & Oberwinkler, F. Species delimitation in downy mildews: the case of Hyaloperonospora in the light of nuclear ribosomal ITS and LSU sequences. Mycol. Res. 113, 308–325 (2009).
Coelho, P. S., Vicente, J. G., Monteiro, A. A. & Holub, E. B. Pathotypic diversity of Hyaloperonospora brassicae collected from Brassica oleracea. Eur. J. Plant Pathol. 134, 763–771 (2012).
Giovannelli, J. L., Farnham, M. W. & Wang, M. Development of sequence characterized amplified region markers linked to downy mildew resistance in broccoli. J. Am. Soc. Hortic. Sci. 127, 597–601 (2002).
Gao, M. et al. High–density Brassica oleracea, linkage map: identification of useful new linkages. Theor. Appl. Genet. 115, 277–287 (2007).
Coelho, P. S. & Monteiro, A. A. Expression of resistance to downy mildew at cotyledon and adult plant stages in Brassica oleracea L. Euphytica 133, 279–284 (2003).
Farinhó, M., Coelho, P., Monteiro, A. & Leitão, J. SCAR and CAPS markers flanking the Brassica oleracea L. Pp523 downy mildew resistance locus demarcate a genomic region syntenic to the top arm end of Arabidopsis thaliana L. chromosome 1. Euphytica 157, 215–221 (2007).
Carlier, J. D., Alabaça, C. A., Coelho, P. S., Monteiro, A. A. & Leitão, J. M. The downy mildew resistance locus Pp523 is located on chromosome C8 of Brassica oleracea L. Plant Breed. 131, 170–175 (2012).
Yu, S. et al. Genetic mapping and localization of a major QTL for seedling resistance to downy mildew in Chinese cabbage (Brassica rapa ssp. pekinensis). Mol. Breed. 23, 573–590 (2009).
Kim, S. et al. Identification of the BrRHP1 locus that confers resistance to downy mildew in Chinese cabbage (Brassica rapa ssp. pekinensis) and development of linked molecular markers. Theor. Appl. Genet. 123, 1183 (2011).
Yu, S. et al. Construction of a sequence–based bin map and mapping of QTLs for downy mildew resistance at four developmental stages in Chinese cabbage (Brassica rapa L. ssp. pekinensis). Mol. Breed. 36, 44 (2016).
Yu, S. C., Zhang, F. L., Zhao, X. Y., Yu, Y. J. & Zhang, D. S. Sequence–characterized amplified region and simple sequence repeat markers for identifying the major quantitative trait locus responsible for seedling resistance to downy mildew in Chinese cabbage (Brassica rapa ssp. pekinensis). Plant Breed. 130, 580–583 (2011).
Nikolaev, S. I. et al. The twilight of Heliozoa and rise of Rhizaria, an emerging supergroup of amoeboid eukaryotes. Proc. Natl Acad. Sci. USA 101, 8066–8071 (2004).
Karling, J. S. The Plasmodiophorales: including a complete host index, bibliography, and a description of diseases caused by species of this order. Mol. Biol. Evol. 22, 582–588 (1968).
Piao, Z., Ramchiary, N. & Yong, P. L. Genetics of clubroot resistance in Brassica species. J. Plant Growth Regul. 28, 252–264 (2009).
Williams, P. H. A system for the determination of races of Plasmodiophora brassicae that infect cabbage and rutabaga. Phytopathology 56, 624–626 (1966).
Buczacki, S. T. et al. Study of physiological specialization in Plasmodiophora brassicae: proposals for attempted rationalization through an international approach. Trans. Br. Mycol. Soc. 65, 295–303 (1975).
Crute, I. R., Gray, A. R., Crisp, P. & Buczacki, S. T. Variation in Plasmodiophora brassicae and resistance to clubroot disease in brassicas and allied crops – a critical review. Plant Breed. 50, 91–104 (1980).
Donald, E. C., Cross, S. J., Lawrence, J. M. & Porter, I. J. Pathotypes of Plasmodiophora brassicae, the cause of clubroot, in Australia. Ann. Appl. Biol. 148, 239–244 (2006).
Dixon, G. R. Variation in Plasmodiophora brassicae. Ann. Appl. Biol. 94, 278–280 (1980).
Voorrips, R. E. Plasmodiophora brassicae: aspects of pathogenesis and resistance in Brassica oleracea. Euphytica 83, 139–146 (1995).
Kageyama, K. & Asano, T. Life cycle of Plasmodiophora brassicae. J. Plant Growth Regul. 28, 203 (2009).
Matsumoto, E., Yasui, C., Ohi, M. & Tsukada, M. Linkage analysis of RFLP markers for clubroot resistance and pigmentation in Chinese cabbage (Brassica rapa ssp. pekinensis). Euphytica 104, 79 (1998).
Ueno, H. et al. Molecular characterization of the CRa gene conferring clubroot resistance in Brassica rapa. Plant Mol. Biol. 80, 621 (2012).
Piao, Z. Y., Deng, Y. Q., Choi, S. R., Park, Y. J. & Lim, Y. P. SCAR and CAPS mapping of CRb, a gene conferring resistance to Plasmodiophora brassicae in Chinese cabbage (Brassica rapa ssp. pekinensis). Theor. Appl. Genet. 108, 1458–1465 (2004).
Kato, T., Hatakeyama, K., Fukino, N. & Matsumoto, S. Fine mapping of the clubroot resistance gene CRb and development of a useful selectable marker in Brassica rapa. Breed. Sci. 63, 116–124 (2013).
Zhang, T. et al. Fine genetic and physical mapping of the CRb gene conferring resistance to clubroot disease in Brassica rapa. Mol. Breed. 34, 1173–1183 (2014).
Kato, T., Hatakeyama, K., Fukino, N. & Matsumoto, S. Identification of a clubroot resistance locus conferring resistance to a Plasmodiophora brassicae classified into pathotype group 3 in Chinese cabbage (Brassica rapa L.). Breed. Sci. 62, 282–287 (2012).
Hatakeyama, K. et al. The tandem repeated organization of NB–LRR genes in the clubroot–resistant CRb locus in Brassica rapa, L. Mol. Genet. Genomics 292, 397–405 (2017).
Suwabe, K. et al. Identification of two loci for resistance to clubroot (Plasmodiophora brassicae woronin) in Brassica rapa L. Theor. Appl. Genet. 107, 997–1002 (2003).
Suwabe, K. et al. Simple sequence repeat–based comparative genomics between Brassica rapa and Arabidopsis thaliana: the genetic origin of clubroot resistance. Genetics 173, 309–319 (2006).
Hirai, M. et al. A novel locus for clubroot resistance in Brassica rapa and ITS linkage markers. Theor. Appl. Genet. 108, 639–643 (2004).
Saito, M. et al. Fine mapping of the clubroot resistance gene, Crr3, in Brassica rapa. Theor. Appl. Genet. 114, 81 (2006).
Hatakeyama, K. et al. Identification and characterization of Crr1a, a gene for resistance to clubroot disease (Plasmodiophora brassicae woronin) in Brassica rapa L. PLoS ONE 8, e54745 (2013).
Chu, M. et al. Fine mapping of Rcr1 and analyses of its effect on transcriptome patterns during infection by Plasmodiophora brassicae. BMC Genomics 15, 1166 (2014).
Yu, F. et al. Identification of genome–wide variants and discovery of variants associated with Brassica rapa clubroot resistance gene Rcr1 through bulked segregant RNA sequencing. PLoS ONE 11, e0153218 (2016).
Huang, Z. et al. Fine mapping of a clubroot resistance gene in Chinese cabbage using SNP markers identified from bulked segregant RNA sequencing. Front. Plant Sci. 8, 1448 (2017).
Pang, W. et al. Identification and mapping of the clubroot resistance gene CRd in Chinese cabbage (Brassica rapa ssp. pekinensis). Front. Plant Sci. 9, 653 (2018).
Figdore, S. S., Ferreira, M. E., Slocum, M. K. & Williams, P. H. Association of RFLP markers with trait loci affecting clubroot resistance and morphological characters in Brassica oleracea L. Euphytica 69, 33–44 (1993).
Grandclément, C. & Thomas, G. Detection and analysis of QTLs based on RAPD markers for polygenic resistance to Plasmodiophora brassicae woron in Brassica oleracea L. Theor. Appl. Genet. 93, 86–90 (1996).
Rocherieux, J. et al. Isolate–specific and broad–spectrum QTLs are involved in the control of clubroot in Brassica oleracea. Theor. Appl. Genet. 108, 1555–1563 (2004).
Moriguchi, K., Kimizuka–Takagi, C., Ishii, K. & Nomura, K. A genetic map based on RAPD, RFLP, isozyme, morphological markers and QTL analysis for clubroot resistance in Brassica oleracea. Breed. Sci. 49, 257–265 (1999).
Nomura, K. et al. Evaluation of F2 and F3 plants introgressed with QTLs for clubroot resistance in cabbage developed by using SCAR markers. Plant Breed. 124, 371–375 (2005).
Voorrips, R. E., Jongerious, M. C. & Kanne, H. J. Mapping of two genes for resistance to clubroot (Plasmodiophora brassicae) in a population of doubled haploid lines of Brassica oleracea by means of RFLP and AFLP markers. Theor. Appl. Genet. 94, 75–82 (1997).
Nagaoka, T. et al. Identification of QTLs that control clubroot resistance in Brassica oleracea and comparative analysis of clubroot resistance genes between B. rapa and B. oleracea. Theor. Appl. Genet. 120, 1335–1346 (2010).
Lee, J. et al. Genotyping–by–sequencing map permits identification of clubroot resistance QTLs and revision of the reference genome assembly in cabbage (Brassica oleracea L.). DNA Res. 23, 29–41 (2016).
Landry, B. S. et al. A genetic map for Brassica oleracea based on RFLP markers detected with expressed DNA sequences and mapping of resistance genes to race 2 of Plasmodiophora brassicae (Woronin). Genome 35, 409–420 (1992).
Manzanares–Dauleux, M. J., Delourme, R., Baron, F. & Thomas, G. Mapping of one major gene and of QTLs involved in resistance to clubroot in Brassica napus. Theor. Appl. Genet. 101, 885–891 (2000).
Werner, S., Diederichsen, E., Frauen, M., Schondelmaier, J. & Jung, C. Genetic mapping of clubroot resistance genes in oilseed rape. Theor. Appl. Genet. 116, 363 (2008).
Fredua–Agyeman, R. & Rahman, H. Mapping of the clubroot disease resistance in spring Brassica napus canola introgressed from European winter canola cv. ‘mendel’. Euphytica 211, 1–13 (2016).
Hasan, M. J. & Rahman, H. Genetics and molecular mapping of resistance to Plasmodiophora brassicae pathotypes 2, 3, 5, 6, and 8 in rutabaga (Brassica napus var. napobrassica). Genome 59, 1 (2016).
Li, L. et al. A genome–wide association study reveals new loci for resistance to clubroot disease in Brassica napus. Front. Plant Sci. 7, 1483 (2016).
Hirani, A. H. et al. Transferring clubroot resistance from Chinese cabbage (Brassica rapa) to canola (B. napus). Can. J. Plant Pathol. 38, 9 (2016).
Liu, Y. et al. Screening of clubroot–resistant varieties and transfer of clubroot resistance genes to Brassica napus using distant hybridization. Breed. Sci. 68, 258–267 (2018).
Tonguc, M. & Griffiths, P. D. Development of black rot resistant interspecific hybrids between Brassica oleracea L. cultivars and brassica accession A19182, using embryo rescue. Euphytica 136, 313–318 (2004).
Snowdon, R. J., Winter, H., Diestel, A. & Sacristán, M. D. Development and characterisation of Brassica napus–sinapis arvensis addition lines exhibiting resistance to Leptosphaeria maculans. Theor. Appl. Genet. 101, 1008–1014 (2000).
Garg, H. et al. High level of resistance to Sclerotinia sclerotiorum, in introgression lines derived from hybridization between wild crucifers and the crop Brassica species B. napus, and B. juncea. Field Crop Res. 117, 51–58 (2010).
Diederichsen, E., Frauen, M., Linders, E. G. A., Hatakeyama, K. & Hirai, M. Status and perspectives of clubroot resistance breeding in crucifer crops. J. Plant Growth Regul. 28, 265–281 (2009).
Shimizu, M. et al. Identification of candidate genes for Fusarium yellows resistance in Chinese cabbage by differential expression analysis. Plant Mol. Biol. 85, 247–257 (2014).
Ma, C. et al. CRISPR/Cas9-mediated multiple gene editing in Brassica oleracea var. capitata using the endogenous tRNA-processing system. Hortic. Res. 6, 20 (2019).
Johnson, R. A critical analysis of durable resistance. Annu. Rev. Phytopathol. 36, 309–330 (1984).
Lindhout, P. The perspectives of polygenic resistance in breeding for durable disease resistance. Euphytica 124, 217–226 (2002).
Pilet, M. L., Delourme, R., Foisset, N. & Renard, M. Identification of loci contributing to quantitative field resistance to blackleg disease, causal agent Leptosphaeria maculans (Desm.) Ces. et De not. in winter rapeseed (Brassica napus L.). Theor. Appl. Genet. 96, 23–30 (1998).
Balesdent, M. H. et al. Genetic control and host range of avirulence towards Brassica napus cvs. Quinta and Jet Neuf in Leptosphaeria maculans. Phytopathology 91, 70–76 (2001).
Uloth, M. B. et al. New sources of resistance to Sclerotinia sclerotiorum, for crucifer crops. Field Crop Res. 154, 40–52 (2013).
Parra, L. et al. Rationalization of genes for resistance to Bremia lactucae in lettuce. Euphytica 210, 309–326 (2016).
Zhao, J. et al. Patterns of differential gene expression in Brassica napus cultivars infected with Sclerotinia sclerotiorum. Mol. Plant Pathol. 10, 635–649 (2009).
Steffen, R., Bernsdorff, F. E. M. & Cai, D. Members of the germin–like protein family in Brassica napus are candidates for the initiation of an oxidative burst that impedes pathogenesis of Sclerotinia sclerotiorum. J. Exp. Bot. 63, 5507–5519 (2012).
Wang, Z. et al. Defense to Sclerotinia sclerotiorum, in oilseed rape is associated with the sequential activations of salicylic acid signaling and jasmonic acid signaling. Plant Sci. 184, 75 (2012).
Wu, J. et al. Comparative transcriptomic analysis uncovers the complex genetic network for resistance to Sclerotinia sclerotiorum in Brassica napus. Sci. Rep. 6, 19007 (2016).
Liang, Y., Strelkov, S. E. & Kav, N. N. V. Oxalic acid–mediated stress responses in Brassica napus L. Proteomics 9, 3156 (2009).
Wen, L. et al. Using proteomic analysis to find the proteins involved in resistance against Sclerotinia sclerotiorum in adult Brassica napus. Eur. J. Plant Pathol. 137, 505–523 (2013).
Wang, Z. et al. Overexpression of Brassica napus MPK4 enhances resistance to Sclerotinia sclerotiorum in oilseed rape. Mol. Plant Microbe 22, 235–244 (2009).
Wang, Z. et al. Overexpression of BnWRKY33 in oilseed rape enhances resistance to Sclerotinia sclerotiorum. Mol. Plant Pathol. 15, 677–689 (2014).
Grison, R. et al. Field tolerance to fungal pathogens of Brassica napus constitutively expressing a chimeric chitinase gene. Nat. Biotechnol. 14, 643 (1996).
Liu, F. et al. Overexpression of barley oxalate oxidase gene induces partial leaf resistance to Sclerotinia sclerotiorum in transgenic oilseed rape. Plant Pathol. 64, 1407–1416 (2015).
Wang, Z. et al. Overexpression of OsPGIP2 confers Sclerotinia sclerotiorum resistance in Brassica napus through increased activation of defense mechanisms. J. Exp. Bot. 69, 3141–3155 (2018).
Bayer, P. E. et al. Variation in abundance of predicted resistance genes in the Brassica oleracea pangenome. Plant Biotechnol. J. 17, 789–800 (2019).
Jenner, C. E. et al. The cylindrical inclusion gene of Turnip mosaic virus encodes a pathogenic determinant to the Brassica resistance gene TuRB01. Mol. Plant Microbe 13, 1102 (2000).
Jenner, C. E., Wang, X. W., Ponz, F. & Walsh, J. A. A fitness cost for Turnip mosaic virus to overcome host resistance. Virus Res. 86, 1–6 (2002).
Jenner, C. E. et al. The dual role of the potyvirus P3 protein of Turnip mosaic virus as a symptom and avirulence determinant in brassicas. Mol. Plant Microbe 16, 777–784 (2003).
Walsh, J. A. et al. Different classes of resistance to Turnip mosaic virus in Brassica rapa. Eur. J. Plant Pathol. 08, 15–2 (2002).
Robaglia, C. & Caranta, C. Translation initiation factors: a weak link in plant RNA virus infection. Trends Plant Sci. 11, 40–45 (2006).
Beauchemin, C., Boutet, N. & Laliberté, J. F. Visualization of the interaction between the precursors of VPg, the viral protein linked to the genome of Turnip mosaic virus, and the translation eukaryotic initiation factor iso 4E in planta. J. Virol. 81, 775–782 (2007).
Yeam, I., Cavatorta, J. R., Ripoll, D. R., Kang, B. C. & Jahn, M. M. Functional dissection of naturally occurring amino acid substitutions in eIF4E that confers recessive potyvirus resistance in plants. Plant Cell 19, 2913–2928 (2007).
Rodríguez–Hernández, A. M. et al. Melon RNA interference (RNAi) lines silenced for Cm–eIF4E show broad virus resistance. Mol. Plant Pathol. 13, 755–763 (2012).
Qian, W. et al. Mapping and candidate–gene screening of the novel Turnip mosaic virus resistance gene retr02 in Chinese cabbage (Brassica rapa L.). Theor. Appl. Genet. 126, 179–188 (2013).
Lehmann, P., Jenner, C. E., Kozubek, E., Greenland, A. J. & Walsh, J. A. Coat protein–mediated resistance to Turnip mosaic virus in oilseed rape (Brassica napus). Mol. Breed. 11, 83–94 (2003).
Jan, F. J., Fagoaga, C., Pang, S. Z. & Gonsalves, D. A single chimeric transgene derived from two distinct viruses confers multi–virus resistance in transgenic plants through homology–dependent gene silencing. J. Gen. Virol. 81, 2103–2109 (2000).
Larkan, N. J., Yu, F., Lydiate, D. J., Rimmer, S. R. & Borhan, M. H. Single R gene introgression lines for accurate dissection of the Brassica - Leptosphaeria pathosystem. Front. Plant Sci. 7, 1771 (2016).
Slusarenko, A. J. & Schlaich, N. L. Downy mildew of Arabidopsis thaliana caused by Hyaloperonospora parasitica (formerly Peronospora parasitica). Mol. Plant Pathol. 4, 159–170 (2003).
Mcdowell, J. M., Williams, S. G., Funderburg, N. T., Eulgem, T. & Dangl, J. L. Genetic analysis of developmentally regulated resistance to downy mildew (Hyaloperonospora parasitica) in Arabidopsis thaliana. Mol. Plant Microbe 18, 1226–1234 (2005).
Coker, T. L. R., Volkan, C., Beynon, J. L. & Gifford, M. L. Spatial dissection of the Arabidopsis thaliana transcriptional response to downy mildew using fluorescence activated cell sorting. Front. Plant Sci. 6, 527 (2015).
Asai, S. et al. Expression profiling during Arabidopsis/downy mildew interaction reveals a highly-expressed effector that attenuates responses to salicylic acid. PLoS Pathog. 10, e1004443 (2014).
Liu, X. et al. Rapid introgression of the fusarium wilt resistance gene into an elite cabbage line through the combined application of a microspore culture, genome background analysis, and disease resistance–specific marker assisted foreground selection. Front. Plant Sci. 8, 354 (2017).
Chung, H. et al. Construction of a genetic map based on high–throughput SNP genotyping and genetic mapping of a TuMV resistance locus in Brassica rapa. Mol. Genet. Genomics 289, 149–160 (2014).
Li, Q. et al. Identification and mapping of a novel Turnip mosaic virus resistance gene TuRBCS01 in Chinese cabbage (Brassica rapa L.). Plant Breed. 134, 221–225 (2015).
Chèvre, A. M. et al. Characterization of Brassica nigra chromosomes and of blackleg resistance in B. napus–B. nigra addition lines. Plant Breed. 115, 113–118 (1996).
Chèvre, A. M. et al. Selection of stable Brassica napus–B. juncea recombinant lines resistant to blackleg (Leptosphaeria maculans) 1. Identification of molecular markers, chromosomal and genomic origin of the introgression. Theor. Appl. Genet. 95, 1104–1111 (1997).
Christianson, J. A., Rimmer, S. R., Good, A. G. & Lydiate, D. J. Mapping genes for resistance to Leptosphaeria maculans in Brassica juncea. Genome 49, 30 (2006).
Wei, D. et al. Quantitative trait loci analyses for resistance to Sclerotinia sclerotiorum, and flowering time in Brassica napus. Mol. Breed. 34, 1797–1804 (2014).
Li, J., Zhao, Z., Hayward, A., Cheng, H. & Fu, D. Integration analysis of quantitative trait loci for resistance to Sclerotinia sclerotiorum, in Brassica napus. Euphytica 205, 483–489 (2015).
Behla, R. et al. Identification of common QTL for resistance to Sclerotinia sclerotiorum, in three doubled haploid populations of Brassica napus (L.). Euphytica 213, 260 (2017).
Farinhó, M. et al. Mapping of a locus for adult plant resistance to downy mildew in broccoli (Brassica oleracea var. italica). Theor. Appl. Genet. 109, 1392–1398 (2004).
Yu, F. et al. Genotyping–by–sequencing reveals three QTL for clubroot resistance to six pathotypes of Plasmodiophora brassicae in Brassica rapa. Sci. Rep. 7, 4516 (2017).
Peng, L. et al. Identification of quantitative trait loci for clubroot resistance in Brassica oleracea with the use of Brassica SNP microarray. Front. Plant Sci. 9, 822 (2018).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (31701927), Central Public-Interest Scientific Institution Basal Research Fund (Y2020PT01; Y2018YJ04), earmarked fund for the Modern Agro-Industry Technology Research System, China (CARS-23), and the Science and Technology Innovation Program of the Chinese Academy of Agricultural Sciences (CAAS-ASTIP-IVFCAAS). We are grateful for the pictures provided by Dr Xiaohui Cheng from the Oil Crops Research Institute, CAAS, Dr Qi Peng from Jiangsu Academy of Agricultural Sciences, and Dr Guoliang Li from IVFCAAS. We apologize to those whose work could not be cited due to space limitations.
Author information
Authors and Affiliations
Contributions
H.L. wrote the draft and created the figures and tables. Z.F., L.Y., Y.Z., and Y.W. critically reviewed and improved the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lv, H., Fang, Z., Yang, L. et al. An update on the arsenal: mining resistance genes for disease management of Brassica crops in the genomic era. Hortic Res 7, 34 (2020). https://doi.org/10.1038/s41438-020-0257-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41438-020-0257-9
This article is cited by
-
Differences in the transcriptional immune response to Albugo candida between white rust resistant and susceptible cultivars in Brassica rapa L.
Scientific Reports (2023)
-
Genetic analysis reveals three novel QTLs underpinning a butterfly egg-induced hypersensitive response-like cell death in Brassica rapa
BMC Plant Biology (2022)
-
Fine mapping and candidate gene analysis of CRA3.7 conferring clubroot resistance in Brassica rapa
Theoretical and Applied Genetics (2022)
-
Perspectives for integrated insect pest protection in oilseed rape breeding
Theoretical and Applied Genetics (2022)