Abstract
Double digest restriction-site associated sequencing (ddRAD-seq) is a flexible and cost-effective strategy for providing in-depth insights into the genetic architecture of germplasm collections. Using this methodology, we investigated the genomic diversity of a panel of 288 diverse tomato (Solanum lycopersicum L.) accessions enriched in ‘da serbo’ (called ‘de penjar’ in Spain) long shelf life (LSL) materials (152 accessions) mostly originating from Italy and Spain. The rest of the materials originate from different countries and include landraces for fresh consumption, elite cultivars, heirlooms, and breeding lines. Apart from their LSL trait, ‘da serbo’ landraces are of remarkable interest for their resilience. We identified 32,799 high-quality SNPs, which were used for model ancestry population structure and non-parametric hierarchical clustering. Six genetic subgroups were revealed, clearly separating most ‘da serbo’ landraces, but also the Spanish germplasm, suggesting a subdivision of the population based on type and geographical provenance. Linkage disequilibrium (LD) in the collection decayed very rapidly within <5 kb. We then investigated SNPs showing contrasted minor frequency allele (MAF) in ‘da serbo’ materials, resulting in the identification of high frequencies in this germplasm of several mutations in genes related to stress tolerance and fruit maturation such as CTR1 and JAR1. Finally, a mini-core collection of 58 accessions encompassing most of the diversity was selected for further exploitation of key traits. Our findings suggest the presence of a genetic footprint of the ‘da serbo’ germplasm selected in the Mediterranean basin. Moreover, we provide novel insights on LSL ‘da serbo’ germplasm as a promising source of alleles for tolerance to stresses.
Similar content being viewed by others
Introduction
Tomato (Solanum lycopersicum L.) is the most economically important vegetable crop, being widely grown and appreciated for its organoleptic and nutritional properties. Its cultivation is spread on a global surface of 4.76 million hectares with a production of 182 million tons1. The domestication of tomato had its origins in the Andean region of South America, fitting a model in which a pre-domestication phase took place in the Andean region, with the domestication being completed in Mesoamerica2,3 and not so long before the discovery of America4. This process led to a series of morphological and physiological changes, the most important of which has been the increase of the size and weight of fruits and the variation of shape3. During the XVI century, tomato was introduced in Europe and mostly in Spain and Italy, where further domestication occurred, resulting in the development of a high diversity of local varieties. For this reason, the Mediterranean basin of Europe is recognized as a secondary center of diversification for tomato5,6.
The last century has been characterized by a boost of breeding activities aimed at introducing and selecting novel variation toward the development of various types of cultivars for different uses and destinations (e.g., fresh consumption or processing industry). The first tomato cultivars developed by growers through selection and propagation are known as heirloom varieties. Heirlooms are characterized by unique features related to their appearance (e.g. color, shape, etc.), flavor, and significance for specific local markets6,7. Similarly, landraces are traditional cultivars developed over time after adapting to natural and cultural environments. Although both have been selected and handed down by farmers, landraces are limited to specific ecogeographical habitats and therefore are adapted to certain pedoclimatic conditions, whereas heirlooms do not necessarily present this characteristic7,8. Furthermore, in several cases, heirlooms have been developed through cross-breeding activities toward the development of resilient cultivars adapted to low input farming7. Anyway, in crops with a high economic impact such as tomato, the release of numerous commercial true-breeding and hybrid varieties has become a common rule in the market. Generally, modern tomato varieties are uniform and highly productive, and are characterized by a reduced gene pool, having been bred for few traits of major interest for the growers and industry, with resulting drawbacks related to the lack of those quality traits positively perceived by consumers such as flavor9. The dynamic changes of the market, concerning the increasing attention to the environment and sustainability, have grown the interest in the ‘Zero’ concept in the food chain (e.g., zero kilometers, zero pesticides, etc.). At the same time, identity, ethical and cultural aspects attract the interest of customers who show greater appreciation for typical products and traditional cuisine. For this reason, in recent years there has been an increasing interest in local landraces, which, although not as productive as commercial cultivars and often vulnerable to major pests8, have a closer tie with the territory and local customs and are better adapted to the small-scale economies.
Among these, ‘da serbo’ (also called ‘de penjar’ or ‘de ramellet’ in Spain) tomato materials are typical long shelf life (LSL) tomatoes landraces that diversified in the Mediterranean region and which have been traditionally cultivated in Southern Italy and Spain. These LSL tomatoes are generally small, mostly round, although a wide range of shapes can be found, with thick cuticle, and different colors, from yellow to red with several shades. The extended shelf-life might have been the primary selection criteria10 and is associated with high resilience, manifested in the tolerance to different abiotic and biotic stresses, as well as to a high soluble solids content providing excellent organoleptic properties11. This syndrome of phenotypic characteristics of the LSL ‘da serbo’ landraces results in a clear differentiation from the standard fresh consumption tomatoes. The particular characteristics of the LSL ‘da serbo’ materials may be also used as a selective trait10,12, in fact, they have been selected in semiarid summer conditions with poor irrigation or rain-fed10. The greater attention to organic farming and sustainability in agriculture has sparked an increasing interest in the ‘da serbo’ LSL cultivars. The potential of these LSL materials and their diversity have not been extensively characterized and are not yet fully exploited in tomato breeding programs. Indeed, apart from the long shelf life and resilience characteristics, ‘da serbo’ LSL germplasm may represent a potential source of other traits of agricultural interest.
The diversity of local landraces and heirlooms of tomato has been investigated using PCR based fragment analysis or custom DNA arrays (Golden-Gate) involving from a few hundred markers13,14,15,16,17 to few thousand of markers (e.g., Illumina Infinium SolCAP)18,19,20. These studies revealed patterns of genetic diversity due to breeding activities, adaptation to the cultivation environment, market type and destination of use. Whole-genome sequencing and re-sequencing projects accelerated the development of high-throughput genotyping methods21. Double digest restriction-site associated DNA (ddRAD-seq) is a method based on the development of multiplexed libraries obtained through the enzymatic digestion of the whole genomic DNA followed by binding to specific adapters (reduced representation libraries). By reducing the portion of the sequenced genome, it generates a large set of SNP markers which can be used to infer very precisely the genetic diversity and population structure of large germplasm collections22. Moreover, the adoption of double restriction enables paired-end sequencing of identical loci across many samples ensuring higher accuracy in the mapping of reads with respect to genotyping by sequencing (GBS) or single RAD digestion (RAD-seq)23. The application of ddRAD-seq technology has been successful in many species including Solanaceae crops24,25; while, so far, its application to tomato for the investigation of germplasm collections has been limited. Indeed, to date, studies are addressed principally to ddRAD-seq in silico optimization protocols23.
A reference genome sequence of tomato has been available since 201226. The first version, obtained by a combination of short-read 454 and a Sanger paired-end read sequencing, consisted of approximately 900 Mb and 34,727 protein-coding gene models. Versions SL2.50 and SL3.0 have been widely used as a reference for SNP calling in various tomato population studies. Very recently, a high quality de novo assembly of the tomato reference genome (SL4.0), obtained through a combination of long-read sequencing methodologies and optical mapping, has been released27. The new version reduced the number of the unknown bases and unplaced contigs by around 2,000 and 30-fold, respectively. Moreover, 4,794 novel genes have been reported compared to the previous version. The new genes information has been implemented by the sequencing of 13 diverse tomato accessions (Shatz and Lippman unpublished, https://solgenomics.net/projects/tomato13/). These novel resources allow a better exploitation of the genomic information on germplasm resources.
In the present study, we analyzed the genome-wide diversity of a collection of 288 tomato accessions using ddRAD-seq genotyping aiming to provide biological knowledge for biodiversity-based breeding. The represented tomato accessions included a wide range of landraces for fresh consumption and LSL ‘da serbo’ retrieved from the Mediterranean basin, as well as representative heirlooms and elite cultivars grown across the world. Population structure and diversity within subpopulations have been inferred by combining Bayesian and non-parametric approaches. Then we investigated in depth the genomic diversity of ‘da serbo’ materials, identifying those alleles subjected to selective sweeps and putatively responsible for the syndrome of phenotypic characteristics of LSL types. The collection studied, as well as a mini-core collection selected for genomic diversity, represents diversified panels to be exploited for future association mapping studies toward the determination of genomic regions responsible of traits underlying resilience and amenable to breeding.
Materials and methods
Plant material and DNA isolation
A diverse set of 288 cultivated tomato (S. lycopersicum) accessions were assembled from the germplasm collections from the Universitat Politècnica de València (UPV; Spain), the Research Center for Vegetable and Ornamental Crops (CREA; Italy), and the Tomato Genetics Resource Center (TGRC; USA). The germplasm panel included materials originating from 27 different countries of Europe, Asia, Africa and America (Fig. 1) and included 152 landraces (77 of the ‘da serbo’ typology and 75 of the standard typology for fresh consumption), mainly retrieved from Spain and Italy, 46 heirloom varieties, 76 elite cultivars and 14 breeding lines. Information about the accessions, including variety name and country of origin is in Supplementary Table 1. Genomic DNA was extracted from leaves of each accession using the DNeasy® Plant Mini Kit (QIAGEN, Germany). DNA quality parameters as well as concentration were measured by absorbance values at 260 nm and 280 nm, using a UV-Vis spectrophotometer (ND-1000; NanoDrop, Thermo Scientific, Wilmington, DE, USA).
RADseq genotyping
In silico analysis on the tomato reference genome was used to select the best combination of two restriction enzymes and the best fragment size distribution to obtain the desired number of loci. Such analysis suggested that the MboI and SphI enzyme pair is capable of producing a higher number of fragments, as already reported in other plant species28. Hence, this combination was used to generate fragments within the 288 tomato samples. Genomic DNA of each cultivar was double digested and incubated at 37 °C for 16–20 h. Fragmented DNA was then purified with AMPureXP beads (Agencourt) and ligated to barcoded adapters. Samples were pooled on multiplexing batches and bead purified. For each pool, targeted fragments distribution was collected on a BluePippin instrument (Sage Science Inc.). The gel eluted fraction was amplified with oligo primers that introduced TruSeq indexes and subsequently bead purified. The resulting libraries were checked with both Qubit 2.0 Fluorometer (Invitrogen, Carlsbad, CA) and Bioanalyzer DNA assay (Agilent Technologies, Santa Clara, CA). Libraries were finally processed with Illumina cBot for cluster generation on the flowcell, following the manufacturer’s instructions and sequenced with V4 chemistry paired end 125 bp mode on HiSeq2500 instrument (Illumina, San Diego, CA).
SNP call, filtering and marker classification
Demultiplexing of raw Illumina reads was carried out using the process_radtags utility included in Stacks v2.029. The quality distribution plot was generated using FASTQ and filtered data were then aligned to reference tomato genome (versions SL3.0 and SL.4.0) using BWA-MEM with default parameters30. Only uniquely aligned reads (i.e., reads with a mapping quality > 4) were selected for downstream analysis. Genomic sequences related to S. lycopersicum (versions SL3.0 and SL4.0) were obtained from Sol Genomics (https://solgenomics.net). Detection of all covered loci from the aligned reads was done using the gstacks program included in Stacks v2.029, while filtering of detected loci was carried out using the included populations program with the option –r = 0.75, in order to retain only loci that are represented in at least 75% of the population. In addition, polymorphic homozygous markers were identified among the accessions using inhouse PERL scripts. Functional annotation of the identified variants associated genes was performed using SnpEff (v 3.1) (http://snpeff.sourceforge.net/). Gene diversity31 and Polymorphic Information Content (PIC)32, which can range from 0.0 to 0.5 for bi-allelic SNPs, were manually calculated.
Population Structure and genetic diversity
Model-based ancestry estimation was obtained using the ADMIXTURE software33 with K ranging from 1 to 15. One thousand bootstrap replicates were run to estimate parameter standard errors. Ten-fold cross-validation (CV) procedure was performed and CV scores were used to determine the best K value. Non-parametric study of population structure was also carried out using AWclust34, with k-means clusters ranging from 1 to 40. The optimal number of clusters was inferred by the Bayesian Information Criterion statistics. Genetic relationships among individual accessions were finally assessed by the construction of a dendrogram based on the allele sharing distance and Ward’s clustering algorithm34. Polymorphic Information Content (PIC), Heterozygosity (H) and Gene Diversity were calculated using Power Marker software35. Multidimensional scaling (MDS) was performed using the ggscatter function implemented (https://rpkgs.datanovia.com/ggpubr/reference/ggscatter.html) in Rstudio36. Variants pruned for LD were used. The relationship between accessions was visualized by plotting scores for the first two PCs.
To identify genetically redundant accessions, a dissimilarity matrix was calculated using VCF2dis.1.0.py script. The dissimilarity index between two accessions was calculated from the entire VCF file as the proportion of unmatching alleles between two accessions: a null value (=0.00000) indicates redundant accessions.
Development of a mini-core collection set and detection of private SNPs
To identify private SNPs in ‘da serbo’ landraces compared with the others, MAF values were computed at individual SNP loci in the two groups separately. Only SNPs that showed contrasting MAF values (> 0.4 in ‘da serbo’ and < 0.2 in the other cultivars) were further analyzed for their biological function using MapMan (http://mapman.gabipd.org/). Moreover, we implemented Core Hunter v. 2.0 to build a mini-core collection based on genotypic data37. The Core Hunter software was run in R with default parameters using the variants pruned for LD (http://www.r-project.org/), which allows choosing sampling intensity and the genetic measures to be used as selection criteria.
Phenotypic analysis
Genotypes of the mini-core set were assessed for main qualitative and quantitative fruit traits. Plants were grown at the experimental organic field of the Research Centre for Vegetable and Ornamental Crops (CREA, Monsampolo Del Tronto located in the Tronto Valley of Marche Region). Traits analyzed included: (i) green shoulder (0, absent; 1, light green; 2, medium green; 3, dark green); (ii) external fruit color (1, yellow; 2, orange; 3, pink; 4, red; 5, purple; 6, brown; 7, green); (iii) fruit predominant shape (1, flat; 2, slightly flattened; 3, circular; 4, rectangular; 5, cylindrical; 6, elliptic; 7, heart shaped; 8, obovate; 9, ovate; 10, pear shaped; 11, pepper shaped); (iv) fruit firmness (1, very soft; 2, soft; 3, medium; 4, hard); (v) average fruit weight on a bulk of 18 fruits (in grams). Principal component analysis was performed using the computer package XLSTAT 2012.1.
Results
SNP discovery and functional annotation
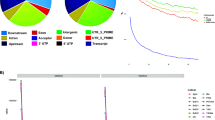
The sequencing of RADs libraries in 288 tomato genotypes produced a total of 1,036,589,344 raw reads, corresponding to an average of 3.5 million read pairs per sample. We identified 277,335 SNPs in the previous annotated genome references (SL3.0), whereas 246,936 SNPs were identified using the new reference genome (SL4.0). The decreased total genome size of SL4.0 with respect to SL3.0 (from 828,076,956 bp to 782,520,033 bp), due to the shorter chromosomes length27, led to identifying ~26.000 SNP less compared with the previous annotation. Differences in length ranged from a minimum of 292,053 bp (Chr 7) to a maximum of 6,991,656 bp (Chr 3). The greatest difference in size was observed for the unlinked SNPs (chromosome 00) which were halved in the new annotation (~10.000 SNP less). Except for chromosomes 6 and 10, the SL4.0 allowed a higher percentage of SNPs distributed along each chromosome. The total number of SNPs and their distribution along chromosomes for SL3.0 and SL4.0 are reported in Supplementary Table 2, whereas gene localization, PIC and heterozygosity within SL4.0 are reported in Table 1. Most of the identified SNPs were localized in intergenic regions spanning the short and long arms of chromosomes (Supplementary Fig. 1). The average density resulted in one SNP every 3.48 kb across the twelve chromosomes, ranging from 1.75 kb (Chr 6) to 5.05 kb (Chr 11). Across the whole set, PIC values ranged from 0.003 to 0.375 (data not shown) with a mean of 0.044. The minimum average PIC values were encountered on chromosome 7 (0.020), while the maximum value was found in Chr 4 (0.083). On average, heterozygosity was 0.049 ranging from 0.003 to 0.500 (data not shown). The observed transitions/transversions ratio was 1.23, with a total of 2,081,361 transitions and 1,688,550 transversions events (data not shown). In particular, among transitions events, C > T and G > A were the most abundant, whereas G > T and C > A abounded within transversions events (Supplementary Fig. 2). Genotypes analyzed had in average a similar number of SNPs ranging from a minimum of 1,354 SNP up to 77,501.
Three of them, harbored a higher number of variants (>50,000). In particular, the breeding line holding the greater number of SNPs (77,501) is LA2934 containing introgressions of the wild species S. pimpinellifolium. The two other Peruvian cultivars LA1313 and PI365927, held 54,166 and 68,247 SNPs, respectively. (Supplementary Fig. 3a). The observed average heterozygosity (in percentage) was also comparable within the germplasm collection with most genotypes showing values of ∼2% and only eight having values above 5% (Supplementary Fig. 3b). The SNP quality control procedure, based on successive filtering steps for call rate, minor allele frequency (MAF) (0.05) and percentage of missing values (20%) yielded a total of 32,779 high-quality SNPs in SL4.0. Raw sequence data were deposited at the National Center for Biotechnology Information Short Read Archive (http://www.ncbi.nlm.nih.gov/sra/) database under the accession number BioProject ID PRJNA638535.
Out of 246,936 SNPs, found using the latest tomato annotation (SL4.0), 10% (23.377 SNPs) were localized in 48% (2.297/4.794) of newly annotated genes. Most of these SNPs (21,991) have a possible modifier effect, followed by 721 with moderate impact, 608 with low impact and 57 with high impact. Most of these SNPs were localized in upstream gene regions (7,380), downstream gene regions (6,868), introns (5,917), 5′ UTR regions (1,006), 3′ UTR regions (820) (Supplementary Table 3). We further analyzed the genes showing sequence polymorphisms with high impact to get an interpretation of the effects of sequence changes. A total of 53 different genes were identified. Most of them had one SNP each (50 genes), whereas only three genes had two. Gene Ontology Enrichment Analysis (GOEA) was performed on this subset and no significant enrichment (FDR ≤ 0.05) was found (data not shown).
Genomic diversity and population structure
The admixture model was implemented to infer population structure. Using all SNP dataset, six different clusters were predicted (Supplementary Fig. 4a) and the population was accordingly divided in six sub-populations (K) (Fig. 2).
On the left, the dendrogram obtained through non-parametric hierarchical clustering. On the right, bar-plot describing the population admixture by the Bayesian approach. Each individual is represented by a thin horizontal line, which is partitioned into K colored segments whose length is proportional to the estimated membership coefficient (q). The population was divided into six (K = 6) groups according to the most informative K value (see Supplementary Fig. S4). Within the non-parametric model, the population was divided into two (K = 2) groups according to the most informative K value (see Supplementary Fig. S4). The colors indicate the accession membership to the groups identified with the Bayesian analysis
The first cluster (named “K1”), contained 10 cultivars from Peru, USA, Spain, France, Russia, Philippines and Falkland Islands, 23 Spanish landraces of which fourteen were ‘da serbo’, two heirlooms and one cultivar from Algeria, whereas 5 heirlooms, 4 landraces for fresh consumption (one from Italy and three from Spain), three breeding lines and one cultivar from Greece were placed in K2. The clusters K3 and K6 were the most abundant in terms of the number of genotypes, with 81 and 96, respectively. In particular, in cluster K3 most of the genotypes came from Italy (17 landraces of which 5 were ‘da serbo’), Spain (17 landraces of which 13 were ‘da serbo’) and France (8 cultivars and 3 landraces). Seventeen heirlooms, two breeding lines and 12 cultivars from various world regions were also included in K3. Similarly, 26 accessions from Italy (of which 10 were ‘da serbo’), 39 from Spain (of which 17 were ‘da serbo’), 11 heirloom, 14 cultivars, 5 unknown and one breeding line were included in K6. The cluster K4 grouped 11 ‘da serbo’ Spanish genotypes with 5 cultivars, one heirloom and a breeding line, whereas 7 landraces from Italy (2 ‘da serbo’ and 5 for fresh use), 10 cultivars, 7 heirlooms (five retrieved from the USA) and 4 breeding lines were included in K5. Nine accessions were classified as admixed, as they showed values for the highest cluster membership coefficient (qi) lower than 0.5.
The Fixation Index (FST) values, measuring the population (K) differentiation based on SNP data, are reported in Table 2. The highest differentiation was found between K2 and other subpopulations, while the lowest divergence was found for clusters 3 and 6 (FST = 0.185). Considering the average q-value at K = 6 (Fig. 3), the analysis showed how breeding lines, ‘da serbo’ types and landraces for fresh consumption were not included in the clusters I, II and IV, respectively.
We complemented the investigation of genetic structure with the non-parametric method of k-means, which identifies clusters of individuals displaying the minimal within-group variance. Linkage disequilibrium-pruned SNPs were also used in this case, in order to avoid autocorrelation among linked loci. Based on the Bayesian Information Criterion, we found that genetic variation in the collection was efficiently summarized by gap statistic suggesting a minimum number of clusters (k) equal to 2 with 42 and 246 accessions, respectively (Supplementary Fig. 4b, Fig. 2).
Notably, a general overlap was observed between the clusters identified by the two-independent analysis. Indeed, sub-clusters at K = 6 were similar to the ones identified by Admixture (Fig. 2, Supplementary Table 4), except for the inclusion of accessions classified as admixed. The first branch of the dendrogram (C1, Supplementary Fig. 5; Supplementary Table 4) contained mostly heirloom and cultivars of diverse origin, and all the accessions belonging to K2 and K5 in the admixture analysis. The remaining K subpopulations and all admixed corresponded to the second branch (C2). Applying the Bayesian method, a better resolution of the genetic relationships of the accessions according to the geographical origin and the destination of use was observed. Indeed, different subgroups (Supplementary Table 4), were formed at the similarity level of 0.2 (Supplementary Fig. 5). In several cases, heirlooms tend to group mostly with cultivars and / or landraces for fresh use. Although not in a strict manner, it was possible to observe in many cases the grouping of accessions retrieved from the same region or a clustering according to biological status. As an example, French accessions tended to cluster together (e.g., C2_A1.2.1.1.1.2) as well as some heirlooms from North America and Central Europe (e.g., C2_B2.2) or the non-European germplasm (e.g., C1B2, C2_A1.1.1.1.2, C1_B1.1.3). Many American heirlooms tended to cluster with Italian landraces for fresh consumption. Homogeneous groups were made by Spanish ‘da serbo’ and/or other landraces (i.e., C2_A1.1.1.1.1.2, C2_A1.2.2.1, C2_A1.2.2.2). Overall, ‘da serbo’ landraces tended to cluster together while the remaining landraces grouped with commercial cultivars and heirlooms.
MDS was obtained after pruning the SNPs dataset for linkage disequilibrium. The resulting 2545 markers confirmed the best number of subpopulations (K) equal to 6. The score plot in the first two components (Fig. 4a) clearly separated Spanish accessions from the rest. The Italian germplasm tended to cluster closely, although more admixtures were found with germplasm from the other European countries and North American heirlooms. Interestingly, several close relationships were found between French and Greek accessions. As expected, Latin American cultivars and Asian ones clustered more distantly from European germplasm.
a Diversity based on geographical provenance; the color of circles reflects the origin of accessions. Countries with more than 8 accessions were considered singularly, the remaining were grouped according to the main region of origin. b Diversity based on biological status. The color legends are in the figure
The second score plot based on the biological status of accessions (Fig. 4b), evidenced how the ‘da serbo’ accessions, in particular, those retrieved from Spain clustered together, while fresh consumption landraces made different tight close clusters. Heirlooms and cultivars were more dispersed than other categories, evidencing a higher variability. A close cluster including Italian landraces, heirlooms and cultivars was also observed.
We calculated r2 between pairs of SNPs using PopLDdecay to estimate the linkage disequilibrium (LD) patterns in our collection and within the different major subgroups (‘da serbo’ landraces, fresh consumption landraces, cultivars and heirlooms). Linkage disequilibrium of the entire population decays very rapidly (within <5 kb). Landraces for fresh consumption and improved cultivars showed the lowest LD values (Fig. 5). By contrast, heirlooms showed the highest LD, with r2 near to 0.4. Overall, landraces showed a higher r2 range compared to the other typologies.
Non-redundant accessions were identified, although some of them were highly similar. For example, the Italian long shelf life accessions Piennolo 21 (DS8) and Pop25 (DS9) had a dissimilarity value of 0.00158, whereas the landraces Varrone (FC33) and Laura (FC34) showed a value of 0.00204 (Supplementary Table 5). The most different accessions were the BGV5592 (DS35) and LA2934 (BL1) with a dissimilarity index of 0.445.
In order to select the most diverse genotypes that represent most of the genetic variation with minimum redundancy, we selected a mini-core collection set from the original collection. To this end, a comparison of statistic and random sampling was implemented using the R software CoreHunter, which allowed us to sample 58 different genotypes that cover the highest genetic diversity of our collection (Supplementary Fig. 6). The mini-core collection set is highly heterogeneous comprising all accession biological statuses, in fact, a total of 23 cultivars, 12 heirlooms, 14 landraces (of which nine were of the ‘da serbo’ type) and 9 breeding lines were included. (Supplementary Table 6). Furthermore, all admixture clusters were represented. Indeed, out of 58 selected genotypes, 13 belonged to K5, whereas 13 were classified as admixed. As far as the remaining genotypes are concerned, the distribution was almost homogeneous in other clusters (K1 = 4; K2 = 8; K3 = 5; K4 = 6; K6 = 9).
The group of selected accessions of the mini-core set was highly representative of the whole phenotypic variability of the collection. For the assessed fruit traits, the PCA in the first two components explained 54.59 % of the total variation (Fig. 6). The accessions were evenly distributed on the two axes of the biplot. The first component accounted for the 32.15% of the total variance and was positively correlated with external color, fruit shape, and fruit firmness and negatively correlated with the green shoulder and fruit weight. The second component accounted for the 22.44% of the variance being positively correlated only with green shoulder and external color.
Code names are reported in Supplementary Table 1. GS green shoulder, EC external color, FS fruit shape, FIR fruit firmness, FW fruit weight
Genome-wide identification of private SNPs within ‘da serbo’ germplasm
In order to detect alleles differentiating ‘da serbo’ accessions from the rest of the germplasm collection, we computed the MAF values at individual SNP loci in both sets. A total of 53 SNPs showed contrasting values in the ‘da serbo’ group (MAF > 0.4) compared with other types (MAF < 0.2) (Supplementary Table 7). Five of them, of which one localized in the intergenic regions (chr 1) and four falling in genic regions (upstream region, downstream region, intron and 3’UTR) (chrs 4, 5 and 9) showed the most contrasting MAF values (MAF > 0.4 in ‘da serbo’ and <0.1 in other groups). Most of these SNPs were localized on chr 9 (24/53), followed by chr 10 (7/53). MapMan software was also used to further understand their biological role (Supplementary Table 8). However, since an input map file for the latest S. lycopersicum annotation is not available yet, we used Mercator online tool38 to associate the 34,071 tomato proteins to MapMan bins. Then, our list was mapped to MapMan bins for data visualization. Interestingly, on the long arm of chr 9, a high frequency for the SNP located in the Kinesin-14 annotated gene was found in ‘da serbo’ genotypes (Solyc09g075480.3, bin 20). This gene encodes for motor proteins acting in vesicular transport on cortical microtubules and regulates cell wall mechanics during cell elongation in plants39. As a cytoskeleton protein, it is likely involved in response to drought stress40. In addition, it has been recently shown that this class of kinesins enters into the nucleus in response to cold stress in rice, suggesting a role in osmotic stress conditions41.
The other genes with high MAF values in ‘da serbo’ genotypes were pre-pro-cysteine proteinase (Solyc04g080960.4, bin19), Cytochrome b561 (Solyc05g041910.3, bin 35) and Guanylate-binding protein (Solyc09g065810.3, bin 35). Cysteine proteinases are known to be involved in protein maturation, degradation, and protein re-built in response to different stress conditions42, whereas Cytochromes b561 (Cyt-b561) are important for plant growth, development, and damage prevention from light excess under drought condition42. By contrast, guanylate-binding proteins act as molecular switches to regulate diverse cellular processes, including bacterial and virus responses, by alternating between active (GTP-bound) and inactive conformation (GDP-bound)43. Investigating the biological role of the remaining genes harboring SNPs with contrasting MAF values, we noticed that the genes included in the bins 17 (Protein biosynthesis), 20 (Cytoskeleton organization) and 21 (Cell wall organization) also seem to have such function in biotic stress response44. One of them (Solyc10g083600.2, bin 17) is a component of ARPF2-ARRS1 assembly complex, which play an essential role in ribosome biogenesis and probably in ribosome stress response45, whereas Solyc11g013490.2 (bin 21) was identified as hexosyltransferase non-classical Arabinogalactan-protein, a family of hydroxyproline-rich glycoproteins ubiquitous in the plant kingdom and involved in several processes, from plant growth and development to reproduction46. Two other coding regions: Solyc10g083610.2 (bin 11) and Solyc05g050280.3-Solyc05g050290.3, bin (5) were implicated in phytohormone action. In particular, Solyc10g083610 encodes for a Raf-like serine/threonine (Ser/Thr) protein kinase annotated as CTR1 (Constitutive triple response), whereas Solyc05g050280 transcribed for Jasmonate Resistant 1 (JAR1). In tomato and Arabidopsis, CTR1 has been reported to be involved in ethylene biosynthesis, being a negative regulator of signal transduction47. When the receptors perceive ethylene, CTR1 kinase activity is shut off, thereby leading to responses. By contrast, JAR1 encodes an enzyme that conjugates jasmonic acid (JA) to isoleucine, which was recently shown to be essential for pathogen defense48.
Single genes were involved in photosynthesis processes (bin 1), lipid metabolism (bin 5), chromatin organization (bin 12), DNA damage response (bin 14), RNA biosynthesis (bin 15), protein modification (bin 18), and solute transported (bin 24), whereas 15 genes did not show any functional description and were grouped in the group with not assigned function (bin 35) (Supplementary Table 8).
We also specifically looked for polymorphisms in nac-nor (non ripening) (Solyc10g006880), rin (ripening inhibitor) (Solyc05g012020) and Cnr (Colorless non-ripening) (Solyc02g077920), since they are related to a strong ripening impairment mainly related to ethylene insensitivity49,50,51. One polymorphism was found in the intergenic region of nac-nor, whereas two were detected in the rin region (one intergenic and one synonymous variant). All the variants detected passed the quality filtering. Regarding Cnr, four different variants, of which one passed quality filtering, were also identified. Only one was found in the upstream region and the minor allele was identified in 13 cultivars, 6 heirlooms, 4 unknown, 3 landraces and in one ‘da serbo’ genotype (data not shown).
Discussion
Genomic investigation of germplasm diversity
Tapping the diversity of crops is a prerequisite for efficient genetic improvement, conservation, and management of germplasm resources. The present work was aimed to study the genomic diversity of a diverse panel of cultivated tomato germplasm and to identify loci under selection and putatively responsible for the syndrome of differences related to LSL cultivars.
As a model species, tomato has been widely exploited for breeding, genomics and biology researches, being among the first vegetable crops having the whole genome sequence released. Within tomato populations genetics, efforts have been principally addressed toward the dissection of the genetic basis of advanced recombinant materials, elite germplasm, and wild species52,53,54,55. Approaches to investigate the diversity of traditional varieties including vintage and landraces have been performed with a low13,14,15,16,17,56, or medium marker-throughput18,19,20 in collections not larger than a hundred of accessions13,14,15,17,19,20 with few exceptions comprising a large proportion of improved cultivars16,18,56. These studies ascertained population stratification due to selection history, geographic adaptation and market destination15,16,17,18,20. Landraces were reported to have a lower allelic diversity compared to contemporary varieties, although the former may be richer in rare alleles17. In fact, modern varieties are developed through introgression breeding increasing the allelic assortment in established genetic backgrounds18,54. On the contrary, local varieties have been less exploited in terms of breeding and cultivation, having been replaced by high yielding varieties, particularly in intensive agricultural systems57. Therefore, the genetic composition of landraces is mainly due to selection driven by local practices in specific agricultural systems as well as environmental factors.
The gene pool could contribute to broadening and improving the genetic base of current modern varieties. A deeper exploration would give more insight to be exploited for tomato improvement. In this regard, we applied ddRAD sequencing protocol detecting 246,936 polymorphic sites and of 32,779 high-quality SNPs used to infer population structure and phylogenetic relationships using parametric and non-parametric computations. The abundance and whole-genome coverage provided by ddRAD based SNPs provide a high measure of genomic diversity in particular at the intra-specific level, where fewer polymorphisms occur compared to the inter-specific level58. The occurring heterozygosity across the whole collection is in agreement with previous evidence reporting values ranging from 1 to 4% in traditional varieties and higher values in the improved varieties3.
The results indicated a certain differentiation within the collection on the basis of biological status and provenance of accessions. Although a certain level of admixture was found between contemporary varieties, heirlooms, and landraces, the ‘da serbo’ LSL types were genetically more distinct. Moreover, a higher number of structured subgroups were found in landraces with respect to contemporary varieties. Indeed, although LSL and fresh consumption landraces represent 52% of the whole collection, fewer alleles were shared with the other typologies. Our observations are in agreement with previous researches that do not report subpopulation structures in varieties developed from advanced breeding programs such as cultivars and heirlooms, evidencing how the subpopulation structure is determined by natural selection and breeding history15. The number of K clusters identified can be considered consistent to define the population structure. Wang and colleagues54, and Bauchet and collaborators56, identified population structure within the typologies studied detecting 9 and 6 clusters in collections of 348 and 300 individuals, respectively. We also investigated the level of redundancy among accessions. The lowest values (<0.002 and <0.003) were found within landraces for fresh consumption as well within LSL, confirming how in situ conservation better maintains the genetic identity of tomato landraces due to reduced gene flow and genetic drift59. The MDS plot confirmed a low gene flow between LSL ‘da serbo’ materials and the other groups of accessions. Furthermore, allele exchange was observed between Italian landraces, heirlooms and elite cultivars, suggesting their greater exploitation in breeding programs. An LD decay within a few kilobases was observed in the 288 accessions studied. Robbins and colleagues evidenced an overall LD decay within a few cM with differences between the fresh market and processing market types14. The same trend was observed by Bauchet and collaborators reporting an LD fluctuating according to the genetic groups studied54. Both authors highlighted the breeding history and gene recombination as the main factors influencing the observed LD values. Moreover, the homozygosity level and mating system are two additional components affecting LD in crops. In fact, in outcrossing species, the LD can decay within hundreds of bp while for highly selfing species the LD may extend to 10 kb60. Moreover, accessions with different biological statuses can exhibit diverse LD values. As an example, in maize, the LD decay of landraces has been proven to be 100-fold less than in elite lines61. The same trend has been observed in heterogenous germplasm of barley61. The mini-core collection set of 58 tomato accessions was built with the objective to maximize genetic diversity with a reduced number of accessions. This mini-core collection represents the whole genetic and phenotypic diversity of the collection studied and could be useful to identify traits of agronomic and qualitative interest in tomato. Phenotyping is a major bottleneck to address in breeding due to costs and time required62. The establishment of a mini-core collection encompassing most of the diversity could be a valid solution to deeply explore the phenotypic diversity, overcoming the bottleneck of using very large collections to identify key traits for genetic improvement.
Variants falling in gene regions characterizing ‘da serbo’ genotypes
Although the LSL are usually related to higher phenotypic performance and stress tolerance, the molecular mechanisms underlying the extended shelf-life and tolerance to drought cultivation conditions are not fully understood. To date, the LSL phenotype has been related to different mutations in genes such as alc, nor and cnr12,63,64,65,66. In particular, the alc mutation has been found in Spanish LSL ‘da serbo’ genotypes, which are commonly called ‘de penjar’ in Catalonia and Valencia and ‘de ramellet’ in the Balearic Islands12,67. In the ‘Alcobaça’ landrace the alc mutation was associated to a reduction of ethylene production (i.e., 25% less than ‘Rutgers’ cultivar65, and mutations alc-like have been suggested to play the same role in Italian ‘da serbo’ LSL landraces10. Similar results comparing the ethylene emission in ‘Ailsa Craig’ and four different ‘de penjar’ accessions, revealed that ethylene emission may be not only related to alc mutation68. A genetic screening within a collection of Italian ‘da serbo’ LSL landraces20, found high variation in ethylene-responsive genes, although no polymorphisms in the ethylene-dependent non-ripening (nor) transcription factor were found, endorsing the idea that genetic determinants of the LSL fruit phenotype may be different between Italian and Spanish landraces. In our study, we identified few polymorphisms in nor-nac, rin and Cnr, mainly localized in intergenic regions except for one variant detected in Cnr. Previously, Casals and collaborators12, identified a polymorphism in the second exon of the gene nac-nor, consisting of a replacement of thymine by adenine in the coding sequence responsible of the alcobaca (alc) mutation. In the same study12, the analysis of the rin region excluded its involvement as a cause of long shelf life in the Spanish ‘de penjar’ varietal types. Bota and colleagues67, also detected the presence of the alc mutation in a collection of Spanish ‘de ramellet” accessions from Majorca island. Our results suggest that, in addition to alc, other variants in rin, nac-nor and Cnr might be involved in the development of LSL ‘da serbo’ shelf life. We cannot exclude that our approach did not cover the region of alc previously studied12 due to the reduced genome representation occurring with ddRAD genotyping. Therefore, target sequencing may be used to verify the presence of such variants in this gene. Some SNPs were instead identified in ethylene signal transduction genes. Indeed, two SNPs were localized in intron regions of Constitutive Triple Response 1 (CTR1), an important negative regulator of ethylene signaling. CTR1 has a central position in the ethylene-response pathway, acting downstream of the ethylene receptors and upstream of EIN2, a membrane-integrated metal transporter-like protein69. In the presence of ethylene, the function of CTR1 is inhibited and EIN2 can translocate in the nucleus inducing the expression of ethylene-responsive genes70. By contrast, in absence of ethylene, CTR1 can phosphorylate EIN2, which becomes inactive and does not induce its response.
In addition, it has been shown that CTR1 is also involved in salt tolerance71. A mutant inactivating the allele-kinase in CTR1 (ctr1-1), exhibited an increased salt tolerance during the germination and post-germination stages, suggesting a role of CTR1 as negative regulator71. Therefore, the two variants identified in CTR1 intron might play a dual role. On the one hand, they may act as enhancers, as already described in many plants such as Arabidopsis and rice72,73 In this scenario, CTR1 might be expressed at a higher level even in the presence of ethylene, inhibiting the expression of related genes, which may lead to the extended shelf life of ‘da serbo’ accessions. On the other hand, intronic variants may induce aberrant splicing, resulting in defective protein products. In this case, CTR1 might have an abnormal structure which can increase the tolerance to osmotic stresses such as salt and drought. However, since much remains unclear in respect to the signal transduction pathways from CTR1 to EIN2 in different plant species, other studies are ongoing to clarify the effect of the two variants identified.
Interesting variants were also identified in JASMONATE RESISTANT 1 (JAR1), a coding-protein having a key role in the biosynthesis of jasmonate48. A variety of functions are associated with JAR1 including resistance to biotic stress. Previous studies evidenced a correlation between jasmonate and reduced ethylene emission in ‘de Peniar’ accessions68, suggesting how the LSL fruit phenotype may be also related to non-ethylene mediated ripening regulation as in the non-ripening tomato mutants10,74,75.
In agreement with surveys of Tranchida-Lombardo and colleagues20, who identified a kinesin harboring a non-synonymous SNP in Italian ‘da serbo’, we found a SNP in the 3’-UTR region of Solyc09g075480.3. The underlying gene has high similarity to kinesin-14, which has a role in cell wall synthesis, acting in the deposition and orientation of cellulose microfibrils76. Other interesting variants were identified in cysteine proteinases (CP), cytochromes b561 (Cyt-b561) and guanylate-binding proteins (GBP). CP are mainly involved in protein rebuilt following external stimuli and in the protection against misfolded or damaged proteins77. The process could involve the rebuilding of cold and heat shock proteins, dehydration-induced proteins and pathogenesis-related proteins. Examples were reported in pea and Arabidopsis for which CP induced by water deficit are responsive to salt stress78,79,80. In tomato, CP are reported to have an active role in biotic and abiotic stress defense and suppression of autonecrosis81,82. Similarly, Cytochromes b561 are intrinsic membrane proteins involved in ascorbate regeneration known for its detoxification during aerobic metabolism and under stress conditions83,84. In watermelon, CYB561 has been reported to be induced by drought and high-light stress, suggesting a role for thermal dissipation of excess light energy, through functional interaction with apoplastic ASC oxidase85. In addition, through the modulation of levels of reactive oxygen species, ascorbate with its CYB561 activities are also implicated in the control of cell expansion, cell division, and programmed cell death85. Regarding GBP, it has been demonstrated how intravacuolar bacteria of infected cells stimulate the recruitment of antimicrobial guanylate binding proteins as part of a coordinated host defense program86.
Overall, these results support that mutations different from those previously known in genes such nac-nor, rin and Cnr may also be related to long shelf life and higher stress tolerance in Mediterranean ‘da serbo’ LSL tomato germplasm. Our results indicate putative genes under selection. The gained information suggests the possibility to further investigate specific genomic regions for the mining of novel alleles to be exploited for the improvement of long shelf life and resilient cultivars.
Conclusions
Using ddRAD-seq for high throughput SNP discovery and the latest tomato genome version for annotation, we showed for the first-time new clues in a large fraction of Mediterranean ‘da serbo’ gene pool, providing furthermore, a list of 2297 newly and never studied target genes with different functions and harboring SNPs. The genomic analysis highlighted a geographical footprint distinguishing the landraces developed in the Mediterranean basin secondary center of diversity of tomato. We then investigated the level of variation underlying long shelf-life landraces, detecting SNPs falling in gene regions other than the previously known alc, nac-nor and cnr. Among the newly identified genes, those involved in biotic and abiotic stress conditions as well as in ethylene and jasmonate pathways may be worth being studied further. We finally provided breeders with a mini-core collection composed of the most diverse genotypes of our germplasm. Overall, the collections studied constitute a promising reservoir of genes for traits of interest, which could be further explored in genome-wide association studies and exploited in new tomato precision breeding programs.
References
Faostat 2018 http://www.fao.org/
Jenkins, J. A. The origin of the cultivated tomato. Econ. Bot. 2, 379–392 (1948).
Blanca, J. et al. Variation revealed by SNP genotyping and morphology provides insight into the origin of the tomato. PLoS ONE 7, e48198 (2012).
Razifard, H. et al. Genomic evidence for complex domestication history of the cultivated tomato in Latin America. Mol. Biol. Evol. 37, 1118–1132 (2020).
Bauchet, G. & Causse, M., Genetic diversity in tomato (Solanum lycopersicum) and its wild relatives. In Genetic Diversity in Plants (INTECH Open Access Publisher, 2012).
Bai, Y. & Lindhout, P. Domestication and breeding of tomatoes: what have we gained and what can we gain in the future? Ann. Bot. 100, 1085–1094 (2007).
Dwivedi, S., Goldman, I. & Ortiz, R. Pursuing the potential of heirloom cultivars to improve adaptation, nutritional, and culinary features of food crops. Agronomy 9, 441 (2019).
Casañas, F., Simó, J., Casals, J. & Prohens, J. Toward an evolved concept of landrace. Front. Plant Sci. 8, 145 (2017).
Klee, H. J. & Tieman, D. M. The genetics of fruit flavour preferences. Nat. Rev. Genet. 19, 347–356 (2018).
Conesa, M. À., Fullana-Pericàs, M., Granell, A. & Galmés, J. Mediterranean long shelf-life landraces: an untapped genetic resource for tomato improvement. Front. Plant Sci. 10, 1651 (2020).
Casals, J., Martí, R., Casañas, F. & Cebolla, J. Sugar-and-acid profile of Penjar tomatoes and its evolution during storage. Sci. Agric. 72, 314–321 (2015).
Casals, J. et al. Genetic basis of long shelf life and variability into Penjar tomato. Genet. Resour. Crop Evol. 59, 219–229 (2012).
van Berloo, R. et al. Diversity and linkage disequilibrium analysis within a selected set of cultivated tomatoes. Theor. Appl. Genet. 117, 89–101(2008).
Robbins, M. D. et al. Mapping and linkage disequilibrium analysis with a genome-wide collection of SNPs that detect polymorphism in cultivated tomato. J. Exp. Bot. 62, 1831–1845 (2011).
Sim, S. C., Robbins, M. D., Deynze, A. V., Michel, A. P. & Francis, D. M. Population structure and genetic differentiation associated with breeding history and selection in tomato (Solanum lycopersicum L.). Heredity 106, 927–935 (2011).
Corrado, G., Piffanelli, P., Caramante, M., Coppola, M. & Rao, R. SNP genotyping reveals genetic diversity between cultivated landraces and contemporary varieties of tomato. BMC Genomics 14, 835 (2013).
Corrado, G., Caramante, M., Piffanelli, P. & Rao, R. Genetic diversity in Italian tomato landraces: implications for the development of a core collection. Sci. Hortic. 168, 138–144 (2014).
Sim, S. C. et al. High-density SNP genotyping of tomato (Solanum lycopersicum L.) reveals patterns of genetic variation due to breeding. PloS ONE 7, e45520 (2012).
Sacco, A. et al. Exploring a tomato landraces collection for fruit-related traits by the aid of a high-throughput genomic platform. PLoS ONE 10, e0137139 (2015).
Tranchida-Lombardo, V. et al. Genetic diversity in a collection of Italian long storage tomato landraces as revealed by SNP markers array. Plant Biosyst. 153, 288–297 (2019).
Pérez de, Castro et al. Application of genomic tools in plant breeding. Curr. Genomics 13, 179–195 (2012).
Peterson, B. K. et al. Double digest RADseq: an inexpensive method for de novo SNP discovery and genotyping in model and non-model species. PLoS ONE 7, e37135 (2012).
Shirasawa, K., Hirakawa, H. & Isobe, S. Analytical workflow of double-digest restriction site-associated DNA sequencing based on empirical and in silico optimization in tomato. DNA Res. 23, 145–153 (2016).
Yang, G. et al. Development of a universal and simplified ddRAD library preparation approach for SNP discovery and genotyping in angiosperm plants. Plant Met. 12, 39 (2016).
Okada, Y. et al. Genome-wide association studies (GWAS) for yield and weevil resistance in sweet potato (Ipomoea batatas (L.) Lam. Plant Cell Rep. 38, 1383–1392 (2019).
The Tomato Genome Consortium. The tomato gene sequence provides insights into fleshy fruit evolution. Nature 485, 635–641 (2012).
Hosmani, P. S. et al. An improved de novo assembly and annotation of the tomato reference genome using single-molecule sequencing, Hi-C proximity ligation and optical maps. Preprint at https://www.biorxiv.org/content/10.1101/767764v1 (2020).
Aguirre, N. et al. Optimizing ddRADseq in non-model species: a case study in Eucalyptus dunnii maiden. Agronomy 9, 484 (2019).
Catchen, J., Hohenlohe, P., Bassham, S., Amores, A. & Cresko, W. Stacks: an analysis tool set for population genomics. Mol. Ecol. 22, 3124–3140 (2013).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Weir, B. S. Genetic Data Analysis II (Sinauer Associates Inc., Sunderland, MA, 1996).
Botstein, D., White, R. L., Skolnick, M. & Davis, R. W. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am. J. Hum. Genet. 32, 314–331 (1980).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Gao, X. & Starmer, J. D. AWclust: point‐and‐click software for non‐parametric population structure analysis. BMC Bioinformatics 9, 77 (2008).
Liu, K. & Muse, S. V. PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21, 2128–2129 (2005).
RStudio Team RStudio: Integrated Development for R (RStudio, Inc., Boston, MA, 2016).
De Beukelaer, H., Smýkal, P., Davenport, G. F., Fack, V. & Core Hunter, I. I. fast core subset selection based on multiple genetic diversity measures using Mixed Replica search. BMC Bioinforma. 13, 312 (2012).
Schwacke, R. et al. MapMan4: a refined protein classification and annotation framework applicable to multi-omics data analysis. Mol. Plant 12, 879–892 (2019).
Kong, Z. et al. Kinesin-4 functions in vesicular transport on cortical microtubules and regulates cell wall mechanics during cell elongation in plants. Mol. Plant 8, 1011–1023 (2015).
Wang, X., Cai, X., Xu, C., Wang, Q. & Dai, S. Drought-responsive mechanisms in plant leaves revealed by proteomics. Int. J. Mol. Sci. 17, 1706 (2016).
Xu, X., Walter, W. J., Liu, Q., Machens, I. & Nick, P. A rice class-XIV kinesin enters the nucleus in response to cold. Sci. Rep. 8, 3588 (2018).
Opiyo, S. O. & Moriyama, E. N. Mining Cytochrome b561 proteins from plant genomes. Int. J. Bioinform. Res. Appl. 6, 209–221 (2010).
Lee, S. et al. The small GTPase, nucleolar GTP-binding protein 1 (NOG1), has a novel role in plant innate immunity. Sci. Rep. 7, 9260 (2017).
Usadel, B. et al. Extension of the visualization tool MapMan to allow statistical analysis of arrays, display of corresponding genes, and comparison with known responses. Plant Physiol. 138, 1195–1204 (2005).
Sáez-Vásquez, J. & Delseny, M. Ribosome biogenesis in plants: from functional 45S ribosomal DNA organization to ribosome assembly factors. Plant Cell 31, 1945–1967 (2019).
Mareri, L., Romi, M. & Cai, G. Arabinogalactan proteins: actors or spectators during abiotic and biotic stress in plants? Plant Biosyst. 153, 173–185 (2018).
Zhong, S., Chang, C. in Ethylene signalling: the CTR1 protein kinase, Vol. 44 (eds Oxford, UK: Wiley-Blackwell) ch. 6 (Annual Plant Reviews, 2012).
Suza, W. P. & Staswick, P. E. The role of JAR1 in jasmonoyl-L-isoleucine production in Arabidopsis wound response. Planta 227, 1221–1232 (2008).
Barry, C. S. & Giovannoni, J. J. Ethylene and fruit ripening. J. Plant Growth Regul. 26, 143–159 (2007).
Giovannoni, J. J. Fruit ripening mutants yield insights into ripening control. Curr. Opin. Plant Biol. 10, 283–289 (2007).
Wang, R. et al. The rin, nor and Cnr spontaneous mutations inhibit tomato fruit ripening in additive and epistatic manners. Plant Sci. 294, 110436 (2020).
Albrecht, E., Escobar, M. & Chetelat, R. Genetic diversity and population structure in the tomato-like nightshades Solanum lycopersicoides and S. sitiens. Ann. Bot. 105, 535–554 (2010).
Xu, J. et al. Phenotypic diversity and association mapping for fruit quality traits in cultivated tomato and related species. Theor. Appl. Genet. 126, 567–581 (2013).
Bauchet, G. et al. Use of modern tomato breeding germplasm for deciphering the genetic control of agronomical traits by Genome Wide Association study. Theor. Appl. Genet. 130, 875–889 (2017).
Rothan, C., Diouf, I. & Causse, M. Trait discovery and editing in tomato. Plant J. 97, 73–90 (2018).
Wang, T. et al. Analysis of genetic diversity and population structure in a tomato (Solanum lycopersicum L.) germplasm collection based on single nucleotide polymorphism. Genet. Mol. Res. 15, 1–12 (2016).
Massaretto, I. L. et al. Recovering tomato landraces to simultaneously improve fruit yield and nutritional quality against salt stress. Front. Plant Sci. 9, 1778 (2018).
Aflitos, S. et al. Exploring genetic variation in the tomato (Solanum section Lycopersicon) clade by whole-genome sequencing. Plant J. 80, 136–148 (2014).
Leimu, R., Mutikainen, P. I. A., Koricheva, J. & Fischer, M. How general are positive relationships between plant population size, fitness and genetic variation? J. Ecol. 94, 942–952 (2006).
Mackay, I. & Powell, W. Methods for linkage disequilibrium mapping in crops. Trends Plant Sci. 12, 57–63 (2007).
Ersoz, E. S., Yu, J., Buckler, E. S. Applications of linkage disequilibrium and association mapping in Genomics-assisted crop improvement, Vol. 1 (eds. Varshney R. K., Tuberosa R.) (Dordrecht, Springer, 2008).
D’Agostino, N. & Tripodi, P. NGS-based genotyping, high-throughput phenotyping and genome-wide association studies laid the foundations for next-generation breeding in horticultural crops. Diversity 9, 38 (2017).
Kopeliovitch, E., Rabinowitch, H. D., Mizrahi, Y. & Kedar, N. Mode of inheritance of Alcobaca, a tomato fruit-ripening mutant. Euphytica 30, 223–225 (1981).
Lobo, M., Bassett, M. J. & Hannah, L. C. Inheritance and characterization of the fruit ripening mutation in ‘alcobaca’ tomato. J. Am. Soc. Hortic. Sci. 109, 741–745 (1984).
Mutschler, M., Guttieri, M., Kinzer, S., Grierson, D. & Tucker, G. Changes in ripening-related processes in tomato conditioned by the alc mutant. Theor. Appl. Genet. 76, 285–292 (1988).
Conesa, M. A. et al. The postharvest tomato fruit quality of long shelf-life Mediterranean landraces is substantially influenced by irrigation regimes. Postharvest Biol. Technol. 93, 114–121 (2014).
Bota, J. et al. Characterization of a landrace collection for Tomàtiga de Ramellet (Solanum lycopersicum L.) from the Balearic Islands. Genet. Resour. Crop Evol. 61, 1131–1146 (2014).
Kumar, R., Tamboli, V., Sharma, R. & Sreelakshmi, Y. NAC-NOR mutations in tomato Penjar accessions attenuate multiple metabolic processes and prolong the fruit shelf life. Food Chem. 259, 234–244 (2018).
Cho, Y. H. & Yoo, S. D. Novel connections and gaps in ethylene signaling from the ER membrane to the nucleus. Front. Plant Sci. 5, 733 (2015).
Ju, C. et al. CTR1 phosphorylates the central regulator EIN2 to control ethylene hormone signaling from the ER membrane to the nucleus in Arabidopsis. Proc. Natl. Acad. Sci. USA 109, 19486–19491 (2012).
Achard, P. et al. Integration of plant responses to environmentally activated phytohormonal signals. Science 311, 91–94 (2006).
Morello, L., Giani, S., Troina, F. & Breviario, D. Testing the IMEter on rice introns and other aspects of intron-mediated enhancement of gene expression. J. Exp. Bot. 62, 533–544 (2011).
Rose, A. B., Carter, A., Korf, I. & Kojima, N. Intron sequences that stimulate gene expression in Arabidopsis. Plant Mol. Biol. 92, 337–346 (2016).
Moore, S., Vrebalov, J., Payton, P. & Giovannoni, J. J. Use of genomics tools to isolate key ripening genes and analyse fruit maturation in tomato. J. Exp. Bot. 53, 2023–2030 (2002).
Vrebalov, J. et al. A MADS-box gene necessary for fruit ripening at the tomato ripening-inhibitor (rin) locus. Science 296, 343–346 (2002).
Zhou, J. L., Qiu, J. & Ye, Z. H. Alteration in secondary wall deposition by overexpression of the Fragile Fiber1 kinesin-like protein in Arabidopsis. J. Integr. Plant Biol. 49, 1235–1243 (2007).
Vallés, D. et al. A cysteine protease isolated from ripe fruits of Solanum granulosoleprosum (Solanaceae). Protein J. 27, 267 (2008).
Guerrero, F. D., Jones, J. T. & Mullet, J. E. Turgor-responsive gene transcription and RNA levels increase rapidly when pea shoots are wilted. Sequence and expression of three inducible genes. Plant Mol. Biol. 15, 11–26 (1990).
Yamaguchi-Shinozaki, K., Koizumi, M., Urao, S. & Shinozaki, K. Molecular cloning and characterization of 9 cDNAs for genes that are responsive to desiccation in Arabidopsis thaliana: sequence analysis of one cDNA clone that encodes a putative transmembrane channel protein. Plant Cell Physiol. 33, 217–224 (1992).
Koizumi, M., Yamaguchi-Shinozaki, K., Tsuji, H. & Shinozaki, K. Structure and expression of two genes that encode distinct drought-inducible cysteine proteinases in Arabidopsis thaliana. Gene 129, 175–182 (1993).
Krüger, J. et al. A tomato cysteine protease required for Cf-2-dependent disease resistance and suppression of autonecrosis. Science 296, 744–747 (2002).
Fan, Y., Yang, W., Yan, Q., Chen, C. & Li, J. Genome-wide identification and expression analysis of the protease inhibitor gene families in tomato. Genes 11, 1 (2020).
Noctor, G. & Foyer, C. H. Ascorbate and glutathione: keeping active oxygen under control. Annu. Rev. Plant Physiol. 49, 249–279 (1998).
Mittler, R. Oxidative stress, antioxidants and stress tolerance. Trends Plant Sci. 7, 405–410 (2002).
Nanasato, Y., Akashi, K. & Yokota, A. Co-expression of cytochrome b561 and ascorbate oxidase in leaves of wild watermelon under drought and high light conditions. Plant Cell Physiol. 46, 1515–1524 (2005).
Feeley, E. M. et al. Galectin-3 directs antimicrobial guanylate binding proteins to vacuoles furnished with bacterial secretion systems. Proc. Natl. Acad. Sci. USA 114, E1698–E1706 (2017).
Acknowledgements
The authors thank the European Union Horizon 2020 Research and Innovation program for funding this research under grant agreement No 774244 (Breeding for Resilient, Efficient and Sustainable Organic Vegetable Production; BRESOV).
Author information
Authors and Affiliations
Contributions
P.T., J.P., and T.C. conceived the project. M.J.D., G.C., S. Sestili, and S. Soler provided germplasm. P.T. performed the genetic analysis, S.E. performed the bioinformatic analysis, G.C., M.J.D., S. Sestili, and S. Soler provided phenotypic data of the core set, G.C. provided fruit photos. S.E. and P.T. jointly wrote the manuscript, T.C., M.J.D., and J.P. carefully revised the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Esposito, S., Cardi, T., Campanelli, G. et al. ddRAD sequencing-based genotyping for population structure analysis in cultivated tomato provides new insights into the genomic diversity of Mediterranean ‘da serbo’ type long shelf-life germplasm. Hortic Res 7, 134 (2020). https://doi.org/10.1038/s41438-020-00353-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41438-020-00353-6
This article is cited by
-
Genomics empowering conservation action and improvement of celery in the face of climate change
Planta (2024)
-
Tomato genomic prediction for good performance under high-temperature and identification of loci involved in thermotolerance response
Horticulture Research (2021)
-
Genome wide association mapping for agronomic, fruit quality, and root architectural traits in tomato under organic farming conditions
BMC Plant Biology (2021)