Abstract
Phosphatidylserine synthase (PSS)-mediated phosphatidylserine (PS) synthesis is crucial for plant development. However, little is known about the contribution of PSS to Na+ homeostasis regulation and salt tolerance in plants. Here, we cloned the IbPSS1 gene, which encodes an ortholog of Arabidopsis AtPSS1, from sweet potato (Ipomoea batatas (L.) Lam.). The transient expression of IbPSS1 in Nicotiana benthamiana leaves increased PS abundance. We then established an efficient Agrobacterium rhizogenes-mediated in vivo root transgenic system for sweet potato. Overexpression of IbPSS1 through this system markedly decreased cellular Na+ accumulation in salinized transgenic roots (TRs) compared with adventitious roots. The overexpression of IbPSS1 enhanced salt-induced Na+/H+ antiport activity and increased plasma membrane (PM) Ca2+-permeable channel sensitivity to NaCl and H2O2 in the TRs. We confirmed the important role of IbPSS1 in improving salt tolerance in transgenic sweet potato lines obtained from an Agrobacterium tumefaciens-mediated transformation system. Similarly, compared with the wild-type (WT) plants, the transgenic lines presented decreased Na+ accumulation, enhanced Na+ exclusion, and increased PM Ca2+-permeable channel sensitivity to NaCl and H2O2 in the roots. Exogenous application of lysophosphatidylserine triggered similar shifts in Na+ accumulation and Na+ and Ca2+ fluxes in the salinized roots of WT. Overall, this study provides an efficient and reliable transgenic method for functional genomic studies of sweet potato. Our results revealed that IbPSS1 contributes to the salt tolerance of sweet potato by enabling Na+ homeostasis and Na+ exclusion in the roots, and the latter process is possibly controlled by PS reinforcing Ca2+ signaling in the roots.
Similar content being viewed by others
Introduction
High concentrations of NaCl in the soil disrupt plant growth, cellular K+/Na+ homeostasis, and metabolic processes and markedly decrease crop yield in irrigated lands1. The increase in Na+ extrusion from the roots, reduction in root-to-shoot Na+ translocation, compartmentalization of Na+ in the vacuole, and maintenance of appropriate cytoplasmic K+ abundance are essential for plant salt tolerance1. For example, many plants activate plasma membrane (PM) Na+/H+ antiporter-mediated Na+ exclusion to maintain Na+ homeostasis in the cytosol under saline conditions1,2. Furthermore, the conserved salt overly sensitive (SOS) pathway regulates PM Na+/H+ antiporter activation in plants; in this pathway, a salt-induced buildup of cytosolic Ca2+ ([Ca2+]cyt) is identified by SOS3/CBL4 calcium sensors, which bind to SOS2/CIPK24 to form a complex2. The SOS2–SOS3 complex ultimately phosphorylates SOS1/NHX7 (a PM Na+/H+ antiporter), which can export Na+ from the cell2. Salt stress rapidly triggers an apoplastic H2O2 burst, which promotes the mediation of Na+ homeostasis in various plant species3,4,5. Chemical inhibition or mutations of PM NADPH oxidase decrease salt tolerance by reducing the salt-induced Ca2+ influx across the PM and the subsequent increase in [Ca2+]cyt3,6. These findings suggest that H2O2 activates PM Ca2+-permeable channels to enhance the signaling of [Ca2+]cyt under saline conditions3,6.
Lipid remodeling plays a critical role in plant salt tolerance because lipids affect membrane integrity, fluidity and membrane protein activity and function7. Alterations in membrane lipid composition act as signaling agents that mediate salt defensive responses in plants1. Phosphatidic acid (PA), phosphoinositide derivatives, and sphingolipids are important regulators of [Ca2+]cyt signaling, ion homeostasis, and salt tolerance in plants8,9. Phosphoinositide-phospholipase C (OsPLC1) is required for the hydrolyzation of phosphatidylinositol-4-phosphate and the formation of a salt-induced [Ca2+]cyt signature, which determines Na+ accumulation in leaf blades and whole-plant tolerance in rice10. PA production is induced rapidly in response to salt stress and plays multiple roles in mediating salt tolerance, such as the regulation of auxin efflux transporters11, mediation of microtubule organization12, stimulation of MPK6 activity to activate Na+/H+ exchange across the PM, and coordination of cellular ROS and pH signaling13,14,15. Recently, glycosyl inositol phosphorylceramide (GIPC) sphingolipids in the PM have been identified as a cell surface Na+ receptor required for NaCl-induced [Ca2+]cyt increase, activation of PM Na+/H+ antiporters, and basal salt tolerance in the model plant species Arabidopsis thaliana16.
Phosphatidylserine (PS) is a negatively charged phospholipid that is synthesized in the luminal leaflet of the endoplasmic reticulum by phosphatidylserine synthase (PSS); it must be flipped into the inner leaflet of the PM to contribute to membrane structure and electrostatic interactions in animals and plants17. Similar to other anionic phospholipids, the PS in the PM contributes to electrostatic interactions with other functional proteins and plays essential regulatory roles in many biological processes in animals and plants17. In Drosophila, PSS-mediated PS production mediates cell growth, lipid storage, and mitochondrial function18. In A. thaliana, AtPSS1, which catalyzes PS production by calcium-dependent base-exchange-type reactions with other phospholipids, is required for the development of microspores, inflorescence meristems, leaves, and internodes19,20. In rice (Oryza sativa L.) plants, PSS is involved in the elongation of cells in the uppermost internode and leaf senescence regulation21,22,23. Experimental manipulation of PS abundance in the PM by knocking down or overexpressing the Arabidopsis AtPSS1 gene leads to the graded activation of the PM auxin signaling pathway, which is regulated by the small GTPase ROP624. Moreover, PS is essential for the proper targeting of ROP6 to the PM, suggesting the essential signaling roles of PS in the mediation of plant development24. Our previous work revealed that salt-induced increases in PS facilitate ion homeostasis and tissue tolerance to salt stress in sweet potato25, suggesting that PSS is an important candidate gene that can be used to improve crop salt tolerance. However, whether PSS regulates salt defensive responses and salt tolerance in plants remains to be elucidated.
Herein, we demonstrated that the overexpression of a sweet potato PSS, IbPSS1, which is an ortholog of Arabidopsis AtPSS1, promotes cellular Na+ homeostasis by activating PM Na+/H+ antiport activity in sweet potato roots. Moreover, we showed that IbPSS1 overexpression enhances PM Ca2+-permeable channel sensitivity to NaCl and H2O2. Taken together, the results show that IbPSS1 acts as a key regulator of Na+ homeostasis, Ca2+ signaling, and salt tolerance in sweet potato.
Results
Identification of IbPSS1 from sweet potato
A PSS gene was isolated from the sweet potato cultivar Xushu 29 (Xu 29). The ORF of this gene is 1272 bp in length and encodes a 423 aa polypeptide (molecular weight: 49.4 kDa) with a predicted pI of 9.12. Multiple sequence alignment of this PSS and its orthologs from different species showed that it is a calcium-dependent base-exchange-type PSS (Fig. 1a). Phylogenetic analysis of the orthologs of PSS from different plant species indicated that this PSS gene from sweet potato is closely related to its orthologs in tomato (SlPSS1;1 and SlPSS1;2) and Arabidopsis (AtPSS1) (Fig. 1b). Thus, we named this gene IbPSS1 (GenBank accession number MN857546).
a Multiple alignment of base-exchange-type PSS protein sequences from Homo sapiens (Hs), Cricetulus griseus (Cg), Ipomoea batatas (Ib) and Arabidopsis thaliana (At). The amino acid residues indicated are important for CgPSS1 catalytic activity (open triangles), free Ser binding/recognition (closed triangle), PSS1 regulation (open circles), and PSS production and/or stability (diamonds). The black boxes indicate conserved residues. The gray bars indicate hypothetical membrane-spanning domains. b Phylogenetic analysis of IbPSS1 and its orthologs from different plant species. PSS1 proteins from the different species are indicated as follows: AtPSS1, Arabidopsis thaliana; GmPSS1;1 and GmPSS1;2, Glycine max; OsPSS1, Oryza sativa; PtPSS1;1 and PtPSS1;2, Populus trichocarpa; SbPSS1, Sorghum bicolor; SlPSS1;1 and SlPSS1;2, Solanum lycopersicum; VvPSS1, Vitis vinifera; and ZmPSS1;1 and ZmPSS1;2, Zea mays. The numbers on the tree indicate bootstrap supports (values < 50% are not shown). The branch lengths were drawn to scale; the bar size indicates the number of amino acid substitutions per site. c The transient expression of IbPSS1 in Nicotiana benthamiana leaves increases PS abundance in the PM region. A typical image of one of three independent experiments is shown. Bar = 30 μm. e Quantification of C2LACT-GFP fluorescence intensity. The columns labeled with “***” indicate significant differences at P < 0.001. n > 100 PM regions from at least 40 cells in three independent experiments.
To confirm whether IbPSS1 contributes to PS synthesis, we transiently coexpressed IbPSS1 with a genetically encoded biosensor of PS (C2LACT-GFP) in Nicotiana benthamiana leaves. This biosensor was extensively validated as a calcium-independent-specific PS reporter17,24. Figure 1c shows that the green fluorescence of C2LACT-GFP was mainly distributed in the marginal region of epidermal cells with or without IbPSS1 expression, suggesting that PS is mainly localized in the PM. However, the relative fluorescence intensity in the PM region of IbPSS1 and C2LACT-GFP coexpressed cells was significantly higher than that of C2LACT-GFP and empty vector (EV) coinoculated cells (Fig. 1d). These results confirmed that IbPSS1 contributes to the synthesis of PS.
Establishment of an efficient root transgenic system in sweet potato
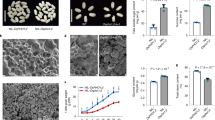
To rapidly determine the role of IbPSS1 in the mediation of Na+ homeostasis, we established an efficient A. rhizogenes-mediated in vivo root transgenic system in sweet potato26 (Fig. 2a). In our system, we introduced the expression vector containing two expression cassettes (the gene of interest + the DsRed marker) into A. rhizogenes strain K599, which was then injected into the basal part of stems of uniform cuttings of sweet potato cultivars. After growing in soil for 3 weeks, the rooted sweet potato seedlings were removed and washed, and many roots with obvious red fluorescence were visible around the injection site (Fig. 2a). The A. rhizogenes-mediated induction of transgenic roots (TRs) exhibited high efficiency in some sweet potato cultivars (DsRed only). For example, a 95% induction rate and an 80% induction rate were observed in the purple sweet potato Xuzishu 8 (Zi 8) and the yellow-flesh sweet potato Xu 29, respectively (Fig. S1).
a Pipeline of the A. rhizogenes-mediated in vivo root transgenic system. TRs: transgenic roots; ARs: adventitious roots. b Schematic of pUBI.U4::IbPSS1-CaMV35S::DsRed expression cassette, representative images of the TRs and ARs in the same seedling and the induction rate of TRs for this construct. c Relative expression level of IbPSS1 in the TRs and ARs with or without NaCl treatment (200 mM for 24 h). d Na+ accumulation in the elongation zones of the TRs and ARs. A typical image (n = 15 from three experiments) is shown. Bar = 0.3 mm. e Quantification of Na+ fluorescence intensity. The columns labeled with “***” indicate significant differences at P < 0.001.
To verify whether the root transgenic system reflects the influence of the genes of interest on Na+ homeostasis in root cells, we selected IbSOS1, which encodes a PM Na+/H+ antiporter in sweet potato, to confirm the effectiveness of the experimental system. We constructed a vector harboring two expression cassettes—one for IbSOS1 and one for the reporter gene DsRed—and then transferred the vector into Zi 8 as described above. After 3 weeks, many TRs with bright red fluorescence were obtained (Fig. S2A). The average transformation rate reached 85% for this vector (Fig. S2B). qRT-PCR data confirmed that IbSOS1 was successfully overexpressed in the TRs (Fig. S2C). We used CoroNa™ Green-based Na+ imaging and noninvasive microelectrode Na+ flux measurements to estimate the influence of IbSOS1 on root Na+ homeostasis in sweet potato5,27,28,29. After 24 h of NaCl stress (200 mM), bright CoroNa™ Green-specific fluorescence was recorded in the elongation zone in most adventitious roots (ARs) (Fig. S2D). However, Na+ accumulation in the elongation zone was markedly lower in the TRs than in the ARs (Fig. S2D, E). In addition, the Na+ efflux recorded in the elongation and mature zones of the TRs was significantly higher than that of the ARs (Figs S2F and G). Moreover, the Na+ efflux did not differ between the DsRed TRs (positive control) and the ARs under saline conditions (Fig. S3). These results clearly showed that our system can efficiently be used to determine the role of genes of interest in mediating Na+ homeostasis in sweet potato roots.
Cellular Na+ homeostasis is altered in IbPSS1-overexpressing sweet potato roots
We further tested whether IbPSS1 influences root Na+ homeostasis by using this root transgenic system. We constructed a vector with two expression cassettes—one for IbPSS1 and one for DsRed—and then transferred the vector into Xu 29. The average induction rate of the TRs reached 60% for this vector in four independent experiments (Fig. 2b). The relative expression level of IbPSS1 in the TRs was higher (3-fold) than that in the ARs under both control and saline conditions (Fig. 2c), suggesting that IbPSS1 was successfully overexpressed in sweet potato roots. However, the IbPSS1 transcript level did not increase in the ARs or TRs in response to NaCl treatment (Fig. 2c), suggesting that this gene was not upregulated by NaCl in the roots of this cultivar. After 24 h of NaCl stress (200 mM), we observed an obviously high accumulation of Na+, as indicated by CoroNaTM Green fluorescence, in the elongation zone of the ARs compared with the TRs (Fig. 2d). In addition, the relative Na+ fluorescence was reduced by 51% in the IbPSS1-overexpressing roots compared with the ARs (Fig. 2e). These results suggest that IbPSS1 overexpression contributes to the inhibition of Na+ accumulation in salinized sweet potato roots.
To explore how IbPSS1 inhibits root Na+ accumulation, we measured the salt-induced Na+ and H+ flux, which reflects the PM Na+/H+ exchange activity, in various plant species27,29 from different zones of the ARs and TRs by using noninvasive microtest technology (NMT). After 24 h of NaCl stress (200 mM), all tested root zones (elongation and mature zones) of the ARs and TRs showed evident Na+ efflux (Fig. 3a), which indicates that NaCl activated Na+ extrusion activity. Interestingly, a more pronounced Na+ efflux was observed in the TRs than in the ARs. The mean Na+ efflux rates in the two root zones of the TRs were 2.5- and 2.3-fold higher than those of the ARs, respectively (Fig. 3a, b). Correspondingly, evident H+ influx was recorded in the two root zones of the ARs and TRs (Fig. 3c), mirroring the Na+/H+ antiporter activity across the PMs of the root cells. Similar to the Na+ extrusion activity, the H+ influx was markedly higher in the two root zones of the TRs than in those of the ARs (Fig. 3d). No difference in the Na+ or H+ flux was observed between the ARs and TRs without NaCl stress (data not shown). These results indicate that IbPSS1 overexpression inhibits cellular Na+ accumulation by activating PM Na+/H+ antiporter activity in sweet potato roots. Interestingly, no significant difference in IbSOS1 transcript levels were found between the ARs and TRs under the control and saline conditions (Fig. S4). Moreover, IbPSS1 overexpression in sweet potato roots also inhibited K+ efflux induced by salt in the root elongation zone (Fig. S5).
a, c Net Na+ and H+ fluxes in the ARs and TRs measured by NMT. The steady-state Na+ and H+ fluxes were measured from the elongation (1.5, 2.0, and 3.0 mm from the tip) and mature root zones (10, 12, and 15 mm from the tip) after 24 h of NaCl treatment (200 mM). n > 20 from ten individual seedlings. The standard errors of the means are represented by the bars. b, d Mean net Na+ efflux and net H+ influx rates in (a) and (b), respectively. The columns labeled with “**” and “***” indicate significant differences between the AR and TR groups at P < 0.01 and P < 0.001, respectively.
IbPSS1 overexpression in sweet potato roots enhances PM Ca2+-permeable channel sensitivity to NaCl and H2O2
The salt-triggered Na+ extrusion to the apoplast in plant roots is generally accompanied by a Ca2+ influx across the PM, which contributes to the [Ca2+]cyt increase and PM Na+/H+ antiporter activation via the SOS pathway3,5,16. To examine whether IbPSS1 influences Ca2+ transport in salinized sweet potato roots, we determined the NaCl-induced transient Ca2+ flux kinetics. NaCl (150 mM) triggered an instantaneous and gradually decreasing Ca2+ efflux in the elongation zone of the ARs. This salt-induced Ca2+ efflux recovered to the control level after 20 min of NaCl treatment (Fig. 4a). We did not observe any evident increase in Ca2+ influx during salt stress in the ARs. However, NaCl induced a much lower Ca2+ efflux in the elongation zone of the TRs than in that of the ARs (Fig. 4a). In TRs, we observed a gradual decrease in NaCl-triggered Ca2+ efflux from 0 to 10 min after the addition of NaCl and evident transition of Ca2+ efflux to Ca2+ influx after 10 min of NaCl treatment (Fig. 4a). These results clearly show that IbPSS1 overexpression enhances PM Ca2+-permeable channel sensitivity to NaCl stress in sweet potato roots.
a Effects of NaCl stress (150 mM) on transient net Ca2+ flux kinetics in the elongation zone (2 mm from the tip) of the TRs and ARs. n = 9 from five individual seedlings. The columns in the insert show the mean rates of Ca2+ flux during the later period of NaCl stress (~10 min). “**” denotes significant differences at P < 0.01. b Effects of H2O2 (10 mM) on transient net Ca2+ flux kinetics in the elongation zones (2 mm from the tip) of the TRs and ARs. n = 12 from six individual seedlings. The columns in the insert show the mean Ca2+ flux rates during the H2O2 treatment (~20 min). “**” denotes significant differences at P < 0.01.
H2O2-activated PM Ca2+ channels contribute to PM Na+/H+ antiporter activation under saline conditions3,5. We further compared the Ca2+ flux kinetics induced by H2O2 in the elongation zone of ARs and TRs. H2O2 (10 mM) triggered an immediate Ca2+ influx in the ARs. The mean Ca2+ influx rate under H2O2 treatment reached 23 pmol cm−2 s−1 (Fig. 4b). However, H2O2 induced a more pronounced Ca2+ influx in the elongation zone of the TRs than in that of the ARs. The mean Ca2+ influx rate in the TRs reached 60 pmol cm−2 s−1 (Fig. 4b). Together, these results indicate that the PM Ca2+-permeable channel sensitivity to H2O2 is also reinforced in the TRs.
IbPSS1 overexpression enhances salt tolerance in transgenic sweet potato at the whole-plant level
To confirm whether IbPSS1 could enhance salt tolerance at the whole-plant level, A. tumefaciens-mediated transformation was used to introduce the pCaMV35S:IbPSS1 construct into embryogenic calli of sweet potato (cultivar Xu 29; Fig. 5a). Three regenerated transgenic lines (L11, L17, and L19) with the highest IbPSS1 transcript levels were used for further physiological characterization (Fig. 5b). NaCl (200 mM) was administered to uniformly rooted wild-type (WT) and IbPSS1-transgenic seedlings for 9 days. The results showed that, compared with the WT, the IbPSS1-transgenic lines exhibited obviously enhanced salt tolerance (Fig. 5c). Compared with the WT, the salinized transgenic lines had a markedly higher survival rate (Fig. 5d). Consistent with the salt-tolerant phenotype, the elongation zone of the salinized roots of the three transgenic lines had a significantly lower Na+ accumulation level than did that of the WT (Fig. 5e, f). In addition, the elongation and mature zones of the salinized roots of the three transgenic lines had significantly higher net Na+ efflux than did those of the WT (Fig. 5g). Taken together, these findings show that IbPSS1 overexpression improves salt tolerance at the whole-plant level by accelerating Na+ exclusion in the roots. Moreover, the PM Ca2+-permeable channel sensitivity to NaCl and H2O2 was enhanced in the roots of the IbPSS1-overexpressing seedlings (Fig. 5h, j).
a Schematic of the pCaMV35S::IbPSS1 expression cassette. b Relative expression level of IbPSS1 in the wild type (WT) and three independent transgenic lines. c Representative growth phenotype of WT and transgenic lines under control and saline conditions (200 mM NaCl for 4 and 9 days). d Quantification of the survival rate from four independent experiments. e Na+ accumulation in the root elongation zones of the WT and transgenic lines. A typical image (of 20) is shown. Bar = 0.3 mm. f Quantification of Na+ fluorescence intensity. g Net Na+ flux measured by NMT in the elongation and mature root zones of the WT and transgenic lines. n = 12 from six individual seedlings. The standard errors of the means are represented by the bars. h Effects of NaCl stress (150 mM) on the Ca2+ flux in the root elongation zone (2 mm from the tip) in the WT and transgenic lines. Ca2+ flux measurements were started after 10 min of NaCl stress. The inserted column shows the mean Ca2+ flux rate during the measurement period. n = 8 from four individual seedlings. i Effects of H2O2 (10 mM) on the transient net Ca2+ flux kinetics in the root elongation zones (2 mm from the tip) of the WT and transgenic lines. n = 8 from four individual seedlings. j Histogram showing the mean rate of Ca2+ flux before (~5 min) and after (15 min) the addition of H2O2. For all the histograms, the columns labeled with “*,” “**,” and “***” indicate significant differences at P < 0.05, P < 0.01, and P < 0.001, respectively, compared with the WT.
Exogenous application of lysophosphatidylserine (LPS) restores the PS content in and rescues the growth defects of AtPSS1 Arabidopsis mutants17. The effects of exogenous LPS on Na+ accumulation and Na+ flux after salinity stress were evaluated for their association with the IbPSS1-modulated cellular processes involving PS. The Na+ fluorescence in the root elongation zone of the LPS-pretreated seedlings (Xu 29, WT) was obviously lower than that in the nontreated Xu 29 seedlings after 48 h of NaCl stress (200 mM) (Fig. 6a, b). These findings suggest that the Na+ accumulation decreased. Consistent with Na+ accumulation, Na+ efflux increased by 65% in the root elongation zone of the LPS-pretreated Xu 29 after 48 h of NaCl stress compared with that in the nontreated roots (Fig. 6c). Moreover, the mean Ca2+ influx rate triggered by NaCl stress (150 mM; the Ca2+ flux measurement was started after 10 min of NaCl treatment; Fig. 6d) and H2O2 (10 mM; the Ca2+ flux measurement was started immediately after H2O2 treatment; Fig. 6e) was reinforced more in the LPS-pretreated roots than in the nontreated roots. These results suggest that PS modulates PM Ca2+-permeable channel sensitivity to NaCl and H2O2.
Sweet potato (WT, Xu 29) roots were pretreated with 50 μM LPS for 24 h and then subjected to NaCl stress (200 mM) for another 48 h. a Effects of LPS on cellular Na+ accumulation. Na+ accumulation in the root elongation zone of LPS-pretreated or nonpretreated seedlings. A typical image (of 22) is shown. Bar = 0.3 mm. b Quantification of Na+ fluorescence intensity. c Effects of LPS on the steady-state flux of Na+. The Na+ flux was measured from the root elongation zone after 48 h of NaCl stress. n = 16 from six individual seedlings. The SEs are represented by the bars. d Effects of LPS on NaCl-induced Ca2+ flux. The Ca2+ flux was measured from the root elongation zone (1.5, 2.5, and 3 mm from the tip) after 10 min of NaCl treatment. n = 12 from six individual seedlings. e Effects of LPS on H2O2-induced Ca2+ flux. The Ca2+ flux was measured from the root elongation zone (1.5, 2.5, and 3 mm from the tip) after the addition of 10 mM H2O2. n = 12 from six individual seedlings. For all the histograms, the columns labeled with “*,” “**,” and “***” indicate significant differences at P < 0.05, P < 0.01, and P < 0.001, respectively.
Discussion
Although the A. tumefaciens-mediated transformation method has been used for many sweet potato genotypes and although various genes have been characterized in detail30, current embryogenic callus-based transformation procedures of sweet potato still limit functional genomics in this important food crop species because they are time consuming and laborious. In the present study, a convenient and highly efficient method in which A. rhizogenes is used to induce the formation of TRs in intact sweet potato cuttings was established (Fig. 2a). This root transgenic system is similar to a previously reported strategy in woody plants26,31. Given that sweet potato is vegetatively propagated, an in vivo root transgenic system is not required for the sterilization of explants prior to A. rhizogenes infection26,31. The fluorescent marker DsRed used here helps to identify TRs from nontransgenic ARs of the same seedling31 (Fig. 2b). The nonoverlapping emission spectra between DsRed and green fluorescence-based physiological analyses enable the widespread application of this system through fluorescence microscopy or confocal microscopy, such as imaging Na+ accumulation by the use of CoroNaTM Green (this study; Figs. 2d, 5e, and 6a) or recoding cytosolic Ca2+ dynamics by the use of Ca2+ fluorescent probes and genetically encoded Ca2+ sensors (GCaMP3 and GCaMP6)32,33.
The cellular functions of genes of interest in different biological backgrounds could be determined within several days by combining this transgenic method with other physiological and molecular analyses (Fig. 2a). This strategy facilitates the determination of the biological function of genes of interest and accelerates the application of these genes to the genetic improvement of sweet potato through CRISPR-based technology. For example, by using the combination of NMT with this root transgenic system within 40 days, we revealed the novel role of IbPSS1 in stimulating PM Na+/H+ antiport activity in salinized sweet potato roots (Figs. 2 and 3). Importantly, the transgenic hairy roots induced by A. rhizogenes were anatomically similar to the primary roots and the ARs34. In addition, in the present study, the cellular processes influenced by the gene of interest (IbPSS1) did not differ between the A. rhizogenes- and A. tumefaciens-mediated transgenic systems (Figs. 2–5). Thus, our root transgenic system serves as a fast, convenient, and reliable tool for functional genomics in sweet potato. Given that each single TR represents an independent transformation event, this system can be a useful tool for testing the gene-editing efficiency among different CRISPR-Cas9/Cas12a/Cas12b variants or for developing regenerated and genome-edited germplasms of this autohexaploid crop species34,35,36.
PS plays an indispensable role in regulating plant development. AtPSS1-mediated alterations in PS contents in the PM are important regulators of small GTPase signaling during Arabidopsis root development24. Genetic disruption of PS biosynthesis decreases heterozygote fertility by inhibiting pollen maturation in Arabidopsis19. In the present study, we found that homologously overexpressed IbPSS1 facilitates Na+ efflux and H+ influx in salinized sweet potato roots (Fig. 3), indicating the enhancement of PM Na+/H+ antiporter activity27,28,29. This enhanced Na+/H+ antiport capability promotes Na+ homeostasis at the cellular level and increases salt tolerance at the whole-plant level (Fig. 5). Evidence from in vitro experiments showed that PS activates PM H+-ATPase activity in isolated PM vesicles of rice culture cells37. Moreover, our previous work showed that the exogenous addition of PS to sweet potato mesophyll cells stimulates H+ extrusion and PM H+-ATPase activity under saline conditions25. The potentially enhanced PM H+-ATPase activity, which generates more electrochemical H+ gradients to promote PM Na+/H+ antiporters1,2, may contribute to the enhanced Na+ extrusion and Na+ homeostasis in IbPSS1-overexpressing roots.
Salt stress increases [Ca2+]cyt concentration by activating PM Ca2+-permeable channels, and the expulsion of excess intracellular Na+ involves the Ca2+-related SOS signaling pathway2,16. H2O2, which is triggered by high-salinity stress, facilitates the activation of PM Ca2+-permeable channels and the formation of a specific [Ca2+]cyt signature, thereby contributing to the increase in PM Na+/H+ antiporter activity3,4,5,6. We revealed that the salt- and H2O2-induced Ca2+ flux patterns were altered in the IbPSS1-overexpressing roots or exogenous LPS-pretreated roots (Figs. 4, 5h–5j, 6d, e), suggesting that PM Ca2+-permeable channels are targets of PS under stressful conditions. IbSOS1 transcript levels exhibited no difference between the IbPSS1-overexpressing roots and the ARs (Fig. S4). Thus, the increased influx of Ca2+ across the PM possibly contributed to the activation of IbSOS1 at the posttranslational level via the SOS pathway, in which the phosphorylation of IbSOS1 may be enhanced by the reinforced Ca2+ signaling2. Given that H2O2 is an important signaling molecule involved in the activation of PM Ca2+-permeable channels in plants under various environmental stresses38, we hypothesize that the PSS-mediated variations in PS abundance in the PM play multiple signaling roles in the plant response to environmental changes. Although we did not observe salt-induced transcriptional changes in IbPSS1 in sweet potato roots (Fig. 2c), high-throughput lipidomic profiling has proven that the PS abundance in different plant tissues changes in response to various environmental stresses25,39,40.
In plant cells, PS is necessary and sufficient for establishing and maintaining the PM electrostatic signature, which may contribute to the specific targeting of proteins to the PM within the electrostatic region17. IbPSS1 overexpression may alter the abundance of PS within cells and its distribution in the PM, thereby shifting the electrostatic gradient and the targeting pattern of specific proteins in sweet potato root cells in response to salt and H2O2 stimuli17. Annexins are soluble proteins with multiple functions in plants41. Several lines of evidence suggest that annexins mediate the PS modulation of cellular responses to salt and H2O2. (1) Like vertebrate annexins, plant annexins contain a conserved PS binding site41. (2) Plant annexins (e.g., AtANN1 and AtANN4 from Arabidopsis) function as Ca2+-permeable channels (or cofactors of channels) that fine tune Ca2+ influx across the PM and the increase in [Ca2+]cyt in response to NaCl and H2O2 stimuli42,43,44. (3) Evidence from proteomic analyses of different cell fractions shows that NaCl stress stimulates the relocation of AtANN1 from the cytosol to root cell membranes45. (4) The ectopic expression of GhANN8b, which encodes an annexin from cotton (Gossypium spp.), stimulates salt-induced Ca2+ influx and Na+/H+ antiport activity across the PM in Arabidopsis roots46. Considering these lines of evidence and our findings, we suggest that annexins are the most probable targets downstream of IbPSS1-mediated PS synthesis and are involved in the regulation of Ca2+ influx and Na+/H+ antiporter capacity across the PM under saline conditions.
Na+ in the soil may be sensed extracellularly, intercellularly, or by transporters or channels at the PM in plant roots1. In Arabidopsis, external Na+ binds to GIPC sphingolipids outside of the PM and activates PM Ca2+-permeable channels to induce downstream responses to salinity, including an increase in [Ca2+]cyt and the activation of PM Na+/H+ antiporters via the SOS pathway16. Thus, GIPC sphingolipids have been identified as cell surface Na+ receptors required for tolerance to salinity in Arabidopsis16. Different Na+ sensing routes may exist and may be responsible for the initiation of multiple salt defensive responses in plants1. Considering that Na+ binds specifically to PS vesicles and that PS mainly localizes in the cytosolic leaflets of the PM17,24,47, PS may function as an intracellular Na+ sensor to modulate downstream responses to salinity, including annexin-mediated Ca2+ influx into the cell, the formation of a specific [Ca2+]cyt signature, the regulation of PM Na+/H+ antiporter activity, and PM H+-ATPase activity via a specific Ca2+-activated molecular pathway48. However, this hypothesis still needs further confirmation.
In conclusion, this study provides a fast, convenient, and reliable transgenic method for functional genomic studies of sweet potato. Our results showed that IbPSS1-mediated PS production is involved in the mediation of salt- and H2O2-induced Ca2+ transport across the PM and revealed that IbPSS1 positively regulates PM Na+/H+ antiporter activity and cellular Na+ homeostasis in the roots under saline conditions, thereby contributing to the enhancement of salt tolerance at the whole-plant level in sweet potato. To the best of our knowledge, this is the first study to investigate the influence of PSS on Na+ homeostasis and salt tolerance in plants. This study highlights the signaling role of PSS-mediated PS synthesis in mediating Na+ sensing and downstream defensive responses. However, further studies using antisense techniques are still needed to determine whether basic PS synthesis is required for the activation of PM Ca2+-permeable channels and PM Na+/H+ antiporters under saline conditions.
Materials and methods
Plant materials and growth conditions
The sweet potato varieties utilized in this study were obtained from the Xuzhou Institute of Agricultural Sciences in Jiangsu Xuhuai district, Xuzhou, Jiangsu, China. For stem cutting propagation, sweet potato seedlings were cultivated in pots (peat moss:loamy soil =1:1) placed inside a clean greenhouse whose temperature was 25 °C and whose photoperiod was 16 h. After enough seedlings were obtained, the stems were removed and used to induce TRs through A. rhizogenes.
Isolation of IbPSS1 and vector construction
Young leaves of Xu 29 were used to extract total RNA by using a TRIzol Reagent Kit. Total RNA was purified, and first-strand cDNA was synthesized by using HiScript II Q Select RT SuperMix (Vazyme). The conserved cloning primer (forward: 5′-ATGGAGCCTAATGGTCATAGAAGGAGTA-3′; reverse: 5′-TCATTGCCGCTTTCTCTTCATCAATTGC-3′) for IbPSS1 was designed according to the homologous gene from Ipomoea triloba (GenBank ID: XM_031245646.1), which is a wild relative of cultivated sweet potato. The cDNA was amplified by PCR using StarMAX DNA Polymerase (Takara, Japan), and the PCR product was subsequently purified and cloned into a pGEM-T vector for sequencing and alignment. IbSOS1 was also isolated from the sweet potato cultivar Xushu 22 (cloning primer: forward-5′-ATGACTTCCATGCTGGTGACG-3′; reverse-5′-CTAGCGAAAAGACAGTGTGCTTGG-3′).
The coding region of IbPSS1 or IbSOS1 and DsRed was inserted into a pCAMBIA0390 expression vector. Three constructs (pCaMV35S::DsRed, pUBI.U4::IbSOS1-CaMV35S::DsRed, and pUBI.U4::IbPSS1-CaMV35S::DsRed) were used for A. rhizogenes-mediated transformation. The coding region of IbPSS1 was inserted into a pCAMBIA1380 expression vector for A. tumefaciens-mediated transformation. Furthermore, two transient expression vectors expressing IbPSS1 and the PS sensor C2LACT-GFP were constructed using pRI201-AN as the backbone vector (pUBI.U4::IbPSS1 and pCaMV35S::C2LACT-GFP).
Phylogenetic analysis
The full-length amino acid sequences of IbPSS1 and its orthologs in other species were obtained from the UniProt and Phytozome databases. The amino acid sequences of the PSS protein were aligned by ClustalW, with the default settings. The conserved sequences were shaded at four levels by GeneDoc. A neighbor-joining phylogenetic tree of PSS proteins was constructed using the MEGA program (version 7.0.14), with 1000 bootstrap replications. The accession numbers of genes utilized in the present study are listed in Table S1.
PS abundance assay using the genetically encoded fluorescent sensor C2LACT-GFP
The pUBI.U4::IbPSS1 and pCaMV35S::C2LACT-GFP vectors were introduced into A. tumefaciens strain GV3101 by using the freeze–thaw method. A. tumefaciens was then resuspended in infiltration buffer (10 mM MES-KOH, 10 mM MgCl2, 200 μM acetosyringone (AS); pH 5.6) and diluted to an OD600 of 0.5. Equal volumes of A. tumefaciens solution containing the two constructs were mixed, and 1 mL of the mixture was infiltrated into the leaves of 4-week-old N. benthamiana plants by using a needleless syringe. EV instead of the pUBI.U4::IbPSS1 construct served as a control. The fluorescence image was visualized 4 days post-inoculation under an Olympus BX63 epifluorescence microscope using the same settings. Image-Pro Plus 6.0 software was then used to quantify the relative fluorescence intensity in the PM region.
A. rhizogenes-mediated transformation
Uniform sweet potato (Xu 29 and Zi 8) cuttings were collected for agro-infiltration using the following procedure. The three constructs were introduced into A. rhizogenes strain K599 and cultured in 5 mL of YPD liquid media consisting of 20 mg/L rifampicin and 50 mg/L kanamycin under shaking conditions (200 rpm) at 28 °C for 12–14 h. After the OD600 value reached 0.5, the cultures were centrifuged at 5000 rpm for 10 min at room temperature and then resuspended in infiltration buffer (10 mM MES-KOH, 10 mM MgCl2, 100 μM AS; pH 5.2). A 1.5 mL aliquot of the A. rhizogenes suspension was then injected into the stem of sweet potato cuttings by using a syringe with a needle. Afterward, the cuttings were transplanted into the soil and grown in a high-humidity environment. The transgenic positive plants were collected for further analysis after 3 weeks of culture in the greenhouse. The induction rate of the TRs was calculated as follows: (number of plants with TRs/total number of injected plants) ×100.
Generation of transgenic sweet potato overexpressing IbPSS1 via A. tumefaciens-mediated transformation
The pCaMV35S::IbPSS1 construct was introduced into A. tumefaciens EHA105, cultured overnight on a shaker (200 rpm, 28 °C) in YPD liquid media consisting of 12.5 mg/L hygromycin and 200 μM AS, and then transformed into embryogenic calli of sweet potato (Xu 29) using a reported previously A. tumefaciens-based method49. Transgenic calli were screened in media containing an appropriate concentration of hygromycin. Transgenic positive lines were verified by genomic DNA PCR analysis. qRT-PCR analysis was then performed to choose IbPSS1-overexpressing transgenic lines for further physiological analysis (IbPSS1 qRT-PCR primers: forward, 5′-CACCACCGTAGGATCTTTCAGG-3′; reverse, 5′-GTGATGATGCTCGCCAGTTTAT-3′). Three transgenic lines were selected and cultivated in the greenhouse for stem cutting propagation.
Salt treatment
Transgenic roots
Sweet potato seedlings with TRs (which fluoresced red under the excitation of a portable fluorescent lamp) were selected and immersed in hydroponic solution (1/4-strength Hoagland solution) for 24 h. Afterward, NaCl stress was started by adding the required amount of NaCl (final concentration of 200 mM) to the hydroponic solution. Before or after 24 h of NaCl stress, the ARs and TRs were collected to measure gene expression, intracellular Na+ accumulation, steady-state Na+ and H+ fluxes, and transient Ca2+ flux kinetics.
Transgenic plants
After obtaining enough seedlings, the shoots (with 4–5 mature leaves) of the WT and three transgenic lines were cut and cultivated in hydroponic solution to initiate AR growth for 10 days. Afterward, uniformly rooted seedlings were subjected to NaCl stress (200 mM) for another 9 days. The phenotype and survival rate were recorded at corresponding time points, after which the roots were collected to measure the intracellular Na+ accumulation, steady-state Na+ flux, and transient Ca2+ flux.
Exogenous LPS treatment
Rooted WT (Xu 29) seedlings were immersed in hydroponic solution in the absence or presence of 50 μM LPS (Avanti Polar Lipids, 850092 P) for 24 h. The required amount of NaCl (200 mM) was then added. The fine roots were collected to measure intracellular Na+ accumulation and Na+ flux after 48 h of NaCl treatment. The Ca2+ flux response to NaCl and H2O2 was also measured after 10 min of treatment.
Visualization of Na+ in root cells
CoroNaTM Green AM (Life Technologies), which is a Na+-specific fluorescent dye, was used to detect the accumulation of Na+ in sweet potato root cells5. The roots were collected after NaCl treatment and placed in a centrifuge tube containing fresh incubation solution (200 mM NaCl, 20 μM CoroNaTM Green AM (Life Technologies), and 0.02% pluronic acid) for 2 h. Afterward, they were washed several times with distilled water. An Olympus BX63 epifluorescence microscope was subsequently used to visualize intracellular Na+ accumulation (as indicated by the green fluorescence). All images were taken using the same settings and exposure times to allow direct comparisons. Image-Pro Plus 6.0 software was used to quantify the relative fluorescence intensity in the different root zones.
Analysis of transcript levels
Total RNA was extracted from the roots using TRIzol reagent (Takara Bio Inc., Japan). qRT-PCR was conducted as described previously50. Primers were designed to target IbSOS1 (forward: 5′-GGACTCCTCAGTGCTACT-3′; reverse: 5′-CATCGTTCATCAAGGATTCTC-3′) and IbPSS1 (forward: 5′-CACCACCGTAGGATCTTTCAGG-3′; reverse: 5′- GTGATGATGCTCGCCAGTTTAT-3′). The relative gene expression level was normalized to the reference gene (IbGAPDH; forward, 5′-ATACTGTGCACGGACAATGG-3′; reverse, 5′-TCAGCCCATGGAATCTCTTC-3′). The 2−∆∆Ct method was then used to calculate the relative expression levels.
Measurement of ion fluxes
The net ion fluxes were measured using the NMT system5,25,27,50. Ion-selective microelectrodes were constructed in accordance with our previously reported procedures27. The ion-selective microelectrodes were calibrated before recording the flux: (1) for Na+, 0.1, 0.5, and 1.0 mM NaCl (pH = 5.7) were used; (2) for H+, solutions with a pH of 5.0, 6.0, and 7.0 were used; and (3) for Ca2+ and K+, 0.1, 0.5 and 1.0 mM CaCl2 or KCl (150 mM NaCl or 10 mM H2O2 was added to the background solution; pH=5.7) was used. Microelectrodes with Nernstian slopes >52 mV per decade for Na+, H+, and K+ (26 mV per decade for Ca2+) were used. The ion flux was calculated in accordance with previously described methods5,25,27,50.
Steady-state Na+, H+ and Ca2+ flux measurements
Fine roots with 3–4 cm apices were collected from NaCl-stressed ARs and TRs and from WT and IbPSS1 transgenic lines. For Na+ and H+ flux recording, the roots were washed several times with the measuring solution (0.5 mM KCl, 0.1 mM MgCl2, 0.1 mM NaCl and 0.1 mM CaCl2; pH=5.7), followed by a 30 min equilibration in the measuring solution5,27. These root segments were then immobilized in the bottom of the measuring chamber that contained 10 mL of fresh measuring solution. Ion fluxes were recorded along the root axis: in the meristem zone (300, 400, and 500 μm from the tip, only for the IbSOS1 experiments), elongation zone (1.5, 2.0, and 2.5 mm from the tip), and mature zone (10, 12, and 15 mm from the tip). Recording was continuously conducted for 3–5 min at each measuring point in the three root zones.
For exogenous LPS experiments, the salt-induced changes in Na+ flux or the salt- and H2O2-induced alterations in Ca2+ flux in the root elongation zone were measured after 48 h or after 10 min of the corresponding treatments. The same measuring solution was used in this series of experiments.
Transient Ca2+ and K+ flux measurements
Fine roots with 3–4 cm apices were collected from ARs and TRs and from WT and IbPSS1 transgenic lines to measure the transient Ca2+ flux. The roots were fixed in the measuring solution (0.5 mM KCl, 0.1 mM MgCl2, 0.1 mM NaCl and 0.1 mM CaCl2; pH=5.7) for 30 min. The Ca2+ flux was recorded from the root elongation zone (2 mm from the tip) for 5 min before salt and H2O2 treatment. Afterward, the salt and H2O2 treatments were applied by adding NaCl (prepared with measuring solution, with a final concentration of 150 mM) and H2O2 (prepared with measuring solution, with a final concentration of 10 mM) to the measuring solution. The transient ion fluxes in the root elongation zone were measured for another 20 min. The NaCl-induced transient K+ flux kinetics in the ARs and TRs were also measured as described above.
Statistical analysis
The data were analyzed using ANOVA. “*”, “**”, and “***” indicate significant differences at P < 0.05, P < 0.01, and P < 0.001, respectively.
References
van Zelm, E., Zhang, Y. & Testerink, C. Salt tolerance mechanisms of plants. Annu. Rev. Plant Biol.71, 403–433 (2020).
Yang, Y. & Guo, Y. Elucidating the molecular mechanisms mediating plant salt stress responses. New Phytol.217, 523–539 (2018).
Sun, J. et al. H2O2 and cytosolic Ca2+ signals triggered by the PM H+-coupled transport system mediate K+/Na+ homeostasis in NaCl-stressed Populus euphratica cells. Plant Cell Environ.33, 943–958 (2010).
Niu, M. et al. Root respiratory burst oxidase homologue-dependent H2O2 production confers salt tolerance on a grafted cucumber by controlling Na+ exclusion and stomatal closure. J. Exp. Bot.69, 3465–3476 (2018).
Liu, Y. et al. Root-zone-specific sensitivity of K+- and Ca2+-permeable channels to H2O2 determines ion homeostasis in salinized diploid and hexaploidy Ipomoea trifida. J. Exp. Bot.70, 1389–1405 (2019).
Ma, L. et al. NADPH oxidase AtrbohD and AtrbohF function in ROS-dependent regulation of Na+/K+ homeostasis in Arabidopsis under salt stress. J. Exp. Bot.63, 305–317 (2012).
Higashi, Y. & Saito, K. Lipidomic studies of membrane glycerolipids in plant leaves under heat stress. Prog. Lipid Res.75, 100990 (2019).
Hong, Y. et al. Plant phospholipases D and C and their diverse functions in stress responses. Prog. Lipid Res.62, 55–74 (2016).
Ali, U., Li, H., Wang, X. & Guo, L. Emerging roles of sphingolipid signaling in plant response to biotic and abiotic stresses. Mol. Plant.11, 1328–1343 (2018).
Li, L. et al. A phosphoinositide-specific phospholipase C pathway elicits stress-induced Ca2+ signals and confers salt tolerance to rice. New Phytol.214, 1172–1187 (2017).
Wang, P. et al. Phosphatidic acid directly regulates PINOID-dependent phosphorylation and activation of the PIN-FORMED2 auxin efflux transporter in response to salt stress. Plant Cell.31, 250–271 (2019).
Zhang, Q. et al. Phosphatidic acid regulates microtubule organization by interacting with MAP65-1 in response to salt stress in Arabidopsis. Plant Cell.24, 4555–4576 (2012).
Yu, L. et al. Phosphatidic acid mediates salt stress response by regulation of MPK6 in Arabidopsis thaliana. New Phytol.188, 762–773 (2010).
Li, W. et al. Tissue-specific accumulation of pH-sensing phosphatidic acid determines plant stress tolerance. Nat. Plants5, 1012–1021 (2019).
Guo, L. et al. Cytosolic glyceraldehyde-3-phosphate dehydrogenases interact with phospholipase Dδ to transduce hydrogen peroxide signals in the Arabidopsis response to stress. Plant Cell.24, 2200–2212 (2012).
Jiang, Z. et al. Plant cell-surface GIPC sphingolipids sense salt to trigger Ca2+ influx. Nature572, 341–346 (2019).
Platre, M. P. et al. A combinatorial lipid code shapes the electrostatic landscape of plant endomembranes. Dev. Cell.45, 465–480 (2018).
Yang, X. et al. Phosphatidylserine synthase regulates cellular homeostasis through distinct metabolic mechanisms. PLoS Genet.5, e1008548 (2019).
Yamaoka, Y. et al. Phosphatidylserine synthase 1 is required for microspore development in Arabidopsis thaliana. Plant J.67, 648–661 (2011).
Liu, C. et al. Phosphatidylserine synthase 1 is required for inflorescence meristem and organ development in Arabidopsis. J. Integr. Plant Biol.55, 682–695 (2013).
Rani, M. H. et al. ES5 is involved in the regulation of phosphatidylserine synthesis and impacts on early senescence in rice (Oryza sativa L.). Plant Mol. Biol.102, 501–515 (2020).
Ma, J. et al. Phosphatidylserine synthase controls cell elongation especially in the uppermost internode in rice by regulation of exocytosis. PLoS ONE11, e0153119 (2016).
Zhu, L. et al. Identification and characterization of SHORTENED UPPERMOST INTERNODE 1, a gene negatively regulating uppermost internode elongation in rice. Plant Mol. Biol.77, 475–487 (2011).
Platre, M. P. et al. Developmental control of plant Rho GTPase nano-organization by the lipid phosphatidylserine. Science364, 57–62 (2019).
Yu, Y. et al. Involvement of phosphatidylserine and triacylglycerol in the response of sweet potato leaves to salt stress. Front Plant Sci.10, 1086 (2019).
Meng, D. et al. Development of an efficient root transgenic system for pigeon pea and its application to other important economically plants. Plant Biotechnol. J.17, 1804–1813 (2019).
Sun, J. et al. NaCl-induced alternations of cellular and tissue ion fluxes in roots of salt-resistant and salt sensitive poplar species. Plant Physiol.149, 1141–1153 (2009).
Cuin, T. A. et al. Assessing the role of root plasma membrane and tonoplast Na+/H+ exchangers in salinity tolerance in wheat: in planta quantification methods. Plant Cell Environ.34, 947–961 (2011).
Wu, H. et al. Root vacuolar Na+ sequestration but not exclusion from uptake correlates with barley salt tolerance. Plant J.100, 55–67 (2019).
Liu, Q. Improvement for agronomically important traits by gene engineering in sweet potato. Breed. Sci.67, 15–26 (2017).
Gomes, C., Dupas, A., Pagano, A., Grima-Pettenati, J. & Paiva, J. A. P. Hairy root transformation: A useful tool to explore gene function and expression in salix spp. recalcitrant to transformation. Front Plant Sci.10, 1427 (2019).
Matthus, E. et al. DORN1/P2K1 and purino-calcium signalling in plants: making waves with extracellular ATP. Ann. Bot.124, 1227–1242 (2020).
Pan, Y. et al. Dynamic interactions of plant CNGC subunits and calmodulins drive oscillatory Ca2+ channel activities. Dev. Cell.48, 710–725 (2019).
Ron, M. et al. Hairy root transformation using Agrobacterium rhizogenes as a tool for exploring cell type-specific gene expression and function using tomato as a model. Plant Physiol.166, 455–469 (2014).
Butler, N. M., Jansky, S. H. & Jiang, J. First generation genome editing in potato using hairy root transformation. Plant Biotechnol. J. https://doi.org/10.1111/pbi.13376. (2020)
Ming., M. et al. CRISPR-Cas12b enables efficient plant genome engineering. Nat. Plants6, 202–208 (2020).
Kasamo, K. Mechanism for the activation of plasma membrane H+-ATPase from rice (Oryza sativa L.) culture cells by molecular species of a phospholipid. Plant Physiol.93, 1049–1052 (1990).
Demidchik, V., Shabala, S., Isayenkov, S., Cuin, T. A. & Pottosin, I. Calcium transport across plant membranes: mechanisms and functions. New Phytol.220, 49–69 (2018).
Sarabia, L. D. et al. Comparative spatial lipidomics analysis reveals cellular lipid remodelling in different developmental zones of barley roots in response to salinity. Plant Cell Environ.43, 327–343 (2019).
Zhang, X., Xu, Y. & Huang, B. Lipidomic reprogramming associated with drought stress priming-enhanced heat tolerance in tall fescue (Festuca arundinacea). Plant Cell Environ.42, 947–958 (2019).
Laohavisit, A. & Davies, J. M. Annexins. New Phytol.89, 40–53 (2011).
Laohavisit, A. et al. Salinity-induced calcium signaling and root adaptation in Arabidopsis require the calcium regulatory protein annexin1. Plant Physiol.163, 253–262 (2013).
Richards, S. L. et al. Annexin 1 regulates the H2O2-induced calcium signature in Arabidopsis thaliana roots. Plant J.77, 136–145 (2014).
Ma, L. et al. The SOS2-SCaBP8 complex generates and fine-tunes an AtANN4-dependent calcium signature under salt stress. Dev. Cell.48, 697–709 (2019).
Lee, S. et al. Proteomic identification of annexins, calcium-dependent membrane binding protein that mediate osmotic stress and abscisic acid signal transduction in Arabidopsis. Plant Cell.16, 1378–1391 (2004).
Mu, C., Zhou, L., Shan, L., Li, F. & Li, Z. Phosphatase GhDsPTP3a interacts with annexin protein GhANN8b to reversely regulate salt tolerance in cotton (Gossypium spp.). New Phytol.223, 1856–1872 (2019).
Kurland, R., Newton, C., Nir, S. & Papahadjopoulos, D. Specificity of Na+ binding to phosphatidylserine vesicles from a 23Na NMR relaxation rate study. Biochim. Biophys. Acta.551, 137–147 (1979).
Yang, Z. et al. Calcium-activated 14-3-3 proteins as a molecular switch in salt stress tolerance. Nat. Commun.10, 1199 (2019).
Kim, S. H., Ahn, Y. O., Ahn, M. J., Lee, H. S. & Kwak, S. S. Down-regulation of β-carotene hydroxylase increases β-carotene and total carotenoids enhancing salt stress tolerance in transgenic cultured cells of sweet potato. Phytochemistry74, 69–78 (2012).
Yu, Y. et al. Melatonin stimulated triacylglycerol breakdown and energy turnover under salinity stress contributes to the maintenance of plasma membrane H+-ATPase activity and K+/Na+ homeostasis in sweet potato. Front Plant Sci.9, 256 (2018).
Acknowledgements
This work was supported by the National Key R & D Program of China (2018YFD1000704, 2018YFD1000700), the earmarked fund for the China Agriculture Research System (CARS-10-B03), the National Natural Science Foundation of China (31871684, 31701483), the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD), the Jiangsu Province Agricultural Science and Technology Innovation Fund (CX18, 3011) and the Graduate Student Scientific Research Innovation Projects in Jiangsu Province (KYCX171612).
Author information
Authors and Affiliations
Contributions
J.S. and Z.Y.L. designed experiments; Y.C.Y., Y.X., X.F.B., L.Z., and Z.Y.P. performed most of the experiments; M.K., Q.H.C., Z.H.T., Q.L., and D.F.M. provided sweet potato materials and suggestions. J.S. wrote the paper.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yu, Y., Xuan, Y., Bian, X. et al. Overexpression of phosphatidylserine synthase IbPSS1 affords cellular Na+ homeostasis and salt tolerance by activating plasma membrane Na+/H+ antiport activity in sweet potato roots. Hortic Res 7, 131 (2020). https://doi.org/10.1038/s41438-020-00358-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41438-020-00358-1
This article is cited by
-
Genome-wide systematic survey and analysis of the RNA helicase gene family and their response to abiotic stress in sweetpotato
BMC Plant Biology (2024)
-
Salinity responses and tolerance mechanisms in underground vegetable crops: an integrative review
Planta (2022)
-
Agrobacterium rhizogenes-mediated hairy root transformation as an efficient system for gene function analysis in Litchi chinensis
Plant Methods (2021)